

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146789.1 + phase: 0
(431 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
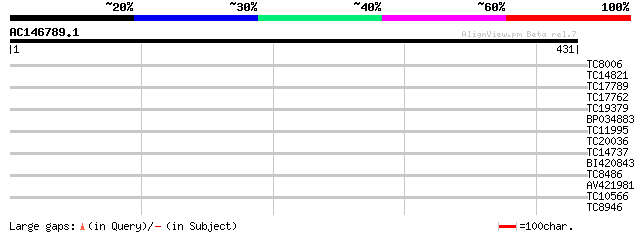
Score E
Sequences producing significant alignments: (bits) Value
TC8006 similar to UP|Q9LVE9 (Q9LVE9) Genomic DNA, chromosome 3, ... 28 0.12
TC14821 homologue to UP|AAR12195 (AAR12195) Molecular chaperone ... 30 0.77
TC17789 similar to PIR|T05768|T05768 subtilisin-like proteinase ... 28 2.9
TC17762 28 3.8
TC19379 homologue to UP|Q945P0 (Q945P0) Transcriptional activato... 28 3.8
BP034883 27 5.0
TC11995 27 6.5
TC20036 similar to UP|Q9LLM5 (Q9LLM5) Outward-rectifying potassi... 27 6.5
TC14737 similar to UP|Q9LEB3 (Q9LEB3) RNA binding protein 47, pa... 27 6.5
BI420843 27 8.5
TC8486 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Ara... 27 8.5
AV421981 27 8.5
TC10566 similar to UP|Q9LI73 (Q9LI73) Chloroplast nucleoid DNA b... 27 8.5
TC8946 homologue to GB|AAM66972.1|21617922|AY088650 pyridoxine b... 27 8.5
>TC8006 similar to UP|Q9LVE9 (Q9LVE9) Genomic DNA, chromosome 3, P1 clone:
MIL23, partial (27%)
Length = 1225
Score = 28.1 bits (61), Expect(2) = 0.12
Identities = 19/67 (28%), Positives = 27/67 (39%)
Frame = -1
Query: 361 YEARSSPDTYDMVSGAVAYADAQLGQEEVMSMTPQQWYEAMTHMREQIAPILTRRRAQRP 420
Y R D YD + + Q E P+ +H R + P RR+ +RP
Sbjct: 745 YRLREGKDGYDPIQNPGSLTRPGSFQPEQAPQIPRGGV-VNSHRRRNLLP--KRRQRKRP 575
Query: 421 RRRHHHQ 427
RRR H +
Sbjct: 574 RRRRHRK 554
Score = 23.1 bits (48), Expect(2) = 0.12
Identities = 10/12 (83%), Positives = 11/12 (91%)
Frame = -3
Query: 334 PQILPPIPGDLP 345
PQ+LPPIP DLP
Sbjct: 863 PQLLPPIP-DLP 831
>TC14821 homologue to UP|AAR12195 (AAR12195) Molecular chaperone Hsp90-1,
partial (29%)
Length = 612
Score = 30.0 bits (66), Expect = 0.77
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = -1
Query: 333 HPQILPPIPGDLPRPANEEQIIAEQW 358
HPQ+ PP P LPRP +++ QW
Sbjct: 123 HPQLCPPRPSSLPRPHHQKS--PSQW 52
>TC17789 similar to PIR|T05768|T05768 subtilisin-like proteinase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (14%)
Length = 940
Score = 28.1 bits (61), Expect = 2.9
Identities = 11/23 (47%), Positives = 14/23 (60%)
Frame = -1
Query: 405 REQIAPILTRRRAQRPRRRHHHQ 427
R I P+L R R +PRR H H+
Sbjct: 553 RHHITPLLHRLREHKPRRLHRHR 485
>TC17762
Length = 463
Score = 27.7 bits (60), Expect = 3.8
Identities = 11/37 (29%), Positives = 20/37 (53%)
Frame = +2
Query: 391 SMTPQQWYEAMTHMREQIAPILTRRRAQRPRRRHHHQ 427
SM+P + + A +++P + + + P R HHHQ
Sbjct: 152 SMSPSREHAATMGNNRRLSPFVLSGQPRAPSRLHHHQ 262
>TC19379 homologue to UP|Q945P0 (Q945P0) Transcriptional activator FHA1,
partial (56%)
Length = 525
Score = 27.7 bits (60), Expect = 3.8
Identities = 12/22 (54%), Positives = 14/22 (63%)
Frame = +2
Query: 405 REQIAPILTRRRAQRPRRRHHH 426
R Q+ I RRR +PRRRH H
Sbjct: 125 RPQLQEIHRRRRPLQPRRRHEH 190
>BP034883
Length = 553
Score = 27.3 bits (59), Expect = 5.0
Identities = 12/35 (34%), Positives = 20/35 (56%), Gaps = 3/35 (8%)
Frame = +2
Query: 155 ELADACRPGHRALGGSVTLLTT---WFLKHFPGFF 186
E+++A +P H G + +T +FL+H P FF
Sbjct: 209 EISEASQPYHSGSGSISSFITASFLFFLRHLPSFF 313
>TC11995
Length = 726
Score = 26.9 bits (58), Expect = 6.5
