

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146792.8 - phase: 0
(266 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
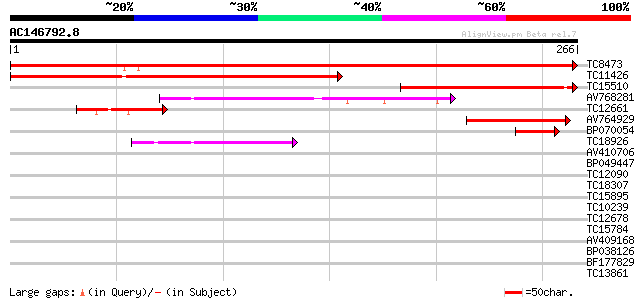
Sequences producing significant alignments: (bits) Value
TC8473 similar to UP|BAD11591 (BAD11591) 27k vesicle-associated ... 475 e-135
TC11426 similar to UP|Q8LAZ8 (Q8LAZ8) Membrane associated protei... 200 2e-52
TC15510 similar to UP|BAD11591 (BAD11591) 27k vesicle-associated... 129 4e-31
AV768281 62 1e-10
TC12661 similar to UP|Q9LS15 (Q9LS15) Gb|AAF26010.1, partial (14%) 54 4e-08
AV764929 47 3e-06
BP070054 47 3e-06
TC18926 similar to AAQ63968 (AAQ63968) VAP27-1, partial (36%) 40 3e-04
AV410706 36 0.008
BP049447 35 0.010
TC12090 similar to GB|AAF05768.1|6180060|AF193027 beta-3b adrene... 35 0.013
TC18307 similar to UP|Q7XYK9 (Q7XYK9) Plastid protein SufE (Frag... 35 0.017
TC15895 34 0.022
TC10239 similar to UP|Q7NEZ8 (Q7NEZ8) Glr3729 protein, partial (8%) 34 0.022
TC12678 weakly similar to UP|Q9LQK7 (Q9LQK7) F5D14.28 protein, p... 34 0.022
TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precurso... 34 0.029
AV409168 34 0.029
BP038126 33 0.038
BF177829 33 0.050
TC13861 weakly similar to UP|Q9S4J5 (Q9S4J5) Serum opacity facto... 27 0.058
>TC8473 similar to UP|BAD11591 (BAD11591) 27k vesicle-associated membrane
protein-associated protein-like, partial (81%)
Length = 1232
Score = 475 bits (1222), Expect = e-135
Identities = 239/269 (88%), Positives = 256/269 (94%), Gaps = 3/269 (1%)
Frame = +1
Query: 1 MAVAEPKLHSEPKVWNFFKLPFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSN--TPLEGS 58
MAVAEPK HSEPKVWN FKLPFR+S+T++ +T ++S ++NLHHHHHHNPN + PLEGS
Sbjct: 142 MAVAEPKPHSEPKVWNLFKLPFRHSSTTANSTGTTSSSSNLHHHHHHNPNLHHPPPLEGS 321
Query: 59 T-SHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQ 117
T SHTS SVSSVARSLLPTRRRLKLDPS+KLYFPYEPGKQVRSAIRIKNTSKS+VAFKFQ
Sbjct: 322 TASHTSGSVSSVARSLLPTRRRLKLDPSSKLYFPYEPGKQVRSAIRIKNTSKSHVAFKFQ 501
Query: 118 TTAPKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKPEKSGLKFKIMSLKVKGSIDYV 177
TTAPKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKPEK+GLKFKIMSLKVKG++DYV
Sbjct: 502 TTAPKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKPEKTGLKFKIMSLKVKGAMDYV 681
Query: 178 PELFDEQKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPKII 237
PELFDEQKDQVAVEQILRV+FLDPERPSP LEKLKRQLADADAALE RKKPAEDAGPKII
