

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
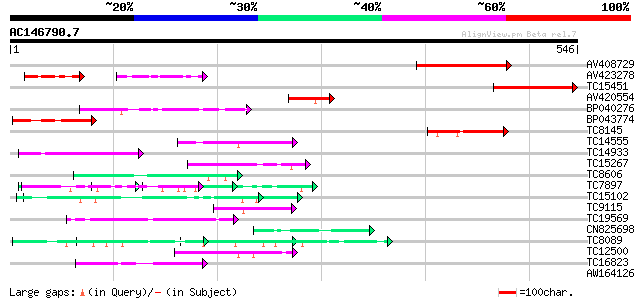
Query= AC146790.7 + phase: 0
(546 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV408729 114 3e-26
AV423278 61 5e-18
TC15451 84 8e-17
AV420554 78 3e-15
BP040276 77 9e-15
BP043774 70 7e-13
TC8145 weakly similar to UP|Q94A38 (Q94A38) AT5g46250/MPL12_3, p... 59 2e-09
TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precu... 50 7e-07
TC14933 homologue to UP|TCMO_GLYEC (Q96423) Trans-cinnamate 4-mo... 46 2e-05
TC15267 similar to UP|EXTN_TOBAC (P13983) Extensin precursor (Ce... 45 2e-05
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 45 4e-05
TC7897 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-li... 44 5e-05
TC15102 similar to UP|Q7X9B3 (Q7X9B3) 9/13 hydroperoxide lyase, ... 44 7e-05
TC9115 similar to PIR|T00425|T00425 photolyase/blue-light recept... 44 7e-05
TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor k... 44 9e-05
CN825698 44 9e-05
TC8089 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinog... 43 1e-04
TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein... 43 1e-04
TC16823 similar to UP|Q8LGA2 (Q8LGA2) Seed maturation-like prote... 42 3e-04
AW164126 35 0.041
>AV408729
Length = 433
Score = 114 bits (286), Expect = 3e-26
Identities = 55/92 (59%), Positives = 69/92 (74%)
Frame = +3
Query: 392 PHMPSPATFFPAAESSLSNVIVNQIDYYFSDINLANDEFLKSNMDEEGWVPITLIANFPR 451
P +P A FF ++ L IVNQIDYYFS+ NL D FL+ NMD++GWVPI LIA F +
Sbjct: 9 PPVPHHAMFFAGSDPQLHTKIVNQIDYYFSNENLVKDTFLRQNMDDQGWVPIKLIAGFKK 188
Query: 452 VKNLTSNIQLILDSMRNSNVVDVQGDKLRRRN 483
V +LT NIQLILD+++ S VV+VQGDK+RRRN
Sbjct: 189 VMHLTDNIQLILDAVQTSYVVEVQGDKIRRRN 284
>AV423278
Length = 389
Score = 61.2 bits (147), Expect(2) = 5e-18
Identities = 43/87 (49%), Positives = 50/87 (57%)
Frame = +2
Query: 104 PLPAPMEKNSNTIASDGGDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAK 163
P P P+ +S SD NAG KKPAW K P + VA E+ PVMGA SWPALSE+AK
Sbjct: 170 PPPCPLIHDS----SDADKAKNAGAPKKPAW-KMPPSDVA-EVSPVMGAVSWPALSEAAK 331
Query: 164 IPGKLPPESSSSKIAPPAAVDGSPSTS 190
+ K P S SS DGS ST+
Sbjct: 332 LSVK--PTSDSS------TTDGSLSTT 388
Score = 46.6 bits (109), Expect(2) = 5e-18
Identities = 32/58 (55%), Positives = 35/58 (60%)
Frame = +1
Query: 15 SPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSSSSSSSS 72
SP +A NN RKNLPSPWAQVVR D+E HQSLPSSSSS+SS
Sbjct: 37 SPNTTTAAANNSP--RKNLPSPWAQVVRCA---DSEPLH------HQSLPSSSSSTSS 177
>TC15451
Length = 662
Score = 83.6 bits (205), Expect = 8e-17
Identities = 46/82 (56%), Positives = 61/82 (74%), Gaps = 2/82 (2%)
Frame = +1
Query: 467 RNSNVVDVQGDKLRRRN-WIDRLTSAQVQADAGSVSPIESRDNSFTADFQTITLDKTTKD 525
R S VV+VQGD LRRRN W+ L SAQ++AD+ S++P SR + ADFQT+++++TT D
Sbjct: 1 RTSTVVEVQGDLLRRRNDWMRWLPSAQLRADSSSLAPGRSRYANLEADFQTLSVEETTND 180
Query: 526 EG-ESSRQSQLSNGSDVAGNIN 546
EG ++ QSQL NGSD AGN N
Sbjct: 181 EGPKTFSQSQLPNGSDAAGNNN 246
>AV420554
Length = 170
Score = 78.2 bits (191), Expect = 3e-15
Identities = 36/54 (66%), Positives = 40/54 (73%), Gaps = 10/54 (18%)
Frame = +1
Query: 269 LPSIPDSSPRDHHRNNNWDARPFVG----------GSSRRGHFGSHPRGDGSYH 312
+PSIPD S RDH+RNNNWDARP VG GSSRRG+FG+HPRGD SYH
Sbjct: 4 VPSIPDPSRRDHYRNNNWDARPLVGGFVPRMNEQRGSSRRGNFGAHPRGDSSYH 165
>BP040276
Length = 556
Score = 76.6 bits (187), Expect = 9e-15
Identities = 61/169 (36%), Positives = 80/169 (47%), Gaps = 3/169 (1%)
Frame = +1
Query: 68 SSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLP---APMEKNSNTIASDGGDGG 124
S +++S P P AA SSP AV VS AP E+ N
Sbjct: 118 SGAAASPNHSPSPSPWSQIVSAAAPPSSPPAVSVSDSSAVNSAPAEETEN---------- 267
Query: 125 NAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVD 184
