

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
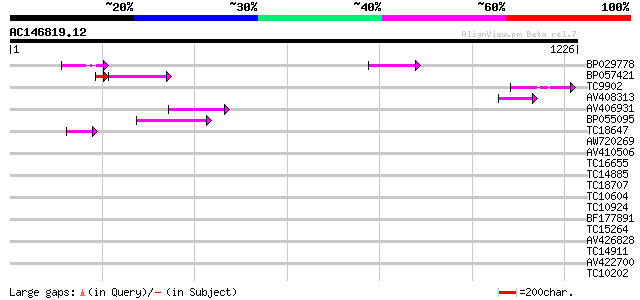
Query= AC146819.12 + phase: 0 /pseudo
(1226 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP029778 78 1e-14
BP057421 55 1e-12
TC9902 similar to UP|BAD07483 (BAD07483) PDR-type ABC transporte... 67 1e-11
AV408313 61 9e-10
AV406931 48 1e-05
BP055095 43 3e-04
TC18647 similar to UP|Q9ASR9 (Q9ASR9) At2g01320/F10A8.20, partia... 42 6e-04
AW720269 35 0.055
AV410506 34 0.16
TC16655 homologue to UP|O99018 (O99018) Chloroplast protease pre... 34 0.16
TC14885 similar to UP|XTH8_ARATH (Q8L9A9) Probable xyloglucan en... 33 0.27
TC18707 homologue to PIR|T02901|T02901 MSP1 protein homolog T13J... 33 0.36
TC10604 similar to AAQ89671 (AAQ89671) At4g33460, partial (36%) 32 0.46
TC10924 homologue to UP|Q8H9D4 (Q8H9D4) 26S proteasome AAA-ATPas... 32 0.46
BF177891 32 0.61
TC15264 homologue to UP|DHSA_ARATH (O82663) Succinate dehydrogen... 32 0.79
AV426828 30 1.8
TC14911 homologue to UP|Q93X55 (Q93X55) Peroxin 6 (Fragment), pa... 30 1.8
AV422700 30 2.3
TC10202 similar to UP|AX28_SOYBN (P13089) Auxin-induced protein ... 29 3.9
>BP029778
Length = 438
Score = 77.8 bits (190), Expect = 1e-14
Identities = 42/111 (37%), Positives = 63/111 (55%)
Frame = -1
Query: 777 EERNDSFF*TTLYHL**SNIFS*YATGNEESTSCRGKVEPIEWCQWFIQTGCSHSSDGCD 836
++ NDS T + + *YAT ++E+ R +VE + C W T C++S +G
Sbjct: 396 KQGNDSSVSTIYGDIPQCQLLC*YATRDKETRHNRNEVETLVKC*WGFCTRCTNSINGVK 217
Query: 837 WCWQNNSDGCTSWKKNQRIYWWDHHNFWLFKEAGNLCKSLWIL*TKLYPFS 887
W W++N +GC+ WKKN RIY + N WL K ++C+S I *TK + FS
Sbjct: 216 WSWKDNFNGCSCWKKNWRIY*G*Y*NIWLSKSTTHICQSSRIC*TK*HTFS 64
Score = 47.8 bits (112), Expect = 1e-05
Identities = 32/102 (31%), Positives = 51/102 (49%), Gaps = 2/102 (1%)
Frame = -2
Query: 113 NYMLDLVEYILKRR--KQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPN 170
NY +D+ + I K+ + +L +L +VSG+ LT L+G +GKT L+ LAG+
Sbjct: 341 NYYVDMPQEIRKQGIIETKLKLLSNVSGVFAPGVLTALMGSSGAGKTTLMDVLAGR---- 174
Query: 171 LKIANEVQFYEQFAGKVSYNGHEMNEFVPQRTAAYVSQNDTH 212
K ++ G + +G+ + R A YV QND H
Sbjct: 173 -KTGGYIE------GDIKISGYPKVQHTFARVAGYVEQNDIH 69
>BP057421
Length = 547
Score = 55.1 bits (131), Expect(2) = 1e-12
Identities = 20/30 (66%), Positives = 29/30 (96%)
Frame = +3
Query: 185 GKVSYNGHEMNEFVPQRTAAYVSQNDTHLG 214
G+++YNGH++NEFVP++TAAY+SQND H+G
Sbjct: 51 GEITYNGHKLNEFVPRKTAAYISQNDVHVG 140
Score = 35.8 bits (81), Expect(2) = 1e-12
Identities = 43/138 (31%), Positives = 65/138 (46%), Gaps = 1/138 (0%)
Frame = +1
Query: 213 LGN*LSEKPWPFQQEFKELDLDMIC*KRCVEERWRKTLFLIRISMSI-*RL*QLRTREQM 271
L *L +KPW Q + K L LDMI * VE + RK ++ +R +++ *RL + ++ +
Sbjct: 136 LEK*L*KKPWISQLDAKGLGLDMIX*VSLVEGK-RKLVYSLRQNLTFS*RLLH*KEQKAV 312
Query: 272 L*QIIF*RFWDWTYVKILW*EMRS*KASLKDKGNVLQ*GRR*SDH*NLYSWMIYLLVWMT 331
I ++W + + EM+ + L K +V R WM Y V +
