

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146794.13 + phase: 0 /pseudo
(818 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
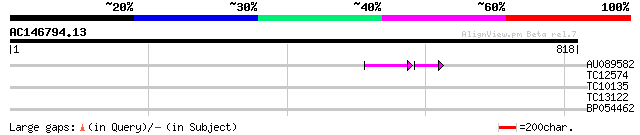
Score E
Sequences producing significant alignments: (bits) Value
AU089582 53 3e-08
TC12574 35 0.048
TC10135 28 4.5
TC13122 similar to UP|Q84Y01 (Q84Y01) Inositol phosphate kinase,... 28 5.8
BP054462 27 10.0
>AU089582
Length = 383
Score = 53.1 bits (126), Expect(2) = 3e-08
Identities = 29/69 (42%), Positives = 40/69 (57%)
Frame = +1
Query: 512 DLYL*YCEAAWCSVEHCVR*RSKVYF*ILEKFARGVGF*VEVEFGLSSANRWSVREDDSV 571
DL * C AWC+ + R RS ++ LE F+ G ++ E+ SS+N WSVRED S
Sbjct: 28 DLLG*DCFLAWCTCVYNFRSRSSIHITFLEVFSNCFGNSIKNEYRFSSSN*WSVREDYSD 207
Query: 572 VRGFVESLC 580
+RG+ LC
Sbjct: 208 LRGYASCLC 234
Score = 22.3 bits (46), Expect(2) = 3e-08
Identities = 16/43 (37%), Positives = 22/43 (50%)
Frame = +2
Query: 584 RWDLG*SSTVDRVHVQQQLPF*YRNGTIRSFVWSEVQNSVMLV 626
R LG D + + L *+ +GTI S VW E+Q + LV
Sbjct: 245 RGXLGSVFVFDGIRI***LSV*HLDGTI*SLVW*EMQVTYRLV 373
>TC12574
Length = 325
Score = 35.0 bits (79), Expect = 0.048
Identities = 14/33 (42%), Positives = 24/33 (72%)
Frame = +2
Query: 19 FRCYMNKIFHAYLDWFVVVFIDDILIYSRSEEE 51
F+ +N IF ++ + F++VFI+DIL Y+ +EE
Sbjct: 2 FKNSVNHIFESFFEHFMIVFINDILSYTEDKEE 100
>TC10135
Length = 1501
Score = 28.5 bits (62), Expect = 4.5
Identities = 22/74 (29%), Positives = 29/74 (38%)
Frame = +1
Query: 653 DESVAEST*ELSRQA*ERPWISGS**CVSKSRSCDRCWSCLEIEEVDAEVYWPVSDIRKS 712
D S+ *+LS Q * W + C+ +C +S L+ YW DI
Sbjct: 943 DSSMGYQL*DLSLQV*RGLWRNCIHHCLDFQLNCFTVFSVLDTN------YWDSMDIHDV 1104
Query: 713 WNSGI*SGFATTSF 726
WN G S F F
Sbjct: 1105 WNFGCCSHFLCHCF 1146
>TC13122 similar to UP|Q84Y01 (Q84Y01) Inositol phosphate kinase, partial
(14%)
Length = 637
Score = 28.1 bits (61), Expect = 5.8
Identities = 15/47 (31%), Positives = 24/47 (50%), Gaps = 1/47 (2%)
Frame = +3
Query: 38 FIDDILIYSRSEEEHAEHLKIMLQVLKEKKMYAKLSK-CEFWLSEVS 83
F D++ + EEE E K KEK+ ++ K C FW++ +S
Sbjct: 141 FFCDVMCQKQQEEEQGEGDKAASPAEKEKEKSVQV*KNCVFWVTRIS 281
>BP054462
Length = 422
Score = 27.3 bits (59), Expect = 10.0
Identities = 15/43 (34%), Positives = 25/43 (57%)
Frame = +3
Query: 697 EVDAEVYWPVSDIRKSWNSGI*SGFATTSFEFA*CVSCVATSE 739
++++ V W +S K W G+ S AT+S A C+SC++ E
Sbjct: 297 KIES*VLWTLSYCSKDWGCGL-ST*ATSSQ*SASCLSCLSPQE 422
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.372 0.167 0.652
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,113,066
Number of Sequences: 28460
Number of extensions: 186411
Number of successful extensions: 2454
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1293
Number of HSP's successfully gapped in prelim test: 58
Number of HSP's that attempted gapping in prelim test: 1125
Number of HSP's gapped (non-prelim): 1409
length of query: 818
length of database: 4,897,600
effective HSP length: 98
effective length of query: 720
effective length of database: 2,108,520
effective search space: 1518134400
effective search space used: 1518134400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146794.13