NA G K+PAWNK P N + VMGA+SWPALS+SA P K PES + +
Sbjct: 268 NANGGKRPAWNK-PSNAAS---SSVMGADSWPALSKSAHSPAKSLPESVKGSVDSSSP-- 429
Query: 185 GSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNI 233
SP+ + SPQRQ + ++ NN LP + +P +R S SN+
Sbjct: 430 -SPNLQGTGSATPSPQRQ-----VRDNAGVNNTLPTQQKPYKR-SNSNL 555
>BP043774
Length = 439
Score = 70.5 bits (171), Expect = 7e-13
Identities = 39/81 (48%), Positives = 49/81 (60%)
Frame = +1
Query: 3 TTQNNHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQS 62
++ N+HSPT+ PSP P RKNLPSPWAQVVRGG ++ + GIH+S
Sbjct: 211 SSSNHHSPTTAPSPVPSGP--------RKNLPSPWAQVVRGG-----DTDFTAAVGIHES 351
Query: 63 LPSSSSSSSSSLTTVDQPPPS 83
PSSSSSSSS T + P+
Sbjct: 352 PPSSSSSSSSLATGAAEQAPA 414
>TC8145 weakly similar to UP|Q94A38 (Q94A38) AT5g46250/MPL12_3, partial
(49%)
Length = 1671
Score = 59.3 bits (142), Expect = 2e-09
Identities = 36/87 (41%), Positives = 56/87 (63%), Gaps = 9/87 (10%)
Frame = +1
Query: 403 AAESSLSN-----VIVNQIDYYFSDINLANDE----FLKSNMDEEGWVPITLIANFPRVK 453
AAE+ LS I+ Q++YYFSD NL ND+ FLK N EG+VP ++IA+F ++K
Sbjct: 310 AAETGLSLDDLKLKIIRQVEYYFSDENLPNDKYLLGFLKKN--REGFVPTSVIASFRKMK 483
Query: 454 NLTSNIQLILDSMRNSNVVDVQGDKLR 480
LT + +I+ +++ S+++ V GD R
Sbjct: 484 RLTRDHSIIVAALKESSLLVVSGDGRR 564
>TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (43%)
Length = 1038
Score = 50.4 bits (119), Expect = 7e-07
Identities = 36/120 (30%), Positives = 51/120 (42%), Gaps = 4/120 (3%)
Frame = +2
Query: 162 AKIPGKLPPESSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLP-- 219
A + + P S ++ APP +P+T Q P SP S+ SS + P
Sbjct: 170 ASVGAQSPSNSPTTSPAPP-----TPTTPQPPAAQSSPPPAQSSPPPVQSSPPPASTPPP 334
Query: 220 --NRPRPMRRISGSNIGPCPAQSSLSNPPTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSP 277
+ P P+ P PA +S P +PPP P P PPA + P L +P +SP
Sbjct: 335 AQSSPPPVSSPPPVQSTPPPAPASTPPPASPPPFSPPPATPPPPAATPPPALTPVPATSP 514
Score = 37.7 bits (86), Expect = 0.005
Identities = 49/188 (26%), Positives = 69/188 (36%), Gaps = 3/188 (1%)
Frame = +2
Query: 64 PSSSSSSSSS--LTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGG 121
PS+S ++S + T QPP + SP A S P V SSP PA
Sbjct: 191 PSNSPTTSPAPPTPTTPQPPAAQSSPPPAQSSPPP---VQSSPPPA-------------- 319
Query: 122 DGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPA 181
+ PA + P PV P S P PP +S +PP
Sbjct: 320 ------STPPPAQSSPP---------PVSSPP--PVQSTPPPAPASTPPPASPPPFSPPP 448
Query: 182 AVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRP-RPMRRISGSNIGPCPAQS 240
A P+ + P ++ P + + K S + P P+ +S S P P+ S
Sbjct: 449 ATPPPPAATPPPALTPVPATSPAPAPAKVKSKSPAPAPAPALAPVVSLSPSE-APGPSLS 625
Query: 241 SLSNPPTP 248
SLS +P
Sbjct: 626 SLSPALSP 649
Score = 35.0 bits (79), Expect = 0.031
Identities = 49/191 (25%), Positives = 70/191 (35%), Gaps = 11/191 (5%)
Frame = +2
Query: 7 NHSPTSVPSPAPPSAETNNPNFIRKNLPS-----PWAQVVRGGSGWDTESQSQ-----SP 56
++SPT+ SPAPP+ T P + + P P Q + +QS SP
Sbjct: 194 SNSPTT--SPAPPTPTTPQPPAAQSSPPPAQSSPPPVQSSPPPASTPPPAQSSPPPVSSP 367
Query: 57 TGIHQSLPSSSSSSSSSLTTVD-QPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNT 115
+ + P + +S+ + PPP+ P AA + V +SP PAP +
Sbjct: 368 PPVQSTPPPAPASTPPPASPPPFSPPPATPPPPAATPPPALTPVPATSPAPAPAKVK--- 538
Query: 116 IASDGGDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSS 175
SK PA P V + P ++ PG P SS S
Sbjct: 539 -------------SKSPAPAPAPALAPVVSLSP-------------SEAPG--PSLSSLS 634
Query: 176 KIAPPAAVDGS 186
PAA D S
Sbjct: 635 PALSPAASDDS 667
Score = 33.9 bits (76), Expect = 0.069
Identities = 27/86 (31%), Positives = 33/86 (37%), Gaps = 2/86 (2%)
Frame = +2
Query: 194 IISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSSLSNPPTPPPL-- 251
+I+ + S S T S + P P + P P QSS TPPP