Sbjct: 313 SLLTILLKYWGLIFARTP*WEMKCTEVYLVAKRSV*PQER*SWGPQRHCLWMRYQRVLIV 492
Query: 332 QQPFKL*NR*NNLSTFSK 349
Q FKL*N + L T K
Sbjct: 493 PQHFKL*NACSRLCTSLK 546
>TC9902 similar to UP|BAD07483 (BAD07483) PDR-type ABC transporter 1, partial
(19%)
Length = 1207
Score = 67.4 bits (163), Expect = 1e-11
Identities = 53/140 (37%), Positives = 71/140 (49%)
Frame = +3
Query: 1083 EKQSSYCRIEHSSS*FCESSFSFKILKTPFRSIQGLLMETTLVLLAESAIQCIKISLHSR 1142
E+Q++Y R + +*F S F +L + GLLMETTLV+LA+S I C ++ LH
Sbjct: 21 EEQAAYTRAG*ACT*FKGSIFRNSVLAAFLDPMPGLLMETTLVILAQSTIHCCEVFLH-- 194
Query: 1143 CLYLFRERVLWPWLQNVHFNQLQ*KETRSP*LHWLNVDYYSTYWH*ECWFCASSGDCRES 1202
F R +W L Q Q KETRS NV S W + + C +SG ++
Sbjct: 195 ---YFHCRDVWNNLLGPR-RQTQ-KETRSVECCGFNV*CCSLPWSAKLFICTASGIS*KN 359
Query: 1203 SFL*GKCCQNVFSTGICFWS 1222
S L* K C NVF +C S
Sbjct: 360 SLL*RKSCWNVFCLTLCICS 419
>AV408313
Length = 342
Score = 61.2 bits (147), Expect = 9e-10
Identities = 34/83 (40%), Positives = 49/83 (58%)
Frame = +2
Query: 1058 NYFFRKRNAIGD*FF*SVQKFRIIQEKQSSYCRIEHSSS*FCESSFSFKILKTPFRSIQG 1117
+Y F R+ *F +QKF +QEKQ++Y R H+ S F S F + IL F + G
Sbjct: 2 SYKFSTRSYYRC*FSSDLQKF*AVQEKQATYSRTGHTCSWFK*SLFPYSILTVIFGPMLG 181
Query: 1118 LLMETTLVLLAESAIQCIKISLH 1140
+LMET LV+LA+ + C ++ LH
Sbjct: 182 MLMETALVILAQPTVHCCEVFLH 250
>AV406931
Length = 438
Score = 47.8 bits (112), Expect = 1e-05
Identities = 48/132 (36%), Positives = 62/132 (46%)
Frame = +1
Query: 343 NLSTFSKELL*SHYNSHH*RLTIFLMTLFYSPMDTLCTKVLVYKCLIFLLQ*VLCVLRGN 402
++ TFS LL H +S H RL F M S M LCT V I VLRG
Sbjct: 25 SMFTFSMALLLYHCSSQHPRLMTFSMISS*SLMAKLCTMVPANMFSISSNLWASNVLRGK 204
Query: 403 QW*TFYKK*HQ*KIRNNIGHIKKSLIYL*QLKSLQMLLSRIMLGKALLMNLLLNLTSLKA 462
TF++K* Q +I+N+ G + + L +L S +L L L +LT L+A
Sbjct: 205 VLLTFFRK*LQRRIKNSTGCAEMNPTDLSLSLNLLRHFSHSILVGNLQRRLQFHLTRLRA 384
Query: 463 IQLL*QQTSMEL 474
I LL* SM L
Sbjct: 385 ILLL*PLKSMVL 420
>BP055095
Length = 538
Score = 43.1 bits (100), Expect = 3e-04
Identities = 54/162 (33%), Positives = 67/162 (41%)
Frame = +1
Query: 275 IIF*RFWDWTYVKILW*EMRS*KASLKDKGNVLQ*GRR*SDH*NLYSWMIYLLVWMTQQP 334
I+ R+ W YV + M+ SL D+ Q R D SWM Y W P
Sbjct: 31 IMLSRYLGWIYVLM**LVMK*EGVSLVDRRRE*QQVRCW*DLQRHSSWMKYQQDWTVPPP 210
Query: 335 FKL*NR*NNLSTFSKELL*SHYNSHH*RLTIFLMTLFYSPMDTLCTKVLVYKCLIFLLQ* 394
+ + *+ * + S H R TL L TKV C FL
Sbjct: 211 SRYASS*DKWFISWM*PW*FLFYSQHQRHLNSSTTLSCFQKVKLYTKVHERTC*SFLSTW 390
Query: 395 VLCVLRGNQW*TFYKK*HQ*KIRNNIGHIKKSLIYL*QLKSL 436
V VL+ + TFYKK*H K RNNIG K SLI + Q +L
Sbjct: 391 VSSVLKEKELLTFYKK*HPRKTRNNIGLEKMSLIDMFQFLNL 516
>TC18647 similar to UP|Q9ASR9 (Q9ASR9) At2g01320/F10A8.20, partial (17%)
Length = 430
Score = 42.0 bits (97), Expect = 6e-04
Identities = 26/68 (38%), Positives = 38/68 (55%), Gaps = 2/68 (2%)
Frame = +1
Query: 124 KRRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKL--DPNLKIANEVQFYE 181
