

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146818.6 + phase: 0 /pseudo
(642 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
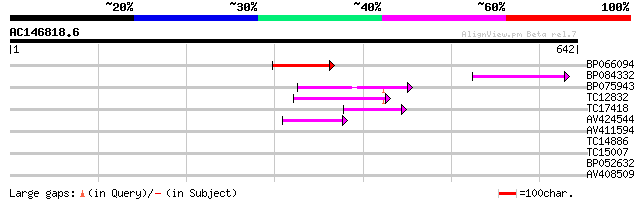
Score E
Sequences producing significant alignments: (bits) Value
BP066094 99 3e-21
BP084332 79 2e-15
BP075943 60 1e-09
TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 54 1e-07
TC17418 similar to UP|Q94AX2 (Q94AX2) AT5g39790/MKM21_80, partia... 41 5e-04
AV424544 40 9e-04
AV411594 30 1.2
TC14886 similar to GB|AAM10290.1|20147147|AY091691 AT3g13930/MDC... 29 2.7
TC15007 similar to UP|Q9FMM3 (Q9FMM3) Gb|AAC80623.1, partial (4%) 28 4.5
BP052632 28 4.5
AV408509 28 5.9
>BP066094
Length = 532
Score = 98.6 bits (244), Expect = 3e-21
Identities = 50/70 (71%), Positives = 57/70 (81%)
Frame = +2
Query: 298 LIKNDFKRGQVDTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMM 357
L++N+ RG+VDTTLF T K IL+VQIYVDDIIFGS N SLCKEFS+ MQ EFE MM
Sbjct: 323 LLENEXVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANPSLCKEFSEMMQAEFEMRMM 502
Query: 358 GELKFFLGIQ 367
GELK+FLGIQ
Sbjct: 503 GELKYFLGIQ 532
>BP084332
Length = 368
Score = 79.3 bits (194), Expect = 2e-15
Identities = 47/109 (43%), Positives = 65/109 (59%)
Frame = +1
Query: 525 NHCYVYSKSIIHFSCKLLYTTTLDETSVGRLSDKC*QYSHLL**YYCYLFVKESNSTFKS 584
NHC +Y + I+ S + ++ LDETS G LS+ QY +LL**+ C ES TFK
Sbjct: 37 NHCTIYCRGRIYLSSNMQHSDALDETSAGGLSNP*EQYPNLL**HCCNFIE*ESYLTFKG 216
Query: 585 QAYINQTPLYQRLCSKRNFKYTIH*Y*TSMG*YIY*AFNYGKI*FYKEK 633
+A+ + LY RLC++ + *Y* SMG Y+Y A + G I*F+ EK
Sbjct: 217 KAH*GKVSLY*RLCTEGRTSSEVC*Y*PSMGRYLYKALSRG*I*FHSEK 363
>BP075943
Length = 547
Score = 59.7 bits (143), Expect = 1e-09
Identities = 38/136 (27%), Positives = 66/136 (47%), Gaps = 5/136 (3%)
Frame = -3
Query: 326 IYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRKDGVYVHQTKYTKE 385
+YVDD++ + + + D+F +GE K+FLG++I + G+ ++Q KY +
Sbjct: 515 LYVDDVLLAGNDMHEIQLVKSSLHDQFRIKDLGEAKYFLGLEIARSTSGIVLNQRKYALQ 336
Query: 386 LLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKL-----YIGMIGSLLYLTASRPDIL 440
L+ + P + + + +GT L Y ++G LLYL +RPDI
Sbjct: 335 LIS----DSGHFGFQPRFYSHGTNSQTLGTNTGTPLTDIGSYRRIVGRLLYLNTTRPDIT 168
Query: 441 FSVCLCARFQSNPKRI 456
F+V ++F S P I
Sbjct: 167 FAVNQLSQFLSAPTDI 120
>TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (9%)
Length = 747
Score = 53.5 bits (127), Expect = 1e-07
Identities = 31/118 (26%), Positives = 63/118 (53%), Gaps = 8/118 (6%)
Frame = +2
Query: 322 LVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQ-RKDG-VYVHQ 379
+++ +YVDD++ N +E + EF+ +G LG+QI++ RKD +++ Q
Sbjct: 89 IILLLYVDDMLVVGPNKDRVQELKAQLAREFDMKDLGPANKILGMQIHRDRKDRRIWLSQ 268
Query: 380 TKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKL------YIGMIGSLLY 431
Y +++L++F ++DC ++TP+ S I + +++ Y +GSL+Y
Sbjct: 269 KNYLQKVLRRFNMQDCNPISTPLPVNYKLSSSMIPSSEAERMEMSRVPYASAVGSLMY 442
>TC17418 similar to UP|Q94AX2 (Q94AX2) AT5g39790/MKM21_80, partial (20%)
Length = 739
Score = 41.2 bits (95), Expect = 5e-04
Identities = 25/72 (34%), Positives = 39/72 (53%), Gaps = 1/72 (1%)
Frame = -1
Query: 379 QTKYTKELLKKFKLEDCKVMNTPM-HPTCT*SKEDIGTKVDQKLYIGMIGSLLYLTASRP 437
Q Y ++LKKFK+ + K ++T + + + G +VD Y +IGS+ YL R
Sbjct: 739 QKIYVDDILKKFKMTNSKYISTTIGGKEIEAGRRNGGKRVDSTYYKSLIGSVRYLNTVRS 560
Query: 438 DILFSVCLCARF 449
DI+ V L +RF
Sbjct: 559 DIVCGVGLRSRF 524
>AV424544
