

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
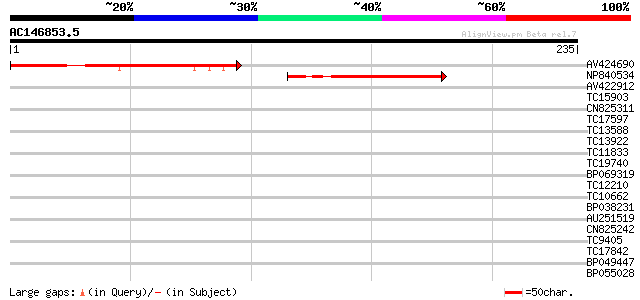
Query= AC146853.5 - phase: 0 /pseudo
(235 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV424690 121 1e-28
NP840534 hypothetical protein [Lotus corniculatus var. japonicus] 79 9e-16
AV422912 34 0.019
TC15903 similar to UP|Q40724 (Q40724) Transcriptional activator ... 33 0.043
CN825311 33 0.056
TC17597 similar to UP|AZF1_YEAST (P41696) Asparagine-rich zinc f... 32 0.073
TC13588 similar to UP|GRP2_PHAVU (P10496) Glycine-rich cell wall... 32 0.12
TC13922 UP|Q99NH6 (Q99NH6) bZIP protein ATF7, partial (5%) 32 0.12
TC11833 32 0.12
TC19740 similar to UP|FKH_DROME (P14734) Fork head protein, part... 31 0.16
BP069319 31 0.16
TC12210 similar to UP|Q867T7 (Q867T7) P67-like superoxide-genera... 31 0.21
TC10662 homologue to UP|CWP1_YEAST (P28319) Cell wall protein CW... 30 0.28
BP038231 30 0.36
AU251519 30 0.47
CN825242 30 0.47
TC9405 homologue to UP|Q84V96 (Q84V96) Aldehyde dehydrogenase 1 ... 29 0.62
TC17842 similar to PIR|B86422|B86422 F1N18.10 protein - Arabidop... 29 0.62
BP049447 29 0.81
BP055028 28 1.1
>AV424690
Length = 416
Score = 121 bits (303), Expect = 1e-28
Identities = 72/115 (62%), Positives = 78/115 (67%), Gaps = 19/115 (16%)
Frame = +3
Query: 1 MSMLGLRDLVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDHNSN-HSLPSSASLSVGFGI 59
MSMLGLRDLV IAP+PSSL H HQ +Q QPIS D NSN + LPSSASLSVGFGI
Sbjct: 93 MSMLGLRDLVFIAPNPSSLHHQHQ-------HQVQPISSDPNSNLNPLPSSASLSVGFGI 251
Query: 60 FPLLTATPCMPQQQSQ------NNEVQE-----NPSNNN-------NFWNLRMCP 96
FPLLTATPCMPQQQ Q NNEVQ N +N N N+WNL+MCP
Sbjct: 252 FPLLTATPCMPQQQHQQHPQNNNNEVQNQECAANNTNTNTNTNTATNYWNLKMCP 416
>NP840534 hypothetical protein [Lotus corniculatus var. japonicus]
Length = 741
Score = 78.6 bits (192), Expect = 9e-16
Identities = 35/66 (53%), Positives = 46/66 (69%)
Frame = +1
Query: 116 IAMMESEENGVYGSEYRVCQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHLKSTWIPSTRR 175
++MM +E V GS CQ+CGN+AKK C + RCRTCC +G+ C TH++STWIP RR
Sbjct: 1 MSMMLGQE--VKGSR---CQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRR 165
Query: 176 REREVE 181
R R +E
Sbjct: 166 RHRLME 183
>AV422912
Length = 460
Score = 34.3 bits (77), Expect = 0.019
Identities = 11/32 (34%), Positives = 17/32 (52%)
Frame = +3
Query: 134 CQDCGNRAKKDCVFRRCRTCCKGRGYDCSTHL 165
C+ CGN A+ C + C++CC C H+
Sbjct: 165 CKQCGNVARSRCPYECCKSCCARNQNPCHIHV 260
>TC15903 similar to UP|Q40724 (Q40724) Transcriptional activator protein
(RITA-1 protein), partial (21%)
Length = 603
Score = 33.1 bits (74), Expect = 0.043
Identities = 24/59 (40%), Positives = 27/59 (45%), Gaps = 8/59 (13%)
Frame = -3
Query: 137 CGNRAKKDC--VFRRCRTCCKGRGY------DCSTHLKSTWIPSTRRREREVEMFAGGG 187
CG R K C + RRCR GR D +HLK + P R RERE GGG
Sbjct: 187 CGTRRSKRC*IMHRRCRDSGGGRNRG*ETRPDLQSHLKPSAPPHRRDRERE----RGGG 23
>CN825311
Length = 522
Score = 32.7 bits (73), Expect = 0.056