Sbjct: 682 PELFDEQKDQVAVEQILRVIFLDPERPSPALEKLKRQLADADAALEARKKPAEDAGPKII 861
Query: 238 GEGLVIDEWKERRERYLAKQQGDVVVDSV 266
GEGLVIDEWKERRERYLAKQQG+VVVDSV
Sbjct: 862 GEGLVIDEWKERRERYLAKQQGEVVVDSV 948
>TC11426 similar to UP|Q8LAZ8 (Q8LAZ8) Membrane associated protein-like
protein, partial (33%)
Length = 608
Score = 200 bits (508), Expect = 2e-52
Identities = 101/156 (64%), Positives = 122/156 (77%)
Frame = +3
Query: 1 MAVAEPKLHSEPKVWNFFKLPFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTS 60
M V K S+ K W+F ++PF +N ++SSS + +H+ HH N + + ++ ST
Sbjct: 126 MEVESEKPGSDVKGWSFCRMPFWQTNNHIPPSSSSSTTSYMHNGHHQNISFQS-VDRSTH 302
Query: 61 HTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTA 120
S +VSSVA+SLLPTRRRL+LDPSNKLYFPYEPGKQVRSAI IKNT K++VAFKFQTTA
Sbjct: 303 QASPTVSSVAKSLLPTRRRLRLDPSNKLYFPYEPGKQVRSAIAIKNTCKTHVAFKFQTTA 482
Query: 121 PKSCFMRPPGAILAPGESIIATVFKFVEQPENNEKP 156
PKSC+MRPPG IL+PGESIIATVFKFVE PENNEKP
Sbjct: 483 PKSCYMRPPGGILSPGESIIATVFKFVEPPENNEKP 590
>TC15510 similar to UP|BAD11591 (BAD11591) 27k vesicle-associated membrane
protein-associated protein-like, partial (33%)
Length = 568
Score = 129 bits (325), Expect = 4e-31
Identities = 63/83 (75%), Positives = 75/83 (89%)
Frame = +3
Query: 184 QKDQVAVEQILRVVFLDPERPSPILEKLKRQLADADAALEERKKPAEDAGPKIIGEGLVI 243
Q+DQVAVEQ+LRV+FLDPERPSP ++KLKRQLA+A+AALE RKKP E+ GP++ GEGLVI
Sbjct: 3 QRDQVAVEQVLRVIFLDPERPSPAMDKLKRQLAEAEAALESRKKPPEETGPRVAGEGLVI 182
Query: 244 DEWKERRERYLAKQQGDVVVDSV 266
DEWKERRERYLA+QQ + VDSV
Sbjct: 183 DEWKERRERYLARQQVE-GVDSV 248
>AV768281
Length = 575
Score = 62.0 bits (149), Expect = 1e-10
Identities = 45/144 (31%), Positives = 76/144 (52%), Gaps = 5/144 (3%)
Frame = +3
Query: 71 RSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRPPG 130
RS + T L +DP +L F +E KQ+ ++++ N + VAFK +TT PK +RP
Sbjct: 105 RSEMSTGDLLHIDPL-ELKFIFELKKQISCSLQLSNQTDEYVAFKVKTTNPKKYCVRPNT 281
Query: 131 AILAPGESIIATVFKFVEQPENNEKPE-KSGLKFKIMSLKVKGSI---DYVPELFDEQKD 186
++ P + TV Q + P+ + KF + S+KV+ + D E+F+++
Sbjct: 282 GVVLPRSTCDVTV---TMQAQKEAPPDMQCKDKFLLQSVKVEDGVTAKDITAEMFNKEAG 452
Query: 187 QVAVEQILRVVFL-DPERPSPILE 209
+ E LRVV++ P+ PSP+ E
Sbjct: 453 HIVEECKLRVVYVAPPQPPSPVAE 524
>TC12661 similar to UP|Q9LS15 (Q9LS15) Gb|AAF26010.1, partial (14%)
Length = 609
Score = 53.5 bits (127), Expect = 4e-08
Identities = 31/48 (64%), Positives = 35/48 (72%), Gaps = 5/48 (10%)
Frame = -1