Sbjct: 164 VIASVGAQSPSNSPTTSPAPPTPTTPQPPAAQSSPPPAQSSPPPVQSSPPPASTPPPAQS 343
Query: 252 PPYPVYQLPPAVSYPNMLPSIPDSSP 277
P PV PP S P P P S+P
Sbjct: 344 SPPPVSSPPPVQSTP---PPAPASTP 412
Score = 29.3 bits (64), Expect = 1.7
Identities = 47/192 (24%), Positives = 57/192 (29%), Gaps = 13/192 (6%)
Frame = +2
Query: 220 NRPRPMRRISGSN-------------IGPCPAQSSLSNPPTPPPLPPYPVYQLPPAVSYP 266
N+PR M+ + N I AQS SN PT P PP P PPA
Sbjct: 89 NQPREMKMMDTHNMHVVLVLGLICIVIASVGAQSP-SNSPTTSPAPPTPTTPQPPAAQSS 265
Query: 267 NMLPSIPDSSPRDHHRNNNWDARPFVGGSSRRGHFGSHPRGDGSYHQNSYSSRREHDRGN 326
P SSP + + P SS
Sbjct: 266 ---PPPAQSSPPPVQSSPPPASTPPPAQSS------------------------------ 346
Query: 327 YANTRDAHAPQPRMPPRGILRPPPPSTAAFLGPQTIGPFPAPVAYPDFYYFPTVPLDPFR 386
P P P + PPP+ A+ P + PF P A P P P
Sbjct: 347 ---------PPPVSSPPPVQSTPPPAPASTPPPASPPPFSPPPATPP----PPAATPPPA 487
Query: 387 GMPVFPHMPSPA 398
PV P+PA
Sbjct: 488 LTPVPATSPAPA 523
>TC14933 homologue to UP|TCMO_GLYEC (Q96423) Trans-cinnamate 4-monooxygenase
(Cinnamic acid 4-hydroxylase) (CA4H) (C4H) (P450C4H)
(Cytochrome P450 73) , partial (57%)
Length = 917
Score = 45.8 bits (107), Expect = 2e-05
Identities = 36/121 (29%), Positives = 53/121 (43%)
Frame = +2
Query: 9 SPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSSSS 68
SP S PSP+ SA + + + + P P G T + + SPT + S SSSS
Sbjct: 101 SPLSSPSPSQNSAASVSSSLLA---PPPSPSSATGSKSAMTSTTATSPTWQNASATSSSS 271
Query: 69 SSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGNAGG 128
+ S+ +++ P S VS+S V SS +P + + S GG GG
Sbjct: 272 AWGSATSSLSHPLNSPRKSSTHRVSNSGPERVTSSSTSSPEKDKTWCSPSTASTGGRCGG 451
Query: 129 S 129
S
Sbjct: 452 S 454
>TC15267 similar to UP|EXTN_TOBAC (P13983) Extensin precursor (Cell wall
hydroxyproline-rich glycoprotein), partial (6%)
Length = 1003
Score = 45.4 bits (106), Expect = 2e-05
Identities = 36/120 (30%), Positives = 51/120 (42%), Gaps = 2/120 (1%)
Frame = +1
Query: 172 SSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGS 231
+++ K+ PP + S+S P +S SP + S NNNL +P P + S
Sbjct: 121 TTTVKLPPP*SPSSPSSSSLSPWLSPSPPSSTTPPPPPSHRTNNNNLSTQPTPSAPSATS 300
Query: 232 NIGPCPAQSSLSNPPTPPPLPPYPVYQLPPAVSYPNMLP--SIPDSSPRDHHRNNNWDAR 289
PA +PP+PPP P + P+ P+ P S P S PR R AR
Sbjct: 301 LASRTPA-----SPPSPPPTTTPPTHN--PSSPSPSAPPWTSSPPSPPRSESRPQVLTAR 459
Score = 36.2 bits (82), Expect = 0.014
Identities = 30/104 (28%), Positives = 39/104 (36%)
Frame = +1
Query: 10 PTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSSSSS 69
P+S P PPS TNN N S PT S PS++S
Sbjct: 205 PSSTTPPPPPSHRTNNNNL------------------------STQPT---PSAPSATSL 303
Query: 70 SSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNS 113
+S + + PPP+ P S SP A +S P+P S
Sbjct: 304 ASRTPASPPSPPPTTTPPTHNPSSPSPSAPPWTSSPPSPPRSES 435
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 44.7 bits (104), Expect = 4e-05
Identities = 51/181 (28%), Positives = 71/181 (39%), Gaps = 18/181 (9%)
Frame = +1
Query: 62 SLPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGG 121
S SSSSSSSSS + P S S + SSS + SS +P +S++ S G
Sbjct: 271 SASSSSSSSSSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSSPSSSSSSSPFSSG- 447
Query: 122 DGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLP-PESSSSKIAPP 180
PA P +G E+ + +PA S K P S SS +P
Sbjct: 448 ----------PAPAPGPPSGAFFELLLLFELLPYPAPSSDDKSSSPSPNGPSPSSSSSPS 597
Query: 181 AAVDGSPSTS--------------QGPIISHSPQRQGSTS---NTKSSSMANNNLPNRPR 223
+++ SPS S +IS+SPQR + S T ++ N P
Sbjct: 598 SSLPSSPSASCCLSSL*DLVSPLVVAMLISNSPQRSAAKSVALTTAEATKVTTTSANAPP 777
Query: 224 P 224
P
Sbjct: 778 P 780
Score = 29.3 bits (64), Expect = 1.7
Identities = 31/131 (23%), Positives = 56/131 (42%)
Frame = +1
Query: 3 TTQNNHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQS 62
