

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146790.10 - phase: 0
(406 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
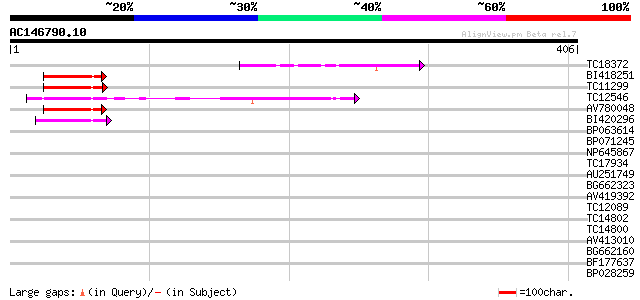
Score E
Sequences producing significant alignments: (bits) Value
TC18372 72 2e-13
BI418251 51 3e-07
TC11299 47 4e-06
TC12546 weakly similar to UP|Q9FJ30 (Q9FJ30) Similarity to heat ... 46 1e-05
AV780048 45 3e-05
BI420296 44 5e-05
BP063614 39 0.001
BP071245 37 0.006
NP645867 proliferating floral organs protein [Lotus japonicus] 36 0.013
TC17934 similar to PIR|T06667|T06667 argininosuccinate synthase ... 34 0.038
AU251749 33 0.086
BG662323 33 0.11
AV419392 32 0.15
TC12089 similar to UP|Q944K0 (Q944K0) At1g18030/T10F20_3, partia... 32 0.25
TC14802 weakly similar to UP|CAD56661 (CAD56661) S locus F-box (... 31 0.42
TC14800 weakly similar to UP|Q9FHP3 (Q9FHP3) Genomic DNA, chromo... 31 0.42
AV413010 30 0.55
BG662160 30 0.72
BF177637 30 0.72
BP028259 29 1.2
>TC18372
Length = 452
Score = 71.6 bits (174), Expect = 2e-13
Identities = 48/136 (35%), Positives = 77/136 (56%), Gaps = 3/136 (2%)
Frame = +1
Query: 165 FSLLINN-KRDISLQNTWISSILNRRVKILRIHSSFYQLPFSALTSHYLFNCTSLEELEL 223
FSL + + D S N WISS+L+R ++ ++H Y L+SH LF+C SL EL+L
Sbjct: 1 FSLCLTSFNHDSSQVNAWISSVLHRGLQ--KLHIQCYDKVL--LSSHSLFSCNSLVELKL 168
Query: 224 VLHVSSTIKFPSISVHFGHLKLLKLYGIFFKIDTSSDC--LTLNLPLLRKFDIKNCNWSG 281
+ T+ P IS +L+ + G+ D+ +D +TL+ P+L+ F+ + C W
Sbjct: 169 --QMKCTLNVP-ISAFLPNLQSFSISGVKLVSDSPTDSKDITLSFPVLKVFEARGCEWLT 339
Query: 282 GKDLIVEAPLLEIVSI 297
+D+ ++APLLE SI
Sbjct: 340 MQDISIQAPLLEKFSI 387
>BI418251
Length = 546
Score = 51.2 bits (121), Expect = 3e-07
Identities = 24/45 (53%), Positives = 32/45 (70%)
Frame = +1
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDD 69
SKLP+ ++ ++LSFLPT+D V T VL KRW + W S+ LD DD
Sbjct: 46 SKLPDEVLCQILSFLPTEDAVATSVLCKRWSSLWL-SVPTLDFDD 177
>TC11299
Length = 498
Score = 47.4 bits (111), Expect = 4e-06
Identities = 22/46 (47%), Positives = 31/46 (66%)
Frame = +1
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDI 70
S LP+ +I +LSFLPT++ V T VLSKRW + W S++ LD +
Sbjct: 94 SALPDEIICHILSFLPTENAVATAVLSKRWTHLW-RSVSALDFSSV 228
>TC12546 weakly similar to UP|Q9FJ30 (Q9FJ30) Similarity to heat shock
transcription factor HSF30, partial (9%)
Length = 755
Score = 45.8 bits (107), Expect = 1e-05
Identities = 59/242 (24%), Positives = 103/242 (42%), Gaps = 4/242 (1%)
Frame = +2
Query: 13 QMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDL 72
+M + ED I S LP++L+ +LSFL +K+ V T VLS+RW+ W S+ L D
Sbjct: 125 KMKKMKMEDGI-STLPDALLCHILSFLTSKEAVATSVLSRRWIPLW-RSVPTLHFKDA-- 292
Query: 73 SYNYNPLCSYCEEICTDYCYDYDKYKLTVCPRCHVCGNHKANLRDAEVYERRVEIDFESG 132
NY+ + D+D K +V R V+ +
Sbjct: 293 --NYHTDIGHA---------DHDIVK--------------------DVRSRHVQSVY--- 370
Query: 133 MKIESGKKEQQFVNFVGRALLLTSISSMELERFSLLINNK----RDISLQNTWISSILNR 188
A++L+ + +++F L +N+ D + + W+++++ R
