

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146792.3 + phase: 0
(518 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
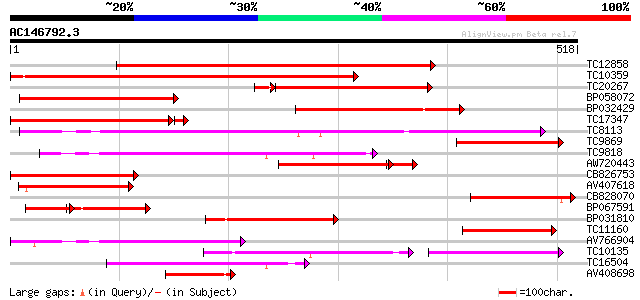
Score E
Sequences producing significant alignments: (bits) Value
TC12858 weakly similar to UP|Q9ZS76 (Q9ZS76) Hexose transporter,... 384 e-107
TC10359 similar to UP|Q94AZ2 (Q94AZ2) AT5g26340/F9D12_17, partia... 377 e-105
TC20267 similar to UP|Q7XA52 (Q7XA52) Monosaccharide transporter... 258 2e-72
BP058072 226 9e-60
BP032429 206 1e-53
TC17347 similar to UP|Q9FMX3 (Q9FMX3) Monosaccharide transporter... 191 4e-51
TC8113 weakly similar to UP|O23213 (O23213) Sugar transporter li... 196 1e-50
TC9869 similar to UP|O65413 (O65413) Glucose transporter (STP12 ... 142 1e-34
TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, ... 142 2e-34
AW720443 127 7e-33
CB826753 132 1e-31
AV407618 121 3e-28
CB828070 115 2e-26
BP067591 69 2e-26
BP031810 112 1e-25
TC11160 similar to UP|Q7XA51 (Q7XA51) Monosaccharide transporter... 110 6e-25
AV766904 97 5e-21
TC10135 93 9e-20
TC16504 weakly similar to UP|Q84KI7 (Q84KI7) Sorbitol transporte... 89 1e-18
AV408698 80 1e-15
>TC12858 weakly similar to UP|Q9ZS76 (Q9ZS76) Hexose transporter, partial
(51%)
Length = 882
Score = 384 bits (986), Expect = e-107
Identities = 187/292 (64%), Positives = 228/292 (78%)
Frame = +3
Query: 98 LVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLY 157
L ASTITR G + +M+ GGL F+ GAL NG A+ +WMLIVGR+LLGFGIG ANQ+VP+Y
Sbjct: 6 LGASTITRIMGMRATMIVGGLFFVSGALFNGLADGIWMLIVGRLLLGFGIGCANQSVPIY 185
Query: 158 LSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITI 217
LSEMAPYKYRG LN+ FQLSITIGI VAN+ NY+FAKI G GWRLSLG VPA+I +
Sbjct: 186 LSEMAPYKYRGGLNMCFQLSITIGIFVANLFNYYFAKILNGQGWRLSLGLGAVPAVIFVV 365
Query: 218 GSLVLPDTPNSMIERGDRDGAKAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQ 277
GSL LPD+P+S++ RG + A+ +L +IRG D++ E D++ ASEA V++PW+ LL+
Sbjct: 366 GSLCLPDSPSSLVARGRHEAARQELVKIRGTTDIEAELKDIITASEALENVKHPWKTLLE 545
Query: 278 RKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLFNSIGFKDDASLMSAVITGVVNVVATCV 337
RKYRPQL AV IPFFQQFTG+NVI FYAP+LF +IGF ASLMSAVI G V+T +
Sbjct: 546 RKYRPQLVFAVCIPFFQQFTGLNVITFYAPILFRTIGFGPTASLMSAVIIGSFKPVSTLI 725
Query: 338 SIYGVDKWGRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIV 389
SI+ VDK+GRR LFLEGGAQMLICQ+ + AI FGTSGNPG LP+WYA+V
Sbjct: 726 SIFVVDKFGRRTLFLEGGAQMLICQIIMTIAIAVTFGTSGNPGQLPKWYAVV 881