++ ++ S +S+ S + PS+ +++P PS + S S S + S
Sbjct: 280 SSSSSSSSSSLSSSSTPSSSSSSP-------PS-------------SSSSSSSSSSSSSS 399
Query: 63 LPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGD 122
S SSSSSSS + P P+ P A + +++ LP P + + +S +
Sbjct: 400 WSSPSSSSSSSPFS-SGPAPAPGPPSGAFF----ELLLLFELLPYPAPSSDDKSSSPSPN 564
Query: 123 GGNAGGSKKPA 133
G + S P+
Sbjct: 565 GPSPSSSSSPS 597
Score = 28.9 bits (63), Expect = 2.2
Identities = 32/109 (29%), Positives = 48/109 (43%), Gaps = 4/109 (3%)
Frame = +1
Query: 172 SSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGS 231
SSSS + + +PS+S S SP S+S++ SSS ++ + P+ S S
Sbjct: 283 SSSSSSSSSLSSSSTPSSS-----SSSPPSSSSSSSSSSSSSSSWSSPS--------SSS 423
Query: 232 NIGPCPAQSSLSNPPTPPPLPP----YPVYQLPPAVSYPNMLPSIPDSS 276
+ P S+ P P P PP + + L + YP PS D S
Sbjct: 424 SSSP------FSSGPAPAPGPPSGAFFELLLLFELLPYP--APSSDDKS 546
>TC7897 weakly similar to UP|ENL2_ARATH (Q9T076) Early nodulin-like protein
2 precursor (Phytocyanin-like protein), partial (31%)
Length = 1482
Score = 44.3 bits (103), Expect = 5e-05
Identities = 44/159 (27%), Positives = 62/159 (38%), Gaps = 18/159 (11%)
Frame = +3
Query: 79 QPPPS-------DDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGNAGGSKK 131
QPPPS SPKA+ S SP SP P + + GG + G+
Sbjct: 525 QPPPSPPPKSSPSPSPKASPPSGSPPTAGEPSPAGTPPSVLPSPAGAPVPAGGPSAGTPP 704
Query: 132 PAWNKQPLNGVAVE--IGPVMGAESWPALSESAKIPGKLPPESSSSKI---------APP 180
A + V+ P GAES P+ S ++ P P +S S AP
Sbjct: 705 SASTSPSPSPVSAPPAASPAPGAES-PSSSPTSNSPAGGPTATSPSPAGDSPAGGPPAPS 881
Query: 181 AAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLP 219
+P ++ GP +S S ++T SSS N+ P
Sbjct: 882 PTTGDTPPSASGPDVSPSGTPSSLPADTPSSSSNNSTAP 998
Score = 43.9 bits (102), Expect = 7e-05
Identities = 33/124 (26%), Positives = 46/124 (36%), Gaps = 6/124 (4%)
Frame = +3
Query: 9 SPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPT------GIHQS 62
SP P PA + P+ PSP + ES S SPT G +
Sbjct: 651 SPAGAPVPAGGPSAGTPPSASTSPSPSPVSAPPAASPAPGAESPSSSPTSNSPAGGPTAT 830
Query: 63 LPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGD 122
PS + S + P D P A+ SP S P P ++N+ A G+
Sbjct: 831 SPSPAGDSPAGGPPAPSPTTGDTPPSASGPDVSPSGTPSSLPADTPSSSSNNSTAPPPGN 1010
Query: 123 GGNA 126
GG++
Sbjct: 1011GGSS 1022
Score = 42.7 bits (99), Expect = 1e-04
Identities = 56/180 (31%), Positives = 74/180 (41%), Gaps = 5/180 (2%)
Frame = +3
Query: 12 SVPSPAPPSAETNNPNFIRKN-LPSP-WAQVVRGGSGWDTESQSQSPTGIHQSLPSSSSS 69
S PS +PP+A +P + LPSP A V GG T PS+S+S
Sbjct: 579 SPPSGSPPTAGEPSPAGTPPSVLPSPAGAPVPAGGPSAGTP-------------PSASTS 719
Query: 70 SSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGNAGGS 129
S S V PP + +P A SSSP + +SP P + T S GD AGG
Sbjct: 720 PSPS--PVSAPPAASPAPGAESPSSSPTS---NSPAGGP----TATSPSPAGD-SPAGGP 869
Query: 130 KKPAWNKQPLNGVAVEIGPVMGAESWPALSES---AKIPGKLPPESSSSKIAPPAAVDGS 186
P + G + S P +S S + +P P SS++ APP GS
Sbjct: 870 PAP----------SPTTGDTPPSASGPDVSPSGTPSSLPADTPSSSSNNSTAPPPGNGGS 1019
Score = 42.0 bits (97), Expect = 3e-04
Identities = 50/177 (28%), Positives = 67/177 (37%), Gaps = 9/177 (5%)
Frame = +3
Query: 129 SKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGK----LPPESSSSKIAPPAAVD 184
S P+ P +G P E PA + + +P +P S+ P A+
Sbjct: 555 SPSPSPKASPPSG-----SPPTAGEPSPAGTPPSVLPSPAGAPVPAGGPSAGTPPSASTS 719
Query: 185 GSPS-TSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSSLS 243
SPS S P S +P + +S+ S+S A P P G PA
Sbjct: 720 PSPSPVSAPPAASPAPGAESPSSSPTSNSPAGGPTATSPSPA--------GDSPA----G 863
Query: 244 NPPTPPPLPPYPVYQLPPAVSYPNMLPS-IPDSSPRD---HHRNNNWDARPFVGGSS 296
PP P P PP+ S P++ PS P S P D NN+ P GGSS
Sbjct: 864 GPPAPSPTTG----DTPPSASGPDVSPSGTPSSLPADTPSSSSNNSTAPPPGNGGSS 1022
