

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146784.10 + phase: 0
(168 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
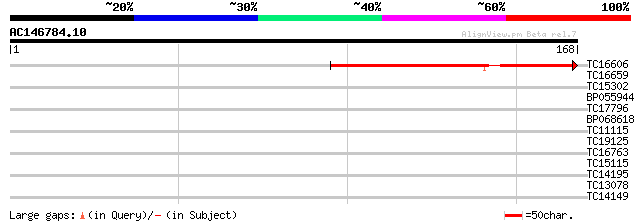
TC16606 127 1e-30
TC16659 similar to UP|Q9LKZ5 (Q9LKZ5) Receptor-like protein kina... 27 1.8
TC15302 homologue to UP|O65673 (O65673) NONCLATHRIN coat protein... 27 2.4
BP055944 26 3.1
TC17796 similar to PIR|T10625|T10625 reticuline oxidase homolog ... 25 5.3
BP068618 25 5.3
TC11115 weakly similar to UP|NFL1_HUMAN (Q14494) Nuclear factor ... 25 5.3
TC19125 UP|GBA1_LOTJA (P49082) Guanine nucleotide-binding protei... 25 9.0
TC16763 similar to UP|Q8LEE3 (Q8LEE3) Lysophospholipase-like pro... 25 9.0
TC15115 similar to UP|PIN1_MALDO (Q94G00) Peptidyl-prolyl cis-tr... 25 9.0
TC14195 UP|Q40215 (Q40215) RAB8A, complete 25 9.0
TC13078 similar to UP|Q7X9Y1 (Q7X9Y1) Gonidia forming protein Gl... 25 9.0
TC14149 homologue to PIR|T45927|T45927 ribosomal protein S3a hom... 25 9.0
>TC16606
Length = 532
Score = 127 bits (318), Expect = 1e-30
Identities = 62/74 (83%), Positives = 66/74 (88%), Gaps = 1/74 (1%)
Frame = +1
Query: 96 DEIPYQELEMALPESASATVVESAEGWDQGWDDDWDDNVAVKSP-VVRHAGSISANGLTS 154
DEIPYQELEMALP SASAT VESAEGWDQGWDDDWDDNVAVKSP VR ISANGLTS
Sbjct: 4 DEIPYQELEMALPVSASATNVESAEGWDQGWDDDWDDNVAVKSPGAVR---GISANGLTS 174
Query: 155 RSSNKDGWEDNWDD 168
RS+NKDGW++NWD+
Sbjct: 175 RSANKDGWDNNWDE 216
>TC16659 similar to UP|Q9LKZ5 (Q9LKZ5) Receptor-like protein kinase 2,
partial (12%)
Length = 1032
Score = 26.9 bits (58), Expect = 1.8
Identities = 12/32 (37%), Positives = 20/32 (62%)
Frame = -1
Query: 116 VESAEGWDQGWDDDWDDNVAVKSPVVRHAGSI 147
+E+A G + +DD +D+V VKSP + S+
Sbjct: 306 LEAAVGDSRAFDDGKEDSVIVKSPCLEPGSSV 211
>TC15302 homologue to UP|O65673 (O65673) NONCLATHRIN coat protein gamma -
like protein (NONCLATHRIN coat protein gamma-like
protein), partial (20%)
Length = 605
Score = 26.6 bits (57), Expect = 2.4
Identities = 11/38 (28%), Positives = 20/38 (51%)
Frame = -2
Query: 11 YNIVFQVTIKQIKSESTELTLDAGKGDCVLHVTVVTPV 48
YN +FQ+++ K S T + G C +H+++ V
Sbjct: 565 YNCLFQISLNLGKESSIRNTTEHTNGICPIHISLAIHV 452
>BP055944
Length = 163
Score = 26.2 bits (56), Expect = 3.1
Identities = 9/18 (50%), Positives = 14/18 (77%)
Frame = -2
Query: 6 LAYCFYNIVFQVTIKQIK 23
L +CFY ++FQ+ I Q+K
Sbjct: 87 LLFCFY*VIFQMNIYQLK 34
>TC17796 similar to PIR|T10625|T10625 reticuline oxidase homolog F21C20.180
- Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(35%)
Length = 594
Score = 25.4 bits (54), Expect = 5.3
Identities = 15/53 (28%), Positives = 22/53 (41%), Gaps = 6/53 (11%)
Frame = +1
Query: 11 YNIVFQVTIKQIKSES------TELTLDAGKGDCVLHVTVVTPVPEASFFLRL 57
+ ++ T+K +K E TLD D VL V P + F+RL
Sbjct: 274 FGVILSYTVKLVKVPEIVTVFRVEKTLDQNATDLVLQWQQVAPTTDDRLFMRL 432
>BP068618
Length = 420
Score = 25.4 bits (54), Expect = 5.3
Identities = 14/45 (31%), Positives = 22/45 (48%), Gaps = 3/45 (6%)
Frame = +2
Query: 55 LRLPSFDKILT---PVNGAYFLIFTVIVFAVTWACCCIFKKKPRD 96
L L F KIL P+ + +++ A+ W C +F + PRD
Sbjct: 224 LDLIYFLKILVGECPIKDMQIWVHPLLIIALGWECESLFIQPPRD 358