>TC10359 similar to UP|Q94AZ2 (Q94AZ2) AT5g26340/F9D12_17, partial (90%)
Length = 1099
Score = 377 bits (969), Expect = e-105
Identities = 187/319 (58%), Positives = 248/319 (77%), Gaps = 1/319 (0%)
Frame = +2
Query: 1 MAGGGIPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPA 60
MAGGG GG ++ +TP V I+CI+AA GGL+FGYD+G+SGGVTSM FL KFFP
Sbjct: 146 MAGGGFATSGGG-DFEAKITPIVIISCILAATGGLMFGYDVGVSGGVTSMPHFLHKFFPD 322
Query: 61 VYRKKNKDKSTNQ-YCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLL 119
VY+K +K + YC+YD+Q L +FTSSLYLA L+S+ AS TR GR+++ML G+
Sbjct: 323 VYKKTVLEKELDSNYCKYDNQGLQLFTSSLYLAGLVSTFFASYTTRTLGRRMTMLIAGVF 502
Query: 120 FLVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSIT 179
F+ G + N A ++ MLIVGRILLG G+GFANQAVPL+LSE+AP + RGALNI FQL+IT
Sbjct: 503 FIAGVVFNVTAQNLAMLIVGRILLGCGVGFANQAVPLFLSEIAPSRIRGALNILFQLNIT 682
Query: 180 IGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAK 239
IGIL AN++NY KIKGGWGWRLSLG A +PA+++T+G+L++ DTPNS+IERG + K
Sbjct: 683 IGILFANLINYGTNKIKGGWGWRLSLGLAGIPAVLLTVGALLVVDTPNSLIERGRTEEGK 862
Query: 240 AQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGI 299
A LK+IRG ++V+ EF +LV AS + +V++P+RNLL+R+ RPQ+ +++ + FQQFTGI
Sbjct: 863 AVLKKIRGTDNVEPEFLELVEASRVAKEVKDPFRNLLKRRNRPQIIISIALQIFQQFTGI 1042
Query: 300 NVIMFYAPVLFNSIGFKDD 318
N IMFYAPVLFN++GFK+D
Sbjct: 1043NAIMFYAPVLFNTLGFKND 1099
>TC20267 similar to UP|Q7XA52 (Q7XA52) Monosaccharide transporter, partial
(31%)
Length = 525
Score = 258 bits (658), Expect(2) = 2e-72
Identities = 123/143 (86%), Positives = 133/143 (92%)
Frame = +2
Query: 244 RIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGINVIM 303
R+RG++DVDEEF+DLV ASEASMQVE+PWRNLLQRKYRP LTMA+LIPFFQQFTGINVIM
Sbjct: 95 RVRGVDDVDEEFSDLVEASEASMQVEHPWRNLLQRKYRPHLTMAILIPFFQQFTGINVIM 274
Query: 304 FYAPVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLICQV 363
FYAPVLF+SIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRR LFL+GG QM+ICQ
Sbjct: 275 FYAPVLFSSIGFKDDASLMSAVITGVVNVVATCVSIYGVDKWGRRKLFLQGGVQMMICQA 454
Query: 364 AVAAAIGAKFGTSGNPGNLPEWY 386
VAAAIGAKFG NPG+LP WY
Sbjct: 455 VVAAAIGAKFGVDWNPGDLPNWY 523
Score = 32.3 bits (72), Expect(2) = 2e-72
Identities = 16/20 (80%), Positives = 17/20 (85%)
Frame = +3
Query: 224 DTPNSMIERGDRDGAKAQLK 243
DTPNSMIERGDR G +A LK
Sbjct: 39 DTPNSMIERGDRSG-QAHLK 95
>BP058072
Length = 520
Score = 226 bits (575), Expect = 9e-60
Identities = 114/145 (78%), Positives = 124/145 (84%)