Sbjct: 371 ------------------AVILSRDFQLPIKKFYLRLNDVCQPFYDPANVSVWVNAVVQR 496
Query: 189 RVKILRIHSSFYQLPFSALTSHYLFNCTSLEELELVLHVSSTIKFPSISVHFGHLKLLKL 248
+++ L I + L +F+C +L L+L + +FP SVHF LK+L L
Sbjct: 497 QLEHLDISLPYPMLSTPRANLSSIFSCRTLVVLKLRGGLELK-RFP--SVHFPCLKVLHL 667
Query: 249 YG 250
G
Sbjct: 668 QG 673
>AV780048
Length = 501
Score = 44.7 bits (104), Expect = 3e-05
Identities = 21/45 (46%), Positives = 29/45 (63%)
Frame = +2
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDD 69
S LP+ ++ +LS L TK+ V T +LSKRW+ W S+ LD DD
Sbjct: 260 SSLPDPILCHILSLLTTKEAVTTSILSKRWIPLW-RSVPTLDFDD 391
>BI420296
Length = 592
Score = 43.9 bits (102), Expect = 5e-05
Identities = 23/55 (41%), Positives = 32/55 (57%)
Frame = +2
Query: 19 EEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWTSSITKLDLDDIDLS 73
EE S LP+ ++ VL F+ TK V+TCVLS RW + W +T L L+ D +
Sbjct: 185 EERDRLSDLPDLVLLHVLKFMSTKQAVQTCVLSTRWKDLW-KGVTTLALNSSDFA 346
>BP063614
Length = 508
Score = 39.3 bits (90), Expect = 0.001
Identities = 15/34 (44%), Positives = 23/34 (67%)
Frame = +2
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRW 58
S LP+ ++ +LSF+ K V+TC+LS RW + W
Sbjct: 176 SDLPDCILLHILSFVKAKAAVQTCILSTRWKDLW 277
>BP071245
Length = 552
Score = 37.0 bits (84), Expect = 0.006
Identities = 15/34 (44%), Positives = 22/34 (64%)
Frame = +1
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRW 58
S LP+ ++ +LSF+ K V+TC LS RW + W
Sbjct: 391 SDLPDCVLLHILSFVNAKYAVQTCTLSTRWKDLW 492
>NP645867 proliferating floral organs protein [Lotus japonicus]
Length = 1350
Score = 35.8 bits (81), Expect = 0.013
Identities = 22/57 (38%), Positives = 31/57 (53%), Gaps = 6/57 (10%)
Frame = +1
Query: 4 GSSKVHKSTQMASVIEEDT------INSKLPESLINRVLSFLPTKDVVRTCVLSKRW 54
GSS + T A+ +T I SKLP+ L++RV++FLPT R + KRW
Sbjct: 97 GSSNLTTPTPTATGTYNNTPWMNSRIWSKLPQKLLDRVIAFLPTPAFFRARSVCKRW 267
>TC17934 similar to PIR|T06667|T06667 argininosuccinate synthase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (11%)
Length = 564
Score = 34.3 bits (77), Expect = 0.038
Identities = 12/35 (34%), Positives = 22/35 (62%)
Frame = -1
Query: 25 SKLPESLINRVLSFLPTKDVVRTCVLSKRWMNRWT 59
++LP+ + +LS LP + +T +LS++W WT
Sbjct: 561 NQLPDGIPGAILSKLPINEAAKTSILSRKWRYMWT 457
>AU251749
Length = 362
Score = 33.1 bits (74), Expect = 0.086
Identities = 14/23 (60%), Positives = 19/23 (81%)
Frame = +2
Query: 25 SKLPESLINRVLSFLPTKDVVRT 47
S LP++++N +LSFLPTK VV T
Sbjct: 146 SALPDTILNFILSFLPTKQVVAT 214
>BG662323
Length = 375
Score = 32.7 bits (73), Expect = 0.11
Identities = 15/50 (30%), Positives = 26/50 (52%), Gaps = 7/50 (14%)
Frame = +1
Query: 12 TQMASVIEED-------TINSKLPESLINRVLSFLPTKDVVRTCVLSKRW 54
T+++ EED +++ LP L+ +L +LP + R C + KRW
Sbjct: 10 TRLSEPGEEDEKEGTIVSVDEVLPGDLLEHILMYLPIASIFRACCVCKRW 159
>AV419392
Length = 318
Score = 32.3 bits (72), Expect = 0.15
Identities = 12/24 (50%), Positives = 18/24 (75%)
