

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
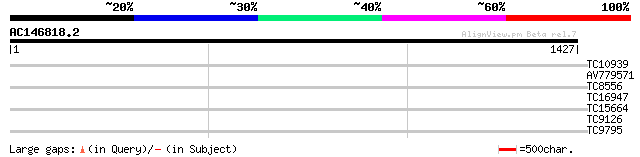
Query= AC146818.2 - phase: 0 /pseudo
(1427 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC10939 similar to UP|Q941N0 (Q941N0) Drought-induced protein, p... 33 0.24
AV779571 31 1.2
TC8556 29 4.6
TC16947 homologue to UP|HIS7_PEA (Q43072) Imidazoleglycerol-phos... 29 4.6
TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, p... 29 6.0
TC9126 29 6.0
TC9795 homologue to UP|Q9ATB8 (Q9ATB8) BiP-isoform D (Fragment),... 28 7.8
>TC10939 similar to UP|Q941N0 (Q941N0) Drought-induced protein, partial (59%)
Length = 564
Score = 33.5 bits (75), Expect = 0.24
Identities = 20/55 (36%), Positives = 34/55 (61%)
Frame = -2
Query: 1171 LMMLA*LVANQFLLLLILLYVYLKILGLFMVMSLVIDV*LEDCYISPPHALI*LL 1225
+M++A LV L+LLILL + + +L LF+ + L++D+ + P LI*L+
Sbjct: 296 VMVMAMLVLLLLLVLLILLVLLVVLLTLFVTVDLILDLVHNPLLVLPSMVLI*LV 132
>AV779571
Length = 538
Score = 31.2 bits (69), Expect = 1.2
Identities = 21/66 (31%), Positives = 40/66 (59%)
Frame = -2
Query: 795 GM*NFLIWNFLSIPLVLLQFPLHIFTLIVKLLTLCLLMTLL*ILLKIMMRFLMKIHLLIL 854
G+ N LI L + L+LL L + +++ LT+ ++ +L*+LLK+ +R + LLI+
Sbjct: 336 GIANLLIIYHLLLSLLLLLLLLLLIVIMILSLTIE*VL-ML*VLLKMRLRLQQLLILLII 160
Query: 855 KKGMMK 860
K +++
Sbjct: 159 LKNLLR 142
>TC8556
Length = 759
Score = 29.3 bits (64), Expect = 4.6
Identities = 18/44 (40%), Positives = 22/44 (49%)
Frame = +2
Query: 1065 MVLSKQVGNGLRSLLSFFMLRDSLKLMQITHYLLKSLLLPTLWF 1108
M L+ Q+G LRS +FML S +L H L L L T F
Sbjct: 500 M*LN*QLGLDLRSFFLYFMLSTSARLQSAVHILDSGLRLRTCLF 631
>TC16947 homologue to UP|HIS7_PEA (Q43072) Imidazoleglycerol-phosphate
dehydratase (IGPD) , partial (31%)
Length = 499
Score = 29.3 bits (64), Expect = 4.6
Identities = 21/58 (36%), Positives = 33/58 (56%), Gaps = 1/58 (1%)
Frame = +2
Query: 800 LIWNFLSIPLVLLQFPLHIFTLIVKLLTLCLL-MTLL*ILLKIMMRFLMKIHLLILKK 856
L W L + +V L+FP I LI L CL+ M L ++ +M+ MK+ LL+L++
Sbjct: 293 LTWMGLELLIVALEFPSLIICLINLLHMDCLMYM*RLLVIYILMIITQMKMSLLLLER 466
>TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, partial (4%)
Length = 670
Score = 28.9 bits (63), Expect = 6.0
Identities = 26/77 (33%), Positives = 35/77 (44%)
Frame = +2
Query: 1336 FMI*SKLLHVYLYSTVITRVHFTYQLIQCFMNAPST*TLIVI*YVKRFKLELCVCYQSPP 1395
F+ S LLH + T I + QL+ CFMN + TLI + + K K + +
Sbjct: 38 FVFLSSLLHCF---TAIVSLLDILQLMLCFMNVLNISTLIAMLFGKSCKPSSFIYFLFQV 208
Query: 1396 NIKQLTCLPRLLVHDNF 1412
IKQ T P L D F
Sbjct: 209 LIKQPTF*PSPLNLDLF 259
>TC9126
Length = 1013
Score = 28.9 bits (63), Expect = 6.0
Identities = 13/32 (40%), Positives = 23/32 (71%)
Frame = +2
Query: 801 IWNFLSIPLVLLQFPLHIFTLIVKLLTLCLLM 832
I +FL+ + L F +++F L+ KLL++CLL+
Sbjct: 611 ITDFLT*VYIRLFFQVYLFQLLEKLLSMCLLL 706
>TC9795 homologue to UP|Q9ATB8 (Q9ATB8) BiP-isoform D (Fragment), partial
(63%)
Length = 981
Score = 28.5 bits (62), Expect = 7.8
Identities = 13/36 (36%), Positives = 26/36 (72%)
Frame = +3
Query: 820 TLIVKLLTLCLLMTLL*ILLKIMMRFLMKIHLLILK 855
T + +L + LLM++ * L+ + +RFL+++ +LIL+
Sbjct: 717 TSLFLILEVVLLMSVS*PLIMVFLRFLLQMEILILE 824
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.370 0.167 0.644
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,701,833
Number of Sequences: 28460
Number of extensions: 460718
Number of successful extensions: 6972
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 3858
Number of HSP's successfully gapped in prelim test: 251
Number of HSP's that attempted gapping in prelim test: 2863
Number of HSP's gapped (non-prelim): 4525
length of query: 1427
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1325
effective length of database: 1,994,680
effective search space: 2642951000
effective search space used: 2642951000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146818.2