Frame = -1
Query: 10 GGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDK 69
GG KEY G LT FVT+TCIVAAMGGLIFGYDIGISGGVTSMDPFL+KFF +VY KK +
Sbjct: 508 GGGKEYTGYLTRFVTVTCIVAAMGGLIFGYDIGISGGVTSMDPFLRKFFHSVYLKKQEKS 329
Query: 70 STNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGF 129
S N+YCQYDSQ LT+FTSSLYLAALLSSLVASTITRRF KLSM GG+ F+V AL+NG
Sbjct: 328 SPNKYCQYDSQVLTIFTSSLYLAALLSSLVASTITRRFSPKLSMFVGGMHFVVVALLNGL 149
Query: 130 ANHVWMLIVGRILLGFGIGFANQAV 154
A VWMLIVGRILLG GIGFANQA+
Sbjct: 148 AQRVWMLIVGRILLGIGIGFANQAL 74
>BP032429
Length = 468
Score = 206 bits (523), Expect = 1e-53
Identities = 101/154 (65%), Positives = 121/154 (77%)
Frame = -2
Query: 262 SEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLFNSIGFKDDASL 321
SEAS +VE+PW+N+ + Y+PQ+ LIPFFQQ TGINVIMFYAPVLF ++GF DDA+L
Sbjct: 467 SEASKKVEHPWKNITKPVYKPQMVFCSLIPFFQQLTGINVIMFYAPVLFKTLGFGDDAAL 288
Query: 322 MSAVITGVVNVVATCVSIYGVDKWGRRALFLEGGAQMLICQVAVAAAIGAKFGTSGNPGN 381
MSAVI+G VNV+AT VSI+ VDK+GRR LFLEGG QMLI Q+AV I KFG SG G+
Sbjct: 287 MSAVISGGVNVLATFVSIFSVDKFGRRLLFLEGGVQMLISQIAVGTMIALKFGVSGE-GS 111
Query: 382 LPEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSE 415
+ A +++ FIC YVA FAWSWGPLGWLVPSE
Sbjct: 110 FTKGEANLLLFFICAYVAAFAWSWGPLGWLVPSE 9
>TC17347 similar to UP|Q9FMX3 (Q9FMX3) Monosaccharide transporter (STP11
protein), partial (30%)
Length = 663
Score = 191 bits (485), Expect(2) = 4e-51
Identities = 92/150 (61%), Positives = 119/150 (79%), Gaps = 1/150 (0%)
Frame = +3
Query: 1 MAGGGIPIGGGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPA 60
MAGG G KE+ G +T FV +TC VAAMGGL+FGYD+GI+GGVTSM+PFL KFFP
Sbjct: 168 MAGGAYVDSGKAKEFDGKVTAFVLVTCFVAAMGGLLFGYDLGITGGVTSMEPFLVKFFPG 347
Query: 61 VYRK-KNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLL 119
VY++ K++ ++YC++D++ LT+FTSSLYLAAL++S AST TR GRK SM GGL
Sbjct: 348 VYKQMKDESGHESEYCKFDNELLTLFTSSLYLAALIASFFASTTTRMLGRKTSMFAGGLF 527
Query: 120 FLVGALINGFANHVWMLIVGRILLGFGIGF 149
FLVGAL+NGFA ++ MLI+GR+LLGFG+G+
Sbjct: 528 FLVGALLNGFAVNIEMLIIGRLLLGFGVGY 617
Score = 27.7 bits (60), Expect(2) = 4e-51
Identities = 11/13 (84%), Positives = 13/13 (99%)
Frame = +2
Query: 151 NQAVPLYLSEMAP 163
NQ+VP+YLSEMAP
Sbjct: 620 NQSVPVYLSEMAP 658
>TC8113 weakly similar to UP|O23213 (O23213) Sugar transporter like
protein, partial (93%)
Length = 1865
Score = 196 bits (497), Expect = 1e-50
Identities = 139/500 (27%), Positives = 229/500 (45%), Gaps = 20/500 (4%)