Query: 32 TTSSSVNN---NLHHHHHHNPNSNTP--LEGSTSHTSNSVSSVARSLL 74
T SS N+ NLHHHH NPN + P EGST+HTS+SVSSVARSLL
Sbjct: 423 TLSSGTNSFPANLHHHHR-NPNLHNPPPFEGSTAHTSSSVSSVARSLL 283
>AV764929
Length = 328
Score = 47.4 bits (111), Expect = 3e-06
Identities = 26/49 (53%), Positives = 31/49 (63%)
Frame = -2
Query: 215 LADADAALEERKKPAEDAGPKIIGEGLVIDEWKERRERYLAKQQGDVVV 263
LADAD A PA GP I G+GLVI +WK+RR R L +QQ +V V
Sbjct: 327 LADADPAARASTAPAAGPGPIIKGQGLVIYKWKKRRYRSLPQQQVEVDV 181
>BP070054
Length = 332
Score = 47.4 bits (111), Expect = 3e-06
Identities = 21/21 (100%), Positives = 21/21 (100%)
Frame = -2
Query: 238 GEGLVIDEWKERRERYLAKQQ 258
GEGLVIDEWKERRERYLAKQQ
Sbjct: 331 GEGLVIDEWKERRERYLAKQQ 269
>TC18926 similar to AAQ63968 (AAQ63968) VAP27-1, partial (36%)
Length = 419
Score = 40.4 bits (93), Expect = 3e-04
Identities = 25/78 (32%), Positives = 42/78 (53%)
Frame = +1
Query: 58 STSHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPYEPGKQVRSAIRIKNTSKSNVAFKFQ 117
S H +S S S + T L++DP +L F ++ KQ+ ++++ N + VAFK +
Sbjct: 91 SARHNPHSFSD-RDSKMSTGYLLRIDPL-ELKFTFQLKKQISCSLQLTNKTDEYVAFKVK 264
Query: 118 TTAPKSCFMRPPGAILAP 135
TT PK +RP ++ P
Sbjct: 265 TTNPKKYCVRPNTGVVLP 318
>AV410706
Length = 427
Score = 35.8 bits (81), Expect = 0.008
Identities = 13/33 (39%), Positives = 24/33 (72%)
Frame = +1
Query: 25 SNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEG 57
S+T++T S ++ ++ HHHHHH P+ ++ L+G
Sbjct: 7 SSTTTTTQPSHTLLHSHHHHHHHPPSKDSCLQG 105
>BP049447
Length = 478
Score = 35.4 bits (80), Expect = 0.010
Identities = 15/40 (37%), Positives = 24/40 (59%), Gaps = 7/40 (17%)
Frame = +1
Query: 26 NTSSTNTTSSSVNN-------NLHHHHHHNPNSNTPLEGS 58
N+ T TT+++ NN +LHHHHHH+ + T ++ S
Sbjct: 160 NSFLTTTTTTTTNNPSQISLQDLHHHHHHHNTTTTQIQNS 279
Score = 27.3 bits (59), Expect = 2.7
Identities = 13/35 (37%), Positives = 19/35 (54%), Gaps = 4/35 (11%)
Frame = +3
Query: 24 NSNTSSTNTTSSSVNNNLHHHHHHN----PNSNTP 54
N N ++ ++ +N L HHHHH+ P SN P
Sbjct: 114 NHNYATL*QRNARINQLLPHHHHHHHHQQPFSNLP 218
>TC12090 similar to GB|AAF05768.1|6180060|AF193027 beta-3b adrenergic
receptor splice variant {Mus musculus;} , partial (6%)
Length = 421
Score = 35.0 bits (79), Expect = 0.013
Identities = 16/40 (40%), Positives = 21/40 (52%), Gaps = 4/40 (10%)
Frame = +3
Query: 40 NLHHHHHHNPNSNTPLEGS----TSHTSNSVSSVARSLLP 75
N HHHHHH+P ++ L S SH S + +A L P
Sbjct: 168 NRHHHHHHHPQTDELLSSSLHAFQSHVSRFIGQLALDLKP 287