Length = 276
Score = 40.4 bits (93), Expect = 9e-04
Identities = 20/74 (27%), Positives = 38/74 (51%)
Frame = +3
Query: 309 DTTLFRGTLKKYILVVQIYVDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQI 368
D +LF V+ +YVDD+I + + + + +F +G LK+FLG+++
Sbjct: 54 DHSLFTKFRDASFTVILVYVDDLILAGNDLNEIQCVKNKLDIQFRIKDLGTLKYFLGLEV 233
Query: 369 NQRKDGVYVHQTKY 382
+ G+++ Q KY
Sbjct: 234 ARSSCGLFLSQRKY 275
>AV411594
Length = 244
Score = 30.0 bits (66), Expect = 1.2
Identities = 20/65 (30%), Positives = 29/65 (43%)
Frame = +2
Query: 363 FLGIQINQRKDGVYVHQTKYTKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLY 422
FL I + K G+ + + KY +LL + CK P+ P+ + D D LY
Sbjct: 50 FLRT*IAKSKKGILLSRRKYALQLLDDTRNMGCKPPPFPLDPSIKLNSTDGELLSDVTLY 229
Query: 423 IGMIG 427
I IG
Sbjct: 230 IRFIG 244
>TC14886 similar to GB|AAM10290.1|20147147|AY091691 AT3g13930/MDC16_5
{Arabidopsis thaliana;}, partial (47%)
Length = 1352
Score = 28.9 bits (63), Expect = 2.7
Identities = 32/102 (31%), Positives = 46/102 (44%), Gaps = 9/102 (8%)
Frame = +2
Query: 92 LQSKMKVLQIYRSLKTFQNLIRKLNLKKVQKHNPLQNLNEPQKHKMKMLLKKLTMTLSK* 151
LQ +K+ I RSLK + L L L PL + P H+ K L+ L LS+
Sbjct: 650 LQ*LLKMRVILRSLKITKLLHLNLVL-------PLPKKHLPHLHQRKRWLRSLPENLSQR 808
Query: 152 F-NPKVHSSTSLHILRI*SLET--------RIVLEEQDHTSN 184
F NP H + L + LE+ + L+EQD T++
Sbjct: 809 FPNPVRHLHLEIAYLLVLLLESWLKRKIYPSLALKEQDLTAS 934
>TC15007 similar to UP|Q9FMM3 (Q9FMM3) Gb|AAC80623.1, partial (4%)
Length = 626
Score = 28.1 bits (61), Expect = 4.5
Identities = 17/61 (27%), Positives = 34/61 (54%)
Frame = +1
Query: 67 CIIQRYYVLKNQCTLNLMTKSLEVKLQSKMKVLQIYRSLKTFQNLIRKLNLKKVQKHNPL 126
C +Q ++ L + ++ KSL++ LQ+ + +QIYR+ T + +++ K +K L
Sbjct: 115 CYLQMFWTLLLRA--GMIRKSLDLVLQAWYQQMQIYRTGTTLKEPLKERRRVKKEKR*VL 288
Query: 127 Q 127
Q
Sbjct: 289 Q 291
>BP052632
Length = 489
Score = 28.1 bits (61), Expect = 4.5
Identities = 24/73 (32%), Positives = 33/73 (44%), Gaps = 3/73 (4%)
Frame = -1
Query: 75 LKNQCTLNLMTKSLEVKLQSKMKVLQIYRSLK---TFQNLIRKLNLKKVQKHNPLQNLNE 131
LKNQ LNLM +++K L ++R + T+ +K KV H L N
Sbjct: 294 LKNQSMLNLMI--------NQLKPLIVWRKIS*T*TYLIKTKKCQQMKV-NHQKLHMKNR 142
Query: 132 PQKHKMKMLLKKL 144
P HK +L K L
Sbjct: 141 PLNHKQPILGKVL 103
>AV408509
Length = 428
Score = 27.7 bits (60), Expect = 5.9
Identities = 31/110 (28%), Positives = 49/110 (44%), Gaps = 5/110 (4%)
Frame = -3
Query: 328 VDDIIFGSTNASLCKEFSK*MQDEFEKSMMGELKFFLGIQINQRK-----DGVYVHQTKY 382
V D I+ TN L +E K +FE+ +G +K+F G Q+ K + + ++
Sbjct: 324 VIDFIY-MTNDELLEE*KKTTMCQFERLDLGLMKYFDGHQVMAYK*IKNMEKCFFYKASM 148
Query: 383 TKELLKKFKLEDCKVMNTPMHPTCT*SKEDIGTKVDQKLYIGMIGSLLYL 432
+ + F L C+ M +ED KVD LY + SL+YL
Sbjct: 147 LQIYYRSFAL--CR-----MGVNKNLLREDGKEKVDDTLYRKLGQSLIYL 19
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.352 0.155 0.517
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,488,375
Number of Sequences: 28460
Number of extensions: 141115
Number of successful extensions: 1145
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1135
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1141
length of query: 642
length of database: 4,897,600
effective HSP length: 96
effective length of query: 546
effective length of database: 2,165,440
effective search space: 1182330240
effective search space used: 1182330240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146818.6