Frame = +2
Query: 10 GGNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDK 69
GGN++ + + IVA+M +I GYD G+ G ++ K++
Sbjct: 77 GGNEDKDTTINKYALACVIVASMVSIISGYDTGVMSGAL------------LFIKEDIGI 220
Query: 70 STNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGF 129
S Q + L + AL+ L A + GR+ ++ +LFLVGA+ G+
Sbjct: 221 SDTQQ--------EVLAGILNICALVGCLAAGKTSDYIGRRYTIFLASILFLVGAVFMGY 376
Query: 130 ANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLN 189
+ +L+ GR + G G+GFA P+Y +E++ RG L ++ I +GI + + N
Sbjct: 377 GPNFAILMFGRCVCGLGVGFALTTAPVYSAELSSASTRGFLTSLPEVCIGLGIFIGYISN 556
Query: 190 YFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAKAQLKRIRG-I 248
YF K+ GWRL LG A +P+L + +G L +P++P ++ +G AK L +
Sbjct: 557 YFLGKLALTLGWRLMLGLAAIPSLGLALGILTMPESPRWLVMQGRLGCAKKVLLEVSNTT 736
Query: 249 EDVDEEFNDLVAAS--------------EASMQVENPWRNLLQRKYRP---QLTMAVLIP 291
E+ + F D++ ++ + S E W+ L R P L AV I
Sbjct: 737 EEAELRFRDIIVSAGFDEKCNDEFVKQPQKSHHGEGVWKELFLRPTPPVRRMLIAAVGIH 916
Query: 292 FFQQFTGINVIMFYAPVLFNSIGF-KDDASLMSAVITGVVNVVATCVSIYGVDKWGRRAL 350
FF+ TGI +M Y P +F G D L++ + TG+ + +S + +D+ GRR L
Sbjct: 917 FFEHATGIEAVMLYGPRIFKKAGVTTKDRLLLATIGTGLTKITFLTISTFLLDRVGRRRL 1096
Query: 351 FLEGGAQMLICQVAVAAAIGAKFGTSGNPGNLPEWYAIVVVLFICIYVAGFAWSWGPLGW 410
A M+ +G W + ++ YVA F P+ W
Sbjct: 1097LQISVAGMIF----GLTILGFSLTMVEYSSEKLVWALSLSIVATYTYVAFFNVGLAPVTW 1264
Query: 411 LVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKF-GLFLFFAFFVLVMSIY 469
+ SEIFPL +R+ S+ V+VN ++ F+ + + G F FA +V ++
Sbjct: 1265VYSSEIFPLRLRAQGNSIGVAVNRGMNAAISMSFISIYKAITIGGAFFLFAGMSVVAWVF 1444
Query: 470 VFFLLPETKGIPIEEMDRVW 489
+F LPETKG +EEM+ V+
Sbjct: 1445FYFCLPETKGKALEEMEMVF 1504
>TC9869 similar to UP|O65413 (O65413) Glucose transporter (STP12 protein),
partial (18%)
Length = 544
Score = 142 bits (358), Expect = 1e-34
Identities = 64/98 (65%), Positives = 81/98 (82%)
Frame = +1
Query: 409 GWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSI 468
GWLVPSEI LE+RSA Q++NV+VNMLFTF +AQ+FL MLCH+KFGLF FFA FV++M+I
Sbjct: 13 GWLVPSEICSLEVRSAGQALNVAVNMLFTFAIAQIFLSMLCHLKFGLFFFFAGFVVIMTI 192
Query: 469 YVFFLLPETKGIPIEEMDRVWKSHPFWSRFVEHGDHGN 506
+V LPETKG+PIEEM+ VW+SH FWS+ V D+ +
Sbjct: 193 FVALFLPETKGVPIEEMNSVWRSHWFWSKIVPAADNND 306
>TC9818 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(62%)
Length = 1162
Score = 142 bits (357), Expect = 2e-34