>TC18307 similar to UP|Q7XYK9 (Q7XYK9) Plastid protein SufE (Fragment),
partial (9%)
Length = 773
Score = 34.7 bits (78), Expect = 0.017
Identities = 21/79 (26%), Positives = 33/79 (41%)
Frame = +2
Query: 21 PFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSVARSLLPTRRRL 80
P SN S ++ + ++ HHHHHH+P PL+ + S SS R P
Sbjct: 137 PSSPSNPSPSSEFQPNHQHHHHHHHHHHPLPCNPLKSFLRSSKKS*SSSNRCRNPKPSTS 316
Query: 81 KLDPSNKLYFPYEPGKQVR 99
+ ++ P P + R
Sbjct: 317 SSYSTGRISNPSIPSSRAR 373
>TC15895
Length = 622
Score = 34.3 bits (77), Expect = 0.022
Identities = 16/42 (38%), Positives = 23/42 (54%), Gaps = 1/42 (2%)
Frame = -1
Query: 19 KLPFRNSNTSSTNTTSSS-VNNNLHHHHHHNPNSNTPLEGST 59
+ PF +++ S T ++N HHHHHH P S PL S+
Sbjct: 283 RAPFTSASHSPTQNQGLM*PHHN*HHHHHHQPGSLPPLNTSS 158
>TC10239 similar to UP|Q7NEZ8 (Q7NEZ8) Glr3729 protein, partial (8%)
Length = 629
Score = 34.3 bits (77), Expect = 0.022
Identities = 31/139 (22%), Positives = 60/139 (42%), Gaps = 8/139 (5%)
Frame = -2
Query: 99 RSAIRIKNTSKSN-VAFKFQTTAPKSCFMRPPGAILAPGESIIATVFKFVEQ--PENNEK 155
R+ I++++ S++ VAFK QT++P + PP ++ P S V + P++ +
Sbjct: 559 RTTIQLRSLSQTTPVAFKIQTSSPNKFLVNPPSGLIPPLSSATFQVILKPQPHLPQSYPR 380
Query: 156 PEKSGLKFKIMSLKVKGSIDYVPELFD-----EQKDQVAVEQILRVVFLDPERPSPILEK 210
K S+ P+ + KD ++ L+V F+ P + +
Sbjct: 379 SPSDRFLIKTAEFNTDSSVPTHPDSINSWFASRPKDFSTIDLKLKVAFVG---PLLLHDA 209
Query: 211 LKRQLADADAALEERKKPA 229
+ R +A L R++PA
Sbjct: 208 VSRGDLEAVRNLIRRRRPA 152
>TC12678 weakly similar to UP|Q9LQK7 (Q9LQK7) F5D14.28 protein, partial
(10%)
Length = 396
Score = 34.3 bits (77), Expect = 0.022
Identities = 23/81 (28%), Positives = 31/81 (37%)
Frame = +1
Query: 6 PKLHSEPKVWNFFKLPFRNSNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNS 65
P + PK W LP +S+TSS PN ++PL G + TS S
Sbjct: 163 PSFSNSPKPWKTLSLPPLHSSTSSETLP---------------PNPSSPLHGHPARTSPS 297
Query: 66 VSSVARSLLPTRRRLKLDPSN 86
S + S P + P N
Sbjct: 298 ASPTSPSSAPPKSSPSHTPQN 360
>TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precursor (Atperox
P31) (ATP41) , partial (38%)
Length = 475
Score = 33.9 bits (76), Expect = 0.029
Identities = 35/133 (26%), Positives = 52/133 (38%), Gaps = 7/133 (5%)
Frame = +1
Query: 34 SSSVNNNLHHHHHHNPNSNTPLEGS-TSHTSNSVSSVARSLLPTRRRLKLDPSNKLYFPY 92
SS + HHHHHH+ +S+ S SHT +S S+ + + PS P
Sbjct: 13 SSKLQQWHHHHHHHSSSSSQHSHSSPPSHTRDSHSTSTNTRAHNSPKSSATPSPTNKSPP 192
Query: 93 EPGKQVRSAIRIKNTSKSNVAFKFQTTAPKSCFMRP---PGAI---LAPGESIIATVFKF 146