Score = 32.3 bits (72), Expect = 0.20
Identities = 28/93 (30%), Positives = 44/93 (47%)
Frame = +3
Query: 3 TTQNNHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQS 62
++ ++SP P+ PS ++P PSP G + SP+G S
Sbjct: 786 SSPTSNSPAGGPTATSPSPAGDSPAG-GPPAPSP----TTGDTPPSASGPDVSPSGTPSS 950
Query: 63 LPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSS 95
LP+ + SSSS+ +T PPP + ++VV SS
Sbjct: 951 LPADTPSSSSNNSTA--PPPGNGG--SSVVPSS 1037
>TC15102 similar to UP|Q7X9B3 (Q7X9B3) 9/13 hydroperoxide lyase, partial
(68%)
Length = 1097
Score = 43.9 bits (102), Expect = 7e-05
Identities = 61/289 (21%), Positives = 98/289 (33%), Gaps = 20/289 (6%)
Frame = +1
Query: 14 PSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSS------S 67
PS + +TN + +R + W Q ++ + +P+ + PSS +
Sbjct: 7 PSQFEKTKQTNRYSSVRSSHQQQWQQHLQTTPKTAAATFL*NPSLEATASPSSEPCTTAT 186
Query: 68 SSSSSSLTTVDQPP------PSDDSPKAAVVSSSPKAVVVS-SPLPAPMEKNSNTIASDG 120
++S++ T PP P P + +SP S S P P +S T S
Sbjct: 187 TTSTTKAATNSSPPESRSTTPPSSEPTCLLAPASPPTPKSSHSSTPPPSRSSSTTPKSKN 366
Query: 121 GDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPP 180
A PA + + +A P K PP ++ P
Sbjct: 367 ATSLTAPSCPPPA-----------------------SPAATAPAPSKTPPNPPTNSSKPS 477
Query: 181 AAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRP--RPMRRISGSNIGPC-- 236
+ P+T+ + S P S +T SS + P P R S S+ P
Sbjct: 478 SCKSSLPNTTPSSLSSAPPSPTTSPISTPSSPENQEKPASTPPSAPPRSTSSSSSSPINT 657
Query: 237 --PAQSSLSNPPTPPPLPPYPVYQLPPAVS-YPNMLPSIPDSSPRDHHR 282
S+ P + PP ++ PP+ S + P I S P R
Sbjct: 658 PRKPSSATKAPASSPPGSELSLHHSPPSASRKSSTTPKISSSEPSHSRR 804
Score = 40.4 bits (93), Expect = 7e-04
Identities = 52/238 (21%), Positives = 82/238 (33%)
Frame = +1
Query: 7 NHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSS 66
N SP S PPS+E P + S S +P S +
Sbjct: 214 NSSPPESRSTTPPSSE-------------PTCLLAPASPPTPKSSHSSTPPPSRSSSTTP 354
Query: 67 SSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGNA 126
S +++SLT PPP+ + A S +P +S P+
Sbjct: 355 KSKNATSLTAPSCPPPASPAATAPAPSKTPPNPPTNSSKPS------------------- 477
Query: 127 GGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVDGS 186
K N P + + P S P+ E+ + P PP APP + +
Sbjct: 478 -SCKSSLPNTTPSSLSSAPPSPTTSPISTPSSPENQEKPASTPPS------APPRS---T 627
Query: 187 PSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSSLSN 244
S+S PI ++P++ S + +SS + L P S+ P + S S+
Sbjct: 628 SSSSSSPI--NTPRKPSSATKAPASSPPGSELSLHHSPPSASRKSSTTPKISSSEPSH 795
Score = 32.7 bits (73), Expect = 0.15
Identities = 28/113 (24%), Positives = 44/113 (38%)
Frame = +1
Query: 3 TTQNNHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQS 62
T N + +S PS S P+ + PSP + S + + P S
Sbjct: 439 TPPNPPTNSSKPSSCKSSLPNTTPSSLSSAPPSPTTSPISTPS---SPENQEKPASTPPS 609
Query: 63 LPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNT 115
P S+SSSSS + ++ P + KA S + + P+ K+S T
Sbjct: 610 APPRSTSSSSS-SPINTPRKPSSATKAPASSPPGSELSLHHSPPSASRKSSTT 765
>TC9115 similar to PIR|T00425|T00425 photolyase/blue-light receptor (PHR2)
[imported] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (55%)
Length = 1018
Score = 43.9 bits (102), Expect = 7e-05
Identities = 28/83 (33%), Positives = 40/83 (47%), Gaps = 3/83 (3%)
Frame = +1
Query: 197 HSPQRQGSTSNTKSSSMANNNLPNRPR--PMRRISGSNIGPCPAQSSLSNPPTPPPLPPY 254
HSP S++ + +NN P R + P S ++ P P NPP+ PP PP
Sbjct: 175 HSPPSSPSSNPNSHHLLPHNNNPTRSKSQPKPPPSPTSHSPPPPHPRRPNPPSNPPSPPT 354
Query: 255 PVYQLPP-AVSYPNMLPSIPDSS 276
P L P A + P+ LP+ P S+
Sbjct: 355 PSAPLTP*APTAPSTLPTPPHSA 423
Score = 28.9 bits (63), Expect = 2.2
Identities = 37/153 (24%), Positives = 52/153 (33%), Gaps = 8/153 (5%)
Frame = +1
Query: 277 PRDHHRNNNWDARPFVGGSSRRGHFGS--HPRGDGSYHQNSYSSRREHDRGNYANTRDAH 334