Frame = +3
Query: 35 VLSFLPTKDVVRTCVLSKRWMNRW 58
+LS++ TKD ++ VLSKRW + W
Sbjct: 6 ILSYVETKDAMQCSVLSKRWRSLW 77
>TC12089 similar to UP|Q944K0 (Q944K0) At1g18030/T10F20_3, partial (25%)
Length = 631
Score = 31.6 bits (70), Expect = 0.25
Identities = 17/53 (32%), Positives = 26/53 (48%)
Frame = -1
Query: 54 WMNRWTSSITKLDLDDIDLSYNYNPLCSYCEEICTDYCYDYDKYKLTVCPRCH 106
W ++ + I L + + YNY PL +C I + Y KYK +C +CH
Sbjct: 532 WSSKLDNYIK*LTANASFIYYNY-PLLQHC--ITITIMHKYQKYKTLICVQCH 383
>TC14802 weakly similar to UP|CAD56661 (CAD56661) S locus F-box (SLF)-S4
protein, partial (10%)
Length = 551
Score = 30.8 bits (68), Expect = 0.42
Identities = 13/28 (46%), Positives = 20/28 (71%)
Frame = +3
Query: 27 LPESLINRVLSFLPTKDVVRTCVLSKRW 54
LP+ LI +LS+LP K +++ V+SK W
Sbjct: 21 LPDELIIEILSWLPVKSLLQFRVVSKTW 104
>TC14800 weakly similar to UP|Q9FHP3 (Q9FHP3) Genomic DNA, chromosome 5,
TAC clone:K14B20, partial (7%)
Length = 782
Score = 30.8 bits (68), Expect = 0.42
Identities = 13/28 (46%), Positives = 20/28 (71%)
Frame = +3
Query: 27 LPESLINRVLSFLPTKDVVRTCVLSKRW 54
LP+ LI +LS+LP K +++ V+SK W
Sbjct: 48 LPDELIIEILSWLPVKSLLQFRVVSKTW 131
>AV413010
Length = 416
Score = 30.4 bits (67), Expect = 0.55
Identities = 15/55 (27%), Positives = 30/55 (54%)
Frame = +3
Query: 3 AGSSKVHKSTQMASVIEEDTINSKLPESLINRVLSFLPTKDVVRTCVLSKRWMNR 57
AG+ + + + E +++S LP+ LI ++L LP ++R +SK W ++
Sbjct: 24 AGAERKMNEQRRSCTEENPSLSSILPDELIIQILLQLPVTSLLRFKSVSKSWFSQ 188
>BG662160
Length = 445
Score = 30.0 bits (66), Expect = 0.72
Identities = 14/28 (50%), Positives = 18/28 (64%)
Frame = +2
Query: 27 LPESLINRVLSFLPTKDVVRTCVLSKRW 54
LPE L +LS+LP K +VR +SK W
Sbjct: 23 LPEELRLEILSWLPVKSLVRFRCVSKSW 106
>BF177637
Length = 480
Score = 30.0 bits (66), Expect = 0.72
Identities = 12/25 (48%), Positives = 15/25 (60%)
Frame = -3
Query: 82 YCEEICTDYCYDYDKYKLTVCPRCH 106
+C C DY YDYD KL++C H
Sbjct: 106 FCFFFCYDYDYDYDFLKLSLCSFDH 32
>BP028259
Length = 502
Score = 29.3 bits (64), Expect = 1.2
Identities = 21/69 (30%), Positives = 34/69 (48%), Gaps = 5/69 (7%)
Frame = -1
Query: 170 NNKRDISLQNTWISSILNRRVKILRIHSSFYQLPFSALTSHYLF---NCTSLEELEL--V 224
+N R ++ Q ++ ++ R + I SSFY F TSHY F C S+ +L
Sbjct: 262 HNTRAVT-QGSFSDIVVQRH*HVYGILSSFYSTTFVRSTSHYCFFHHTCISIFDLTTPDF 86
Query: 225 LHVSSTIKF 233
+H + T +F
Sbjct: 85 MHDTPTCRF 59
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,825,424
Number of Sequences: 28460
Number of extensions: 124075
Number of successful extensions: 849
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 838
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 845
length of query: 406
length of database: 4,897,600
effective HSP length: 92
effective length of query: 314
effective length of database: 2,279,280
effective search space: 715693920
effective search space used: 715693920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146790.10