Identities = 92/326 (28%), Positives = 158/326 (48%), Gaps = 17/326 (5%)
Frame = +2
Query: 28 IVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKKNKDKSTNQYCQYDSQTLTMFTS 87
++A+M ++ GYDIG+ G A+Y K++ S + + +
Sbjct: 251 MLASMTSILLGYDIGVMSGA------------AIYIKRDLKVSDVK--------IEILLG 370
Query: 88 SLYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGRILLGFGI 147
+ L +L+ S +A + GR+ +++F G +F VGAL+ GF+ + W L+ GR + G GI
Sbjct: 371 IINLYSLIGSGLAGRTSDWIGRRYTIVFAGAIFFVGALLMGFSPNYWFLMFGRFIAGIGI 550
Query: 148 GFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWGWRLSLGG 207
G+A P+Y +E++P RG L ++ I GIL+ + N+ F+K+ GWR+ LG
Sbjct: 551 GYALMIAPVYTAEVSPASSRGFLTSFPEVFINGGILLGYISNFAFSKLSLKVGWRMMLGV 730
Query: 208 AMVPALIITIGSLVLPDTPNSMIERG--------------DRDGAKAQLKRIRGIEDVDE 253
+P++I+ +G L +P++P ++ RG + A+ +L I+ + E
Sbjct: 731 GALPSVILGVGVLAMPESPRWLVMRGRLGDAIKVLNKTSDSPEEAQLRLADIKRAAGIPE 910
Query: 254 EFNDLVAASEASMQVENPWRNLL---QRKYRPQLTMAVLIPFFQQFTGINVIMFYAPVLF 310
D V E W+ L R + A+ I FFQQ +GI+ ++ Y+P +F
Sbjct: 911 SCTDDVVEVSKRSTGEGVWKELFLYPTPAIRHIVIAALGIHFFQQASGIDAVVLYSPTIF 1090
Query: 311 NSIGFKDDASLMSAVITGVVNVVATC 336
G K D + A + V V TC
Sbjct: 1091EKAGIKSDTDKLLATV--AVGFVKTC 1162
>AW720443
Length = 406
Score = 127 bits (320), Expect(2) = 7e-33
Identities = 63/106 (59%), Positives = 83/106 (77%), Gaps = 1/106 (0%)
Frame = +2
Query: 246 RGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTMAVLIPFFQQFTGINVIMFY 305
RG ++++ EF+DLV A+ + QV++P+RN+L+RK RPQL +A+ + FQQFTGIN IMFY
Sbjct: 2 RGTDNIELEFSDLVEATRVAKQVKHPFRNILKRKNRPQLVIAIALQIFQQFTGINAIMFY 181
Query: 306 APVLFNSIGFKDDASLMSAVITGVVNVVATCVSIYGVDK-WGRRAL 350
APVLFN+ GFK+DA+L SAVI G VN V+T VSIY VD+ W AL
Sbjct: 182 APVLFNTFGFKNDAALYSAVIVGAVNCVSTIVSIYSVDRLWT*NAL 319
Score = 30.0 bits (66), Expect(2) = 7e-33
Identities = 13/28 (46%), Positives = 18/28 (63%)
Frame = +3
Query: 345 WGRRALFLEGGAQMLICQVAVAAAIGAK 372
+GRR L LE G QML+ + +A +G K
Sbjct: 300 FGRRMLLLEAGFQMLLSLMVIAVVLGIK 383
>CB826753
Length = 494
Score = 132 bits (332), Expect = 1e-31
Identities = 65/119 (54%), Positives = 87/119 (72%), Gaps = 2/119 (1%)
Frame = +3
Query: 1 MAGGGIPIGG-GNKEYPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFP 59
MAGG + G G +++ +TP + I+CI+AA GGL+FGYD+G+SGGVTSM FLKKFFP
Sbjct: 138 MAGGCLTSGSDGGRDFEAKITPIIIISCIMAATGGLMFGYDVGVSGGVTSMPHFLKKFFP 317
Query: 60 AVYRK-KNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFGG 117
V+RK K +++ + YC+YD+Q L +FTSSLYLA L + AS TRR GR+L+ML G