P+ HH + S + H S H S + S+ H + N
Sbjct: 76 PKYHHHHQR-KTTTLTRNHSHQNHHHSQ*HHSPFHSPPSSPSSNPNSHHLLPHNNNPTRS 252
Query: 335 APQPRMPPRGILRPPPPSTAAFLGPQTIGPFPAPVAYPDFYYFPTVPLDPFRGMPVFPHM 394
QP+ PP PPP P + P P + P PT P +P PH
Sbjct: 253 KSQPKPPPSPTSHSPPPPHPRRPNPPSNPPSPPTPSAPLTP*APTAP----STLPTPPHS 420
Query: 395 PSP-----ATFFPAAESSLSN-VIVNQIDYYFS 421
+P AT +A + LS N+ +Y S
Sbjct: 421 AAPPSSGSATTSASATTRLSTPPTTNRSQFYLS 519
>TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor kinase 1;1,
partial (16%)
Length = 486
Score = 43.5 bits (101), Expect = 9e-05
Identities = 43/167 (25%), Positives = 69/167 (40%), Gaps = 1/167 (0%)
Frame = +1
Query: 55 SPTGIHQSLPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSN 114
SP G+ L +SSSSSSS PPP S+SP + P P + +N
Sbjct: 19 SPCGL---LLHTSSSSSSS----YSPPPPPSQQPTHSPSTSPTSTTTPPPSPTKVTENPP 177
Query: 115 TIASDGGDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPES-S 173
T S + S++P+ P N + + P S S P PP + +
Sbjct: 178 TAPSTSTKSPTSSASEEPS----PPNPSTSTTQQPKHSPTSPPASPSPSPPSTKPPTAMA 345
Query: 174 SSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPN 220
S +P +A P+ P S +P +T++ K+ S +++ P+
Sbjct: 346 SPSTSPLSATKSHPTPPAAPSASSTP--PPTTTSLKTMSSPSSSTPS 480
Score = 36.2 bits (82), Expect = 0.014
Identities = 29/111 (26%), Positives = 40/111 (35%), Gaps = 2/111 (1%)
Frame = +1
Query: 7 NHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSS 66
+ PT PS +P S T P+ + P A S + S+ SP S
Sbjct: 88 SQQPTHSPSTSPTSTTTPPPSPTKVTENPPTAPSTSTKSPTSSASEEPSPPNPSTSTTQQ 267
Query: 67 SSSSSSSLTTVDQPPPSDDSPKAAVV--SSSPKAVVVSSPLPAPMEKNSNT 115
S +S P P P A+ S+SP + S P P S+T
Sbjct: 268 PKHSPTSPPASPSPSPPSTKPPTAMASPSTSPLSATKSHPTPPAAPSASST 420
Score = 35.0 bits (79), Expect = 0.031
Identities = 29/110 (26%), Positives = 47/110 (42%), Gaps = 3/110 (2%)
Frame = +1
Query: 3 TTQNNHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQS 62
++ +++SP PS P + + +P PSP + T ++S PT
Sbjct: 52 SSSSSYSPPPPPSQQPTHSPSTSPTSTTTPPPSPTKVTENPPTAPSTSTKS--PTSSASE 225
Query: 63 LPSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKA---VVVSSPLPAPM 109
PS + S+S T QP S SP A+ S P ++SP +P+
Sbjct: 226 EPSPPNPSTS---TTQQPKHSPTSPPASPSPSPPSTKPPTAMASPSTSPL 366
Score = 32.0 bits (71), Expect = 0.26
Identities = 28/82 (34%), Positives = 37/82 (44%), Gaps = 11/82 (13%)
Frame = +1
Query: 3 TTQNNHSPTSVP---SPAPPSAE--------TNNPNFIRKNLPSPWAQVVRGGSGWDTES 51
T Q HSPTS P SP+PPS + + +P K+ P+P A S
Sbjct: 259 TQQPKHSPTSPPASPSPSPPSTKPPTAMASPSTSPLSATKSHPTPPA----------APS 408
Query: 52 QSQSPTGIHQSLPSSSSSSSSS 73
S +P SL + SS SSS+
Sbjct: 409 ASSTPPPTTTSLKTMSSPSSST 474
>CN825698
Length = 704
Score = 43.5 bits (101), Expect = 9e-05
Identities = 37/117 (31%), Positives = 45/117 (37%)
Frame = +1
Query: 235 PCPAQSSLSNPPTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSPRDHHRNNNWDARPFVGG 294
P P+ SS S PP PPP PP P LPP PS D P R + D R F
Sbjct: 103 PPPSPSSASAPPPPPPPPPPP--SLPP--------PSDSDLPPPPARRRDRRDDRDFDRP 252
Query: 295 SSRRGHFGSHPRGDGSYHQNSYSSRREHDRGNYANTRDAHAPQPRMPPRGILRPPPP 351
+RR Y+ S +S DR R + +P + PPPP
Sbjct: 253 PNRR-----------DYYDRSNNSPPPKDRDRDFKRRRSPSPPSQRDRDRRHSPPPP 390
>TC8089 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinogalactan
protein 1 precursor, partial (43%)
Length = 890
Score = 43.1 bits (100), Expect = 1e-04
Identities = 54/219 (24%), Positives = 78/219 (34%), Gaps = 6/219 (2%)
Frame = +2
Query: 65 SSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGG 124
S SSS Q P + + + SS P SP P N G
Sbjct: 179 SRQRCSSSPWLLPPQTPTTSPAFSPSTPSSPPSTTTSPSPTSPPRSTN--------GPQS 334
Query: 125 NAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAVD 184
+ S P W N P S P+ S S P S+ S APP+