Identities = 15/30 (50%), Positives = 19/30 (63%)
Frame = -2
Query: 21 HHHQQNQNQNQNQNQPISHDHNSNHSLPSS 50
HHH +QNQNQN + H H+ +HS P S
Sbjct: 98 HHHHHHQNQNQNFH---FHFHHHSHSKPFS 18
>TC17597 similar to UP|AZF1_YEAST (P41696) Asparagine-rich zinc finger
protein AZF1, partial (3%)
Length = 399
Score = 32.3 bits (72), Expect = 0.073
Identities = 23/75 (30%), Positives = 35/75 (46%), Gaps = 17/75 (22%)
Frame = +1
Query: 19 LQHHHQQNQNQNQNQNQPIS----------------HDHNSNHSLPSSASLSVGFGIFPL 62
+Q + + N NQNQNQNQ S +++N+N+ + SS S S+ P
Sbjct: 154 IQDNREINSNQNQNQNQSSSSSPMNSPSPRSNASNNNNYNNNNCMSSSNSNSISQIQIPT 333
Query: 63 LTATP-CMPQQQSQN 76
TP +P+ S N
Sbjct: 334 SPHTPKTLPRSDSNN 378
>TC13588 similar to UP|GRP2_PHAVU (P10496) Glycine-rich cell wall structural
protein 1.8 precursor (GRP 1.8), partial (6%)
Length = 517
Score = 31.6 bits (70), Expect = 0.12
Identities = 20/55 (36%), Positives = 25/55 (45%), Gaps = 8/55 (14%)
Frame = -2
Query: 14 PSPSSLQHHHQQNQNQNQ--------NQNQPISHDHNSNHSLPSSASLSVGFGIF 60
P P + HHH Q+ Q Q N P HD +S+H L AS + G G F
Sbjct: 357 PHPHAHHHHHYQSPPQQQQTLPIRAPNPQFPKPHDPSSHHYLYPFASSARGGGPF 193
>TC13922 UP|Q99NH6 (Q99NH6) bZIP protein ATF7, partial (5%)
Length = 511
Score = 31.6 bits (70), Expect = 0.12
Identities = 12/16 (75%), Positives = 13/16 (81%)
Frame = +3
Query: 19 LQHHHQQNQNQNQNQN 34
L HHHQ+NQN N NQN
Sbjct: 126 LLHHHQKNQNWNHNQN 173
>TC11833
Length = 562
Score = 31.6 bits (70), Expect = 0.12
Identities = 21/60 (35%), Positives = 30/60 (50%), Gaps = 7/60 (11%)
Frame = +2
Query: 2 SMLGLRDLVLIAPSPSSLQHHHQQNQNQNQNQNQPISH-------DHNSNHSLPSSASLS 54
S++G R L + +Q+H QNQNQNQ QN +S+ N +H+ P S S
Sbjct: 251 SIIGKRTL-----AEFQIQNHQNQNQNQNQIQNPVLSNLLLRSVKPRNFHHASPIDFSTS 415
>TC19740 similar to UP|FKH_DROME (P14734) Fork head protein, partial (4%)
Length = 622
Score = 31.2 bits (69), Expect = 0.16
Identities = 13/27 (48%), Positives = 16/27 (59%)
Frame = +3
Query: 15 SPSSLQHHHQQNQNQNQNQNQPISHDH 41
+P QH HQQ+Q+Q Q Q QP H
Sbjct: 111 NPQHQQHQHQQHQHQFQQQQQPQHFHH 191
>BP069319
Length = 494
Score = 31.2 bits (69), Expect = 0.16
Identities = 14/23 (60%), Positives = 16/23 (68%)
Frame = -2
Query: 212 SHSSNSNGTTPKSFATSSCHQGA 234
SH+S SN T P+SF TSS H A
Sbjct: 484 SHTSTSNTTPPRSFETSSSHPDA 416
>TC12210 similar to UP|Q867T7 (Q867T7) P67-like superoxide-generating NADPH
oxidase, partial (3%)
Length = 416
Score = 30.8 bits (68), Expect = 0.21
Identities = 18/65 (27%), Positives = 26/65 (39%), Gaps = 6/65 (9%)
Frame = +1
Query: 20 QHHHQQNQNQNQNQNQPISHDHNSNH-SLPSSASL-----SVGFGIFPLLTATPCMPQQQ 73
+HHHQQ QN + + H H L S S+ S+ F +P + P PQ
Sbjct: 217 KHHHQQEHEQNHHYQGANPYQHTYQHPQLVRSPSVLQRLKSINFYSYPFRSQDPSHPQTH 396
Query: 74 SQNNE 78
+
Sbjct: 397 EHEEQ 411
>TC10662 homologue to UP|CWP1_YEAST (P28319) Cell wall protein CWP1
precursor, partial (6%)
Length = 937
Score = 30.4 bits (67), Expect = 0.28
Identities = 16/38 (42%), Positives = 21/38 (55%)
Frame = +1