Sbjct: 318 EVFRKTKEEEEFNSNYCKYDNQMLQLFTSSLYLAGLTVTFFASYTTRRLGRRLTMLVSG 494
>AV407618
Length = 334
Score = 121 bits (303), Expect = 3e-28
Identities = 58/108 (53%), Positives = 73/108 (66%), Gaps = 3/108 (2%)
Frame = +2
Query: 9 GGGNKE---YPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFPAVYRKK 65
GG K Y TP+ TC V A+GG +FGYD+G+SGGVTSMD FLK+FFP VYR+K
Sbjct: 11 GGPGKRAHLYEHRFTPYFAFTCFVGALGGSLFGYDLGVSGGVTSMDDFLKEFFPKVYRRK 190
Query: 66 NKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSM 113
+ YC+YD Q LT+FTSSLY++AL + AS +TR GRK S+
Sbjct: 191 HTHLKETDYCKYDDQILTLFTSSLYISALFMTFFASYLTRNKGRKASI 334
>CB828070
Length = 530
Score = 115 bits (288), Expect = 2e-26
Identities = 56/99 (56%), Positives = 73/99 (73%), Gaps = 3/99 (3%)
Frame = +1
Query: 422 RSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFLLPETKGIP 481
RSA QS+ V VN+ FTFL+AQ FL M+C +KFG+F+FF+ + L+MS++VFFLLPETK +P
Sbjct: 1 RSAGQSLTVCVNLAFTFLIAQAFLSMMCSLKFGIFIFFSGWALIMSVFVFFLLPETKNVP 180
Query: 482 IEEM-DRVWKSHPFWSRFVEHG--DHGNGVEMGKGAPKN 517
IEEM +RVWK H W RFVE+ + V GKG +N
Sbjct: 181 IEEMVERVWKQHWLWKRFVENDYVEDDQKVADGKGPTRN 297
>BP067591
Length = 358
Score = 68.6 bits (166), Expect(2) = 2e-26
Identities = 36/76 (47%), Positives = 47/76 (61%)
Frame = +3
Query: 53 FLKKFFPAVYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLS 112
FLK ++ RK N YC YDSQ LT+FTSSLYLA L++SL+AS + GRK +
Sbjct: 126 FLKNSSRSILRKA-AGSDVNMYCVYDSQVLTLFTSSLYLAGLVTSLLASRVRAWIGRKNT 302
Query: 113 MLFGGLLFLVGALING 128
++ GG FL G + G
Sbjct: 303 IILGGATFLGGGAVKG 350
Score = 67.4 bits (163), Expect(2) = 2e-26
Identities = 30/45 (66%), Positives = 37/45 (81%)
Frame = +2
Query: 15 YPGNLTPFVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKKFFP 59
+ +T + ITCIVAA GGLIFGYD+GI+GGVT+M PFL+KFFP
Sbjct: 11 FDSKITLAILITCIVAASGGLIFGYDVGIAGGVTTMLPFLEKFFP 145
>BP031810
Length = 393
Score = 112 bits (280), Expect = 1e-25
Identities = 57/122 (46%), Positives = 85/122 (68%), Gaps = 1/122 (0%)
Frame = +3
Query: 180 IGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDGAK 239
+G AN++NY ++ WGWRLSLG A VPA ++ IG L+ P+TPNS++E+G + A+
Sbjct: 9 LGNFDANLVNYATEQLHP-WGWRLSLGLATVPATVMFIGGLLCPETPNSLVEQGRLEEAR 185
Query: 240 AQLKRIRGIEDVDEEFNDLVAASEASMQVENPWRNLLQRKYRPQLTM-AVLIPFFQQFTG 298
L+R+RG +VD E+ D+V AS + +++NP++NLL +K RPQ + A+ IP FQQ TG
Sbjct: 186 QVLERVRGTPNVDAEYEDIVEASVEAQKIKNPFQNLLLKKNRPQFVIGALAIPAFQQLTG 365