Sbjct: 335 PSAPSTTPPWTTSSQN---THQSPPSRTSS-PSTSSSTT---SAPRSSTRSPTAPPSP-- 487
Query: 185 GSPSTSQGPIISHSPQRQGSTSNTKSSSMANN--NLPNRPRPMRRISGSNIGPCPAQSSL 242
P ++ P +P STS T ++ +++ P P+R + P + SS
Sbjct: 488 --PQCTKPPAPPRAPP-DSSTSPTSAAGKSDSAPRTTTAPSPLRSSNP*RKSPTTSPSSR 658
Query: 243 S----NPPTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSP 277
S PP PL P+P+ ++ P + + S P SP
Sbjct: 659 SVRFFPPPQQKPLHPHPLSRISPPLCPSTVARSSPTLSP 775
Score = 43.1 bits (100), Expect = 1e-04
Identities = 54/217 (24%), Positives = 78/217 (35%), Gaps = 13/217 (5%)
Frame = +2
Query: 165 PGKLPPESSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRP 224
P P S S+ +PP+ SPS + P ++ PQ + S T + ++ N P
Sbjct: 227 PTTSPAFSPSTPSSPPSTTT-SPSPTSPPRSTNGPQSPSAPSTTPPWTTSSQNTHQSPPS 403
Query: 225 MRRISGSNIGPCPAQSSLSNPPTPPPLPPY------PVYQLPPAVSYPNMLPSIPDSSPR 278
++ A S + PT PP PP P P + + P DS+PR
Sbjct: 404 RTSSPSTSSSTTSAPRSSTRSPTAPPSPPQCTKPPAPPRAPPDSSTSPTSAAGKSDSAPR 583
Query: 279 DH------HRNNNWDARPFVGGSSRRGHFGSHPRGDGSYHQNSYSSRREHDRGNYANTRD 332
+N P SSR F P+ H + S +
Sbjct: 584 TTTAPSPLRSSNP*RKSPTTSPSSRSVRFFPPPQ-QKPLHPHPLS----RISPPLCPSTV 748
Query: 333 AHAPQPRMPPRGILRPPPPSTAAFLGPQTIG-PFPAP 368
A + PPR P P+T+ T+G PFPAP
Sbjct: 749 ARSSPTLSPPRRTRTQPSPTTS------TVG*PFPAP 841
Score = 42.7 bits (99), Expect = 1e-04
Identities = 56/218 (25%), Positives = 84/218 (37%), Gaps = 29/218 (13%)
Frame = +2
Query: 3 TTQNNHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQS--QSPTGIH 60
T + S T+ PSP P TN P +P W T SQ+ QSP
Sbjct: 257 TPSSPPSTTTSPSPTSPPRSTNGPQSPSAPSTTP---------PWTTSSQNTHQSPPSRT 409
Query: 61 QSLPSSSSSSSSSLTTVDQ-------------------PPPSDDSPKAAV--VSSSPKAV 99
S +SSS++S+ ++ PP S SP +A S+P+
Sbjct: 410 SSPSTSSSTTSAPRSSTRSPTAPPSPPQCTKPPAPPRAPPDSSTSPTSAAGKSDSAPRTT 589
Query: 100 VVSSPLPA--PMEKNSNTIASDGGDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPA 157
SPL + P K+ T S +KP + PL+ ++ + P A S P
Sbjct: 590 TAPSPLRSSNP*RKSPTTSPSSRSVRFFPPPQQKPL-HPHPLSRISPPLCPSTVARSSPT 766
Query: 158 LSESAKIPGKLPPESSSSKIAPPAAVDG----SPSTSQ 191
LS PP + ++ +P + G +PST+Q
Sbjct: 767 LS---------PPRRTRTQPSPTTSTVG*PFPAPSTTQ 853
>TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein {Danio
rerio;} , partial (5%)
Length = 678
Score = 43.1 bits (100), Expect = 1e-04
Identities = 39/128 (30%), Positives = 52/128 (40%), Gaps = 9/128 (7%)
Frame = +1
Query: 159 SESAKIPGKLPPESSSSKIAPPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNL 218
++ A +P P S + PP +P + IIS + G S S S
Sbjct: 28 ADRAPLPPPSTPPKSLKALPPPPPPPSAPFIAGSAIISKVSKSVGDVSLPCSPSPPPPQP 207
Query: 219 P------NRPRPMRRISGSNI--GPCPAQSSLSNPPTPP-PLPPYPVYQLPPAVSYPNML 269
P P P I+GS I G SL +PP PP P P Y V +PP+ P +L
Sbjct: 208 PPPSLPPTTPPPASVIAGSAIMSGEASFPCSLPSPPQPPPPPPKYGVTTIPPS---PPLL 378
Query: 270 PSIPDSSP 277
S D +P
Sbjct: 379 MSATDRAP 402
Score = 39.3 bits (90), Expect = 0.002
Identities = 55/251 (21%), Positives = 85/251 (32%), Gaps = 5/251 (1%)
Frame = +1
Query: 9 SPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIHQSLPSSSS 68
S P P PPS + + P P A + G + S+S SLP S S
Sbjct: 25 SADRAPLP-PPSTPPKSLKALPPPPPPPSAPFIAGSAIISKVSKSVGDV----SLPCSPS 189
Query: 69 SSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDGGNAGG 128
PPP P + P +V+ S + + G+
Sbjct: 190 P----------PPPQPPPPSLPPTTPPPASVIAGSAIMS-------------GEASFPCS 300
Query: 129 SKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSS-SKIAPPAAVDGSP 187
P P V P P L SA LPP S+ + + P+
Sbjct: 301 LPSPPQPPPPPPKYGVTTIPPS-----PPLLMSATDRAPLPPPSTHLTSLTTPSPPSSRT 465
Query: 188 STSQGPIISH----SPQRQGSTSNTKSSSMANNNLPNRPRPMRRISGSNIGPCPAQSSLS 243
+ +Q P+ S SP + + + +++ P P+ +++ P +LS
Sbjct: 466 TITQTPLPSPPQPPSPPPKYGVATIPPPPLPMSSIDRTPLPLLSTPLTSLDAPPPTPTLS 645