K K +L++VSG K RL ++GP SGKT LL LAG+L P L ++ ++F
Sbjct: 82 KSSKSVRFLLKNVSGEAKPGRLLAIMGPSGSGKTTLLNVLAGQLAASPRLHLSGLLEFNG 261
Query: 182 QFAGKVSY 189
+ K +Y
Sbjct: 262 KPGSKNAY 285
>AW720269
Length = 524
Score = 35.4 bits (80), Expect = 0.055
Identities = 15/45 (33%), Positives = 30/45 (66%)
Frame = +3
Query: 123 LKRRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKL 167
L ++ + +NIL+ VS + + S + ++GP +GK+ LL +AG++
Sbjct: 27 LTQKPKPVNILKSVSFVARSSEIVAVVGPSGTGKSTLLRIIAGRV 161
>AV410506
Length = 433
Score = 33.9 bits (76), Expect = 0.16
Identities = 21/58 (36%), Positives = 33/58 (56%), Gaps = 10/58 (17%)
Frame = +1
Query: 1 MESDEISLMKWDSIQRLPTVARLRRGLL-TTPEGDS---------NEIDVHKIGLQER 48
++ DE +L KW +I++LPT RLR ++ T EGD E+DV K+ + +R
Sbjct: 262 VDEDEEAL-KWAAIEKLPTYDRLRTSIIQTIAEGDQPQGGNRMQHKEVDVTKLDMNDR 432
>TC16655 homologue to UP|O99018 (O99018) Chloroplast protease precursor,
partial (39%)
Length = 934
Score = 33.9 bits (76), Expect = 0.16
Identities = 16/39 (41%), Positives = 23/39 (58%)
Frame = +3
Query: 147 LLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFAG 185
LL+GPP +GKT+L A+AG+ + +F E F G
Sbjct: 765 LLIGPPGTGKTLLAKAIAGEAGVPFFSISGSEFVEMFVG 881
>TC14885 similar to UP|XTH8_ARATH (Q8L9A9) Probable xyloglucan
endotransglucosylase/hydrolase protein 8 precursor
(At-XTH8) (XTH-8) , partial (93%)
Length = 1257
Score = 33.1 bits (74), Expect = 0.27
Identities = 18/62 (29%), Positives = 26/62 (41%), Gaps = 1/62 (1%)
Frame = +3
Query: 804 NEESTSCRGKVEPIEW-CQWFIQTGCSHSSDGCDWCWQNNSDGCTSWKKNQRIYWWDHHN 862
N + + RG +E W F+ + S D C W ++ C S YWWD +N
Sbjct: 672 NADDWATRGGLEKTNWKLAPFVSSYKDFSVDACQW--EDPFPACVSTTTK---YWWDQYN 836
Query: 863 FW 864
W
Sbjct: 837 AW 842
>TC18707 homologue to PIR|T02901|T02901 MSP1 protein homolog T13J8.110 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(29%)
Length = 771
Score = 32.7 bits (73), Expect = 0.36
Identities = 15/29 (51%), Positives = 20/29 (68%)
Frame = +3
Query: 138 GILKHSRLTLLLGPPNSGKTILLLALAGK 166
G+LK R LL GPP +GKT+L A+A +
Sbjct: 288 GLLKPCRGILLFGPPGTGKTMLAXAIANE 374
>TC10604 similar to AAQ89671 (AAQ89671) At4g33460, partial (36%)
Length = 498
Score = 32.3 bits (72), Expect = 0.46
Identities = 17/45 (37%), Positives = 25/45 (54%)
Frame = +2
Query: 125 RRKQQLNILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDP 169
R+ + +L+D S + + +LLGP GK+ LL LAG L P
Sbjct: 179 RQTNDVRVLRDCSLRIPSGQFWMLLGPNGCGKSTLLKILAGLLAP 313
>TC10924 homologue to UP|Q8H9D4 (Q8H9D4) 26S proteasome AAA-ATPase subunit
RPT4a, partial (49%)
Length = 648
Score = 32.3 bits (72), Expect = 0.46
Identities = 14/31 (45%), Positives = 20/31 (64%)
Frame = +2
Query: 140 LKHSRLTLLLGPPNSGKTILLLALAGKLDPN 170
+K + LL GPP +GKT+L A+A +D N
Sbjct: 551 IKPPKGVLLYGPPGTGKTLLARAIASNIDAN 643
>BF177891