Query: 191 GCSGVKRQKGLLGSSQNAAATSHSSNSNGTTPKSFATS 228
G S V K G+ +AAA S S+ NG+ P SF T+
Sbjct: 616 GASSVSHPKSQDGNRVDAAAASCGSDDNGSIPDSFYTN 729
>BP038231
Length = 513
Score = 30.0 bits (66), Expect = 0.36
Identities = 11/20 (55%), Positives = 14/20 (70%)
Frame = +1
Query: 20 QHHHQQNQNQNQNQNQPISH 39
QHHH Q+Q+Q+QN P H
Sbjct: 226 QHHHHQHQHQHQNHTLPHLH 285
>AU251519
Length = 412
Score = 29.6 bits (65), Expect = 0.47
Identities = 19/53 (35%), Positives = 21/53 (38%)
Frame = -3
Query: 21 HHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIFPLLTATPCMPQQQ 73
HHH QNQN H H+ H L S A S G P P P Q+
Sbjct: 293 HHHPQNQN----------HSHHQIHRLASKAGESAGSSPQPREANFPNPP*QE 165
>CN825242
Length = 698
Score = 29.6 bits (65), Expect = 0.47
Identities = 16/42 (38%), Positives = 22/42 (52%)
Frame = +2
Query: 47 LPSSASLSVGFGIFPLLTATPCMPQQQSQNNEVQENPSNNNN 88
L +S+ S + LLT+ P P + SQNN P NNN+
Sbjct: 2 LSNSSFSSHSLSLSSLLTSPP--PMEHSQNNPTSSYPQNNNH 121
>TC9405 homologue to UP|Q84V96 (Q84V96) Aldehyde dehydrogenase 1
precursor , partial (73%)
Length = 1249
Score = 29.3 bits (64), Expect = 0.62
Identities = 15/40 (37%), Positives = 20/40 (49%)
Frame = +1
Query: 21 HHHQQNQNQNQNQNQPISHDHNSNHSLPSSASLSVGFGIF 60
HHH N + + P DH +HS S+ SLS+ F F
Sbjct: 13 HHH----NNKEKKQ*PHRSDHGCSHSFLSAFSLSLSFFFF 120
>TC17842 similar to PIR|B86422|B86422 F1N18.10 protein - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (6%)
Length = 445
Score = 29.3 bits (64), Expect = 0.62
Identities = 16/37 (43%), Positives = 19/37 (51%), Gaps = 10/37 (27%)
Frame = +3
Query: 21 HHHQQNQ-----NQNQNQNQPIS-----HDHNSNHSL 47
HHH QNQ N N N N P S + H +N+SL
Sbjct: 3 HHHMQNQTLIFNNNNNNNNAPSSTNSSMYSHYNNNSL 113
>BP049447
Length = 478
Score = 28.9 bits (63), Expect = 0.81
Identities = 14/33 (42%), Positives = 17/33 (51%), Gaps = 4/33 (12%)
Frame = +1
Query: 14 PSPSSLQ----HHHQQNQNQNQNQNQPISHDHN 42
PS SLQ HHH N Q QN P ++H+
Sbjct: 202 PSQISLQDLHHHHHHHNTTTTQIQNSPTMNNHH 300
>BP055028
Length = 530
Score = 28.5 bits (62), Expect = 1.1
Identities = 15/41 (36%), Positives = 25/41 (60%)
Frame = +2
Query: 6 LRDLVLIAPSPSSLQHHHQQNQNQNQNQNQPISHDHNSNHS 46
LR+LV P+PS + H Q Q++N+ I+ H+SN++
Sbjct: 56 LRNLV---PAPSVKRMHSLQQHQHKQHKNKAITPIHSSNNN 169
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.128 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,283,771
Number of Sequences: 28460
Number of extensions: 90481
Number of successful extensions: 949
Number of sequences better than 10.0: 104
Number of HSP's better than 10.0 without gapping: 869
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 920
length of query: 235
length of database: 4,897,600
effective HSP length: 87
effective length of query: 148
effective length of database: 2,421,580
effective search space: 358393840
effective search space used: 358393840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 54 (25.4 bits)
Medicago: description of AC146853.5