Query: 299 IN 300
N
Sbjct: 366 NN 371
>TC11160 similar to UP|Q7XA51 (Q7XA51) Monosaccharide transporter, partial
(16%)
Length = 538
Score = 110 bits (275), Expect = 6e-25
Identities = 51/86 (59%), Positives = 70/86 (81%)
Frame = +1
Query: 414 SEIFPLEIRSAAQSVNVSVNMLFTFLVAQVFLIMLCHMKFGLFLFFAFFVLVMSIYVFFL 473
SE+FPLEIRSAAQS+ V VNM FT LVAQ+FL+ LC++K+G+FL F ++VMS ++FFL
Sbjct: 1 SELFPLEIRSAAQSIVVCVNMTFTALVAQLFLMSLCNLKYGIFLLFGGLIVVMSCFIFFL 180
Query: 474 LPETKGIPIEEMDRVWKSHPFWSRFV 499
LPETK +PIEE+ ++++H FW + V
Sbjct: 181 LPETKQVPIEEIYLLFENHWFWKKIV 258
>AV766904
Length = 611
Score = 97.4 bits (241), Expect = 5e-21
Identities = 62/219 (28%), Positives = 107/219 (48%), Gaps = 4/219 (1%)
Frame = +3
Query: 1 MAGGGIPIGGGNKEYPGNLTP----FVTITCIVAAMGGLIFGYDIGISGGVTSMDPFLKK 56
MA G G +++ P F I+A+M ++ GYDIG+ G
Sbjct: 12 MAEGEAAAGKSLRDFDPQKKPKRNKFALACAILASMTSILLGYDIGVMSGAV-------- 167
Query: 57 FFPAVYRKKNKDKSTNQYCQYDSQTLTMFTSSLYLAALLSSLVASTITRRFGRKLSMLFG 116
+Y K++ S Q + + + + + + + S +A + GR+ +++
Sbjct: 168 ----IYIKRDLKISDVQ--------IEILSGIMNIYSPIGSYIAGRTSDWIGRRYTIVLA 311
Query: 117 GLLFLVGALINGFANHVWMLIVGRILLGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQL 176
GL+F VGA++ GF+ + L+ GR + G GIGFA P+Y SE++P RG L ++
Sbjct: 312 GLIFFVGAILMGFSPNYAFLMFGRFVAGVGIGFAFLIAPVYTSEISPTLSRGFLTSLPEV 491
Query: 177 SITIGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALII 215
+ GIL+ + NY F+K+ GWRL LG +P++ +
Sbjct: 492 FLNAGILIGYISNYTFSKLPLRLGWRLMLGIGAIPSIFL 608
>TC10135
Length = 1501
Score = 93.2 bits (230), Expect = 9e-20
Identities = 62/198 (31%), Positives = 102/198 (51%), Gaps = 6/198 (3%)
Frame = +2
Query: 178 ITIGILVANVLNYFFAKIKGGWGWRLSLGGAMVPALIITIGSLVLPDTPNSMIERGDRDG 237
IT G ++ ++N F G W W L G A PA+I + L LP++P + +G +
Sbjct: 32 ITGGQFLSYLINLAFTNAPGTWRWML--GVAAAPAIIQIVLMLSLPESPRWLYRKGREEE 205
Query: 238 AKAQLKRIRGIEDVDEEFNDLVAASEASMQVENPWR-----NLLQRKYRPQLTMAVLIPF 292
+K+ LK+I EDVD E L + E+ ++ + L R L + + F
Sbjct: 206 SKSILKKIYAPEDVDAEIEALKESVESEIEESKTSKISMMTLLKTTTVRRGLYAGMGLQF 385
Query: 293 FQQFTGINVIMFYAPVLFNSIGFKDD-ASLMSAVITGVVNVVATCVSIYGVDKWGRRALF 351
FQQF GIN +M+Y+P + GF + +L+ ++IT +N + +SIY +DK GR+ L
Sbjct: 386 FQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSGLNAFGSILSIYFIDKTGRKKL- 562
Query: 352 LEGGAQMLICQVAVAAAI 369
A + +C V ++ A+
Sbjct: 563 ----ALISLCGVVLSLAL 604
Score = 58.5 bits (140), Expect = 3e-09
Identities = 29/125 (23%), Positives = 61/125 (48%), Gaps = 1/125 (0%)