Length = 390
Score = 32.0 bits (71), Expect = 0.61
Identities = 21/53 (39%), Positives = 29/53 (54%), Gaps = 4/53 (7%)
Frame = +1
Query: 122 ILKRRKQQL----NILQDVSGILKHSRLTLLLGPPNSGKTILLLALAGKLDPN 170
IL R+ +L N+L+ V GIL L GPP +GKT++ ALA + N
Sbjct: 25 ILPMRRPELFTHGNLLRPVKGIL-------LFGPPGTGKTLMAKALATEAGAN 162
>TC15264 homologue to UP|DHSA_ARATH (O82663) Succinate dehydrogenase
[ubiquinone] flavoprotein subunit, mitochondrial (FP)
(Flavoprotein subunit of complex II) , partial (77%)
Length = 1643
Score = 31.6 bits (70), Expect = 0.79
Identities = 13/37 (35%), Positives = 17/37 (45%)
Frame = +3
Query: 821 QWFIQTGCSHSSDGCDWCWQNNSDGCTSWKKNQRIYW 857
QWF+Q G S C WCW W++ + YW
Sbjct: 192 QWFLQCGGS*IRRDCCWCW---------WRRVESRYW 275
>AV426828
Length = 401
Score = 30.4 bits (67), Expect = 1.8
Identities = 20/58 (34%), Positives = 31/58 (52%), Gaps = 9/58 (15%)
Frame = +1
Query: 119 VEYILKRRKQQLN-------ILQDVSGILKHSRLT--LLLGPPNSGKTILLLALAGKL 167
+ ++ K R Q L+ I+ + + +RL LL GPP +GKT +LA+A KL
Sbjct: 10 IPWVEKYRPQSLDDVAAHRDIVDTIDRLTTENRLPHLLLYGPPGTGKTSTILAVARKL 183
>TC14911 homologue to UP|Q93X55 (Q93X55) Peroxin 6 (Fragment), partial (32%)
Length = 1418
Score = 30.4 bits (67), Expect = 1.8
Identities = 18/54 (33%), Positives = 27/54 (49%)
Frame = +2
Query: 137 SGILKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFAGKVSYN 190
SG+ K S + LL GPP +GKT+L A+A + N + + G+ N
Sbjct: 308 SGLRKRSGV-LLYGPPGTGKTLLAKAVATECSLNFLSVKGPELINMYIGESEKN 466
>AV422700
Length = 472
Score = 30.0 bits (66), Expect = 2.3
Identities = 17/47 (36%), Positives = 25/47 (53%)
Frame = +2
Query: 140 LKHSRLTLLLGPPNSGKTILLLALAGKLDPNLKIANEVQFYEQFAGK 186
LK R LL GPP +GKT L+ A+ + +L I + + AG+
Sbjct: 299 LKWPRGLLLYGPPGTGKTSLVRAVVQECGAHLTIISPHSVHRAHAGE 439
>TC10202 similar to UP|AX28_SOYBN (P13089) Auxin-induced protein AUX28,
partial (60%)
Length = 664
Score = 29.3 bits (64), Expect = 3.9
Identities = 12/32 (37%), Positives = 16/32 (49%)
Frame = -2
Query: 1191 WFCASSGDCRESSFL*GKCCQNVFSTGICFWS 1222
W C +S SF GK + S+G CF+S
Sbjct: 318 WICPTSSSRSRRSFFRGKVKHQIHSSGCCFFS 223
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.352 0.155 0.576
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,809,008
Number of Sequences: 28460
Number of extensions: 439493
Number of successful extensions: 6829
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 5029
Number of HSP's successfully gapped in prelim test: 192
Number of HSP's that attempted gapping in prelim test: 1460
Number of HSP's gapped (non-prelim): 5645
length of query: 1226
length of database: 4,897,600
effective HSP length: 101
effective length of query: 1125
effective length of database: 2,023,140
effective search space: 2276032500
effective search space used: 2276032500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146819.12