>TC11115 weakly similar to UP|NFL1_HUMAN (Q14494) Nuclear factor erythroid 2
related factor 1 (NF-E2 related factor 1) (NFE2-related
factor 1) (Nuclear factor, erythroid derived 2, like 1)
(Transcription factor 11) (Transcription factor HBZ17)
(Transcription factor LCR-F1) (Locus con, partial (3%)
Length = 622
Score = 25.4 bits (54), Expect = 5.3
Identities = 10/19 (52%), Positives = 13/19 (67%), Gaps = 2/19 (10%)
Frame = +1
Query: 117 ESAEGWD--QGWDDDWDDN 133
+ EG D Q W+DDWDD+
Sbjct: 190 DKEEGKDVTQQWEDDWDDD 246
>TC19125 UP|GBA1_LOTJA (P49082) Guanine nucleotide-binding protein alpha-1
subunit (GP-alpha-1), complete
Length = 1597
Score = 24.6 bits (52), Expect = 9.0
Identities = 8/18 (44%), Positives = 9/18 (49%)
Frame = +3
Query: 71 YFLIFTVIVFAVTWACCC 88
YF+ I F W CCC
Sbjct: 1458 YFIFTFHIAFPCPWLCCC 1511
>TC16763 similar to UP|Q8LEE3 (Q8LEE3) Lysophospholipase-like protein,
partial (22%)
Length = 594
Score = 24.6 bits (52), Expect = 9.0
Identities = 9/28 (32%), Positives = 14/28 (49%)
Frame = -2
Query: 87 CCIFKKKPRDEIPYQELEMALPESASAT 114
CC +P D++PYQ+ + S T
Sbjct: 137 CCTDPARPADQLPYQQFHAEQRTAQSQT 54
>TC15115 similar to UP|PIN1_MALDO (Q94G00) Peptidyl-prolyl cis-trans
isomerase 1 (Rotamase Pin1) (PPIase Pin1) (MdPin1) ,
partial (98%)
Length = 737
Score = 24.6 bits (52), Expect = 9.0
Identities = 14/34 (41%), Positives = 17/34 (49%)
Frame = -2
Query: 53 FFLRLPSFDKILTPVNGAYFLIFTVIVFAVTWAC 86
FFLRL SF A+FL + +FAV C
Sbjct: 103 FFLRLLSFSARSAAEIDAFFLF*SESIFAVAVTC 2
>TC14195 UP|Q40215 (Q40215) RAB8A, complete
Length = 1171
Score = 24.6 bits (52), Expect = 9.0
Identities = 12/27 (44%), Positives = 17/27 (62%)
Frame = +1
Query: 57 LPSFDKILTPVNGAYFLIFTVIVFAVT 83
LP+F + VN YF +F ++ FAVT
Sbjct: 958 LPTF*MLCFKVNILYFSLF*LVEFAVT 1038
>TC13078 similar to UP|Q7X9Y1 (Q7X9Y1) Gonidia forming protein GlsA, partial
(9%)
Length = 564
Score = 24.6 bits (52), Expect = 9.0
Identities = 10/26 (38%), Positives = 20/26 (76%)
Frame = +3
Query: 137 KSPVVRHAGSISANGLTSRSSNKDGW 162
KS +++ +++ANG++S SS++D W
Sbjct: 93 KSTDDQNSTAVAANGVSSSSSDQDIW 170
>TC14149 homologue to PIR|T45927|T45927 ribosomal protein S3a homolog -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(87%)
Length = 1191
Score = 24.6 bits (52), Expect = 9.0
Identities = 12/30 (40%), Positives = 15/30 (50%), Gaps = 4/30 (13%)
Frame = -2
Query: 72 FLIFTVIVFA----VTWACCCIFKKKPRDE 97
FL+ V +F+ V CCC F PR E
Sbjct: 152 FLLIWVAIFSQLREVRHCCCCCFVANPRFE 63
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.135 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,163,439
Number of Sequences: 28460
Number of extensions: 39852
Number of successful extensions: 310
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 296
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 300
length of query: 168
length of database: 4,897,600
effective HSP length: 84
effective length of query: 84
effective length of database: 2,506,960
effective search space: 210584640
effective search space used: 210584640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146784.10