Query: 244 NPPTPPPLPPY 254
PP PPP PP+
Sbjct: 646 PPPPPPPTPPH 678
Score = 31.6 bits (70), Expect = 0.34
Identities = 47/223 (21%), Positives = 78/223 (34%), Gaps = 29/223 (13%)
Frame = +1
Query: 68 SSSSSSLTTVDQPPPSDDSP---KAAVVSSSPKAV----VVSSPLPAPMEKNSNTIASDG 120
S+ SL + PPP +P +A++S K+V + SP P P + ++
Sbjct: 55 STPPKSLKALPPPPPPPSAPFIAGSAIISKVSKSVGDVSLPCSPSPPPPQPPPPSL---- 222
Query: 121 GDGGNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKI--A 178
P P + +A + G S+P S P PP+ + I +
Sbjct: 223 -----------PPTTPPPASVIAGS-AIMSGEASFPCSLPSPPQPPPPPPKYGVTTIPPS 366
Query: 179 PPAAVDGSPSTSQGPIISHSPQRQGSTSNTKSSSMANNNLPNRPRPMR---RISGSNIGP 235
PP + + P +H + + +++ LP+ P+P + + I P
Sbjct: 367 PPLLMSATDRAPLPPPSTHLTSLTTPSPPSSRTTITQTPLPSPPQPPSPPPKYGVATIPP 546
Query: 236 CPAQSSLSN-----------------PPTPPPLPPYPVYQLPP 261
P S + PPTP PP P PP
Sbjct: 547 PPLPMSSIDRTPLPLLSTPLTSLDAPPPTPTLSPPPPPPPTPP 675
>TC16823 similar to UP|Q8LGA2 (Q8LGA2) Seed maturation-like protein, partial
(29%)
Length = 711
Score = 42.0 bits (97), Expect = 3e-04
Identities = 36/127 (28%), Positives = 56/127 (43%)
Frame = +1
Query: 64 PSSSSSSSSSLTTVDQPPPSDDSPKAAVVSSSPKAVVVSSPLPAPMEKNSNTIASDGGDG 123
PSSS +S +S + P S AA +SSP A + S P+ + T++S G+
Sbjct: 61 PSSSHASPTSHPILSTHPSHHLSSSAAAAASSPPATPMGSR--RPLLPHPLTMSSPPGN- 231
Query: 124 GNAGGSKKPAWNKQPLNGVAVEIGPVMGAESWPALSESAKIPGKLPPESSSSKIAPPAAV 183
P + A P A S P+++ S++ PPE+ S+ +PP +V
Sbjct: 232 --------------PPSTAASPSSPTCSAASSPSITPSSQRVSPTPPETP*SRPSPPCSV 369
Query: 184 DGSPSTS 190
P TS
Sbjct: 370 CSPPITS 390
Score = 30.4 bits (67), Expect = 0.77
Identities = 26/93 (27%), Positives = 37/93 (38%)
Frame = +1
Query: 1 MVTTQNNHSPTSVPSPAPPSAETNNPNFIRKNLPSPWAQVVRGGSGWDTESQSQSPTGIH 60
+++T +H +S + A S R LP P G+ T + SPT
Sbjct: 97 ILSTHPSHHLSSSAAAAASSPPATPMGSRRPLLPHPLTMSSPPGNPPSTAASPSSPTCSA 276
Query: 61 QSLPSSSSSSSSSLTTVDQPPPSDDSPKAAVVS 93
S PS + SS T + P S SP +V S
Sbjct: 277 ASSPSITPSSQRVSPTPPETP*SRPSPPCSVCS 375
>AW164126
Length = 387
Score = 34.7 bits (78), Expect = 0.041
Identities = 22/72 (30%), Positives = 28/72 (38%), Gaps = 5/72 (6%)
Frame = +2
Query: 336 PQPRMPPRGILRPPPPSTAA-----FLGPQTIGPFPAPVAYPDFYYFPTVPLDPFRGMPV 390
P P P PPPPS + + P P P+PV +P +YY P P P
Sbjct: 152 PSPSPKPYYYKSPPPPSPSPPPPYYYKSP----PPPSPVPHPPYYYKSPPPPSPIPHPPY 319
Query: 391 FPHMPSPATFFP 402
+ P P P
Sbjct: 320 YYSSPPPPVKSP 355
Score = 30.8 bits (68), Expect = 0.59
Identities = 20/71 (28%), Positives = 28/71 (39%)
Frame = +3
Query: 235 PCPAQSSLSNPPTPPPLPPYPVYQLPPAVSYPNMLPSIPDSSPRDHHRNNNWDARPFVGG 294
P P PP+P P PPY Y+ PP S PS+ ++ HH ++ +
Sbjct: 27 PPPYYYKSPPPPSPVPHPPY-YYKSPPPPS-----PSLNHTTTSPHHHPHHLQNHTTINH 188
Query: 295 SSRRGHFGSHP 305
H HP
Sbjct: 189 LHHLHHLLHHP 221
Score = 28.9 bits (63), Expect = 2.2
Identities = 18/51 (35%), Positives = 19/51 (36%), Gaps = 5/51 (9%)
Frame = +2
Query: 235 PCPAQSSLSNPPTPPPLPPYPVYQLPPAVSYPNMLP-----SIPDSSPRDH 280
P P +PP P P PP P Y P P P S P SP H
Sbjct: 158 PSPKPYYYKSPPPPSPSPPPPYYYKSPPPPSPVPHPPYYYKSPPPPSPIPH 310
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.309 0.128 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,816,230
Number of Sequences: 28460
Number of extensions: 281669
Number of successful extensions: 7160
Number of sequences better than 10.0: 702
Number of HSP's better than 10.0 without gapping: 3815
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5910
length of query: 546
length of database: 4,897,600
effective HSP length: 95
effective length of query: 451
effective length of database: 2,193,900
effective search space: 989448900
effective search space used: 989448900
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146790.7