P RSA +S + + T P S P P A +P + +T
Sbjct: 193 PPPAPPRSA-----SSSTTASTPTAATPPSSSPQLPSTKPNATPTSTSPSPATPSTSSSE 357
Query: 147 VEQPENNEKPEKS 159
++ P N+ P S
Sbjct: 358 LKPPSNSLAPTPS 396
>AV409168
Length = 243
Score = 33.9 bits (76), Expect = 0.029
Identities = 18/45 (40%), Positives = 27/45 (60%)
Frame = +1
Query: 25 SNTSSTNTTSSSVNNNLHHHHHHNPNSNTPLEGSTSHTSNSVSSV 69
SN S+ ++ S NNN HHHHH P+ + S+S TSN+ ++
Sbjct: 13 SNQSTMDSKSPRTNNN-HHHHHLLPSPRS--SSSSSSTSNNGGAI 138
>BP038126
Length = 548
Score = 33.5 bits (75), Expect = 0.038
Identities = 27/86 (31%), Positives = 35/86 (40%), Gaps = 21/86 (24%)
Frame = +3
Query: 15 WNFFKLPFRNSNTSS-----TNTTSSSVNNNL----------------HHHHHHNPNSNT 53
+NF + P RN T + T +SSV N + HHHHHH +T
Sbjct: 216 FNFTQAPRRNFPTQTFRNECKETLASSVTNKVIGVGFGL*TLPTRTPSHHHHHHTHPLST 395
Query: 54 PLEGSTSHTSNSVSSVARSLLPTRRR 79
PL T V++V S P R R
Sbjct: 396 PLRTFT----GVVATVCASSKPPRAR 461
>BF177829
Length = 134
Score = 33.1 bits (74), Expect = 0.050
Identities = 11/14 (78%), Positives = 12/14 (85%)
Frame = -2
Query: 42 HHHHHHNPNSNTPL 55
HHHHH NP+SN PL
Sbjct: 58 HHHHHQNPSSNPPL 17
>TC13861 weakly similar to UP|Q9S4J5 (Q9S4J5) Serum opacity factor precursor
(Fragment), partial (9%)
Length = 632
Score = 27.3 bits (59), Expect(2) = 0.058
Identities = 9/16 (56%), Positives = 11/16 (68%)
Frame = -2
Query: 33 TSSSVNNNLHHHHHHN 48
T + + LHHHHHHN
Sbjct: 229 TQLLLQHMLHHHHHHN 182
Score = 24.3 bits (51), Expect(2) = 0.058
Identities = 13/42 (30%), Positives = 23/42 (53%), Gaps = 1/42 (2%)
Frame = -2
Query: 42 HHHHHHNPNSNT-PLEGSTSHTSNSVSSVARSLLPTRRRLKL 82
HHHHH+N ++ +E SH ++ + + +R+LKL
Sbjct: 199 HHHHHNNKRRHS*TIEA*ASH----FHALLENKVWVKRKLKL 86
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.130 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,681,331
Number of Sequences: 28460
Number of extensions: 75587
Number of successful extensions: 2558
Number of sequences better than 10.0: 279
Number of HSP's better than 10.0 without gapping: 1542
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2112
length of query: 266
length of database: 4,897,600
effective HSP length: 89
effective length of query: 177
effective length of database: 2,364,660
effective search space: 418544820
effective search space used: 418544820
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 54 (25.4 bits)
Medicago: description of AC146792.8