Identities = 12/37 (32%), Positives = 22/37 (59%)
Frame = +1
Query: 70 RDFYTSYLGRANVLVGTEDPEEVEELERVRTYCVRCY 106
R F T+YLGR V+ T + ++ EE+ + + +R +
Sbjct: 298 RSFQTNYLGRLTVIHNTLEGKKKEEILFCQVFVLRVF 408
>TC20036 similar to UP|Q9LLM5 (Q9LLM5) Outward-rectifying potassium channel
KCO1, partial (34%)
Length = 470
Score = 26.9 bits (58), Expect = 6.5
Identities = 10/21 (47%), Positives = 15/21 (70%)
Frame = -1
Query: 409 APILTRRRAQRPRRRHHHQDQ 429
+P R + Q+P++ HHHQDQ
Sbjct: 245 SPSTYR*QIQQPQQSHHHQDQ 183
>TC14737 similar to UP|Q9LEB3 (Q9LEB3) RNA binding protein 47, partial (32%)
Length = 803
Score = 26.9 bits (58), Expect = 6.5
Identities = 22/95 (23%), Positives = 37/95 (38%), Gaps = 6/95 (6%)
Frame = +3
Query: 343 DLPRPANEEQIIAEQWQRYEARSSPDTYDMVSGAVAYADAQLGQEEVMSMTPQQWYEAMT 402
DL + A +Q +QW + + Y A A ++ M M PQ + +
Sbjct: 150 DLNQNAPPQQQQPQQWAAQQQQWMAMQYP--------ATAMAMMQQQMMMYPQHYMPYVH 305
Query: 403 H------MREQIAPILTRRRAQRPRRRHHHQDQDQ 431
H ++Q A Q+ ++HHHQ Q +
Sbjct: 306 HHYQPHHQQQQAAAAAAGAVPQQQHQQHHHQQQQK 410
>BI420843
Length = 377
Score = 26.6 bits (57), Expect = 8.5
Identities = 12/24 (50%), Positives = 15/24 (62%)
Frame = +1
Query: 400 AMTHMREQIAPILTRRRAQRPRRR 423
AMTH R + ++ RRR RPR R
Sbjct: 208 AMTHQRRRCTGMMLRRRFGRPRWR 279
>TC8486 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Arabidopsis
thaliana;}, partial (25%)
Length = 856
Score = 26.6 bits (57), Expect = 8.5
Identities = 14/26 (53%), Positives = 16/26 (60%)
Frame = +3
Query: 402 THMREQIAPILTRRRAQRPRRRHHHQ 427
TH R+ +L RRR Q PRRR HQ
Sbjct: 192 THPRQPA--LLRRRRHQDPRRRPEHQ 263
>AV421981
Length = 468
Score = 26.6 bits (57), Expect = 8.5
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = +3
Query: 278 PRHPTDVRDLPPPSIVQMFID 298
P HPT +R LPPP+I ++
Sbjct: 129 PLHPTLLRHLPPPTISTSLVE 191
>TC10566 similar to UP|Q9LI73 (Q9LI73) Chloroplast nucleoid DNA binding
protein-like, nucellin-like protein, partial (25%)
Length = 774
Score = 26.6 bits (57), Expect = 8.5
Identities = 18/47 (38%), Positives = 22/47 (46%)
Frame = +2
Query: 167 LGGSVTLLTTWFLKHFPGFFSVDLNTDYLEKYPVAARWKLEKGHGEG 213
+GG + LL F+KHF GF+ D L P R K HG G
Sbjct: 509 VGGKMLLLQCIFIKHFFGFYFFSEFCDVL*CSPEGKR----K*HGGG 637
>TC8946 homologue to GB|AAM66972.1|21617922|AY088650 pyridoxine
biosynthesis protein-like {Arabidopsis thaliana;} ,
partial (45%)
Length = 503
Score = 26.6 bits (57), Expect = 8.5
Identities = 13/27 (48%), Positives = 18/27 (66%), Gaps = 2/27 (7%)
Frame = +3
Query: 405 REQIAPIL--TRRRAQRPRRRHHHQDQ 429
R+Q P+L +R R+ PRRRHH + Q
Sbjct: 132 RDQEIPLLRQSRPRSDAPRRRHHGRRQ 212
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.139 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,487,995
Number of Sequences: 28460
Number of extensions: 125211
Number of successful extensions: 720
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 699
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 718
length of query: 431
length of database: 4,897,600
effective HSP length: 93
effective length of query: 338
effective length of database: 2,250,820
effective search space: 760777160
effective search space used: 760777160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146789.1