Frame = +2
Query: 383 PEWYAIVVVLFICIYVAGFAWSWGPLGWLVPSEIFPLEIRSAAQSVNVSVNMLFTFLVAQ 442
P + ++ + +Y+ F+ G + W++ SEI+P R + + + +V+Q
Sbjct: 872 PSKFGWAALVGLALYIITFSPGMGTVPWVINSEIYPFRYRGVCGGIASTTVWISNLIVSQ 1051
Query: 443 VFLIMLCHMKFG-LFLFFAFFVLVMSIYVFFLLPETKGIPIEEMDRVWKSHPFWSRFVEH 501
FL + + F+ F +V +V +PETKG+P+EE++++ + +F +
Sbjct: 1052SFLSLTQTIGTAWTFMMFGILAVVAIFFVIVFVPETKGVPMEEVEKMLEQRSLQFKFWQK 1231
Query: 502 GDHGN 506
D G+
Sbjct: 1232KDSGS 1246
>TC16504 weakly similar to UP|Q84KI7 (Q84KI7) Sorbitol transporter, partial
(37%)
Length = 903
Score = 89.4 bits (220), Expect = 1e-18
Identities = 56/200 (28%), Positives = 103/200 (51%), Gaps = 14/200 (7%)
Frame = +2
Query: 89 LYLAALLSSLVASTITRRFGRKLSMLFGGLLFLVGALINGFANHVWMLIVGRILLGFGIG 148
L+++AL + + A I GR+ +++ ++FLVG+++ G+ +L++G + G G+G
Sbjct: 224 LHMSALPACMTAGRIADYIGRRYTIILTAIIFLVGSVLMGYGPSYPILMIGLSISGIGLG 403
Query: 149 FANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWGWRLSLGGA 208
F+ P+Y +E++P +RG L ++S GI + V NYFF K+ GWRL L
Sbjct: 404 FSLIVAPVYCAEISPPSHRGFLTSFLEVSTNAGIALGYVSNYFFGKLSIRLGWRLMLVVH 583
Query: 209 MVPALIITIGSLVLPDTPNSMIERG--------------DRDGAKAQLKRIRGIEDVDEE 254
VP+L + L ++P ++ +G ++ A+ +LK I+ +DE+
Sbjct: 584 AVPSLALVALMLNWVESPRWLVMQGRVGEARKVLLLVSNTKEEAEQRLKEIKVAVGIDEK 763
Query: 255 FNDLVAASEASMQVENPWRN 274
++ +QV N RN
Sbjct: 764 ------CTQDIVQVPNRTRN 805
>AV408698
Length = 199
Score = 79.7 bits (195), Expect = 1e-15
Identities = 42/64 (65%), Positives = 48/64 (74%)
Frame = +1
Query: 143 LGFGIGFANQAVPLYLSEMAPYKYRGALNIGFQLSITIGILVANVLNYFFAKIKGGWGWR 202
LG GIGF NQAVPLYLSEMAP K RGA+N FQ + GIL+AN++NYF KI GWR
Sbjct: 7 LGGGIGFGNQAVPLYLSEMAPAKTRGAVNQLFQFTTCAGILIANLVNYFTEKIH-PHGWR 183
Query: 203 LSLG 206
+SLG
Sbjct: 184 ISLG 195
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.143 0.444
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,233,979
Number of Sequences: 28460
Number of extensions: 132722
Number of successful extensions: 1282
Number of sequences better than 10.0: 80
Number of HSP's better than 10.0 without gapping: 1237
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1252
length of query: 518
length of database: 4,897,600
effective HSP length: 94
effective length of query: 424
effective length of database: 2,222,360
effective search space: 942280640
effective search space used: 942280640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146792.3


