

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146759.10 + phase: 1 /partial
(178 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
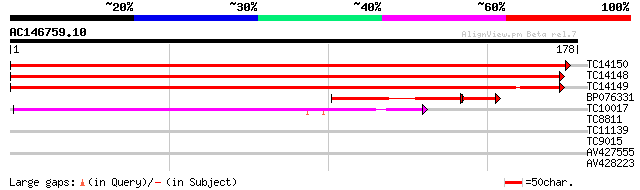
TC14150 homologue to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S3... 339 1e-94
TC14148 similar to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S3 (... 335 2e-93
TC14149 homologue to PIR|T45927|T45927 ribosomal protein S3a hom... 335 2e-93
BP076331 48 3e-08
TC10017 UP|RR3_LOTJA (Q9BBP8) Chloroplast 30S ribosomal protein ... 43 3e-05
TC8811 similar to UP|AAP04330 (AAP04330) G-protein-coupled recep... 30 0.24
TC11139 26 4.5
TC9015 similar to UP|Q9SS90 (Q9SS90) F4P13.19 protein, partial (... 25 7.7
AV427555 25 7.7
AV428223 25 7.7
>TC14150 homologue to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S3
(AT5g35530/MOK9_14), partial (91%)
Length = 1075
Score = 339 bits (870), Expect = 1e-94
Identities = 170/176 (96%), Positives = 173/176 (97%)
Frame = +3
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY
Sbjct: 261 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 440
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 120
GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG
Sbjct: 441 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 620
Query: 121 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDIEVPVPV 176
VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEE+EYIRPAAV A ++EVPV V
Sbjct: 621 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEDEYIRPAAVSAEEVEVPVAV 788
>TC14148 similar to UP|Q9FJA6 (Q9FJA6) 40S ribosomal protein S3
(AT5g35530/MOK9_14), partial (91%)
Length = 1032
Score = 335 bits (860), Expect = 2e-93
Identities = 168/174 (96%), Positives = 172/174 (98%)
Frame = +3
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
EKGRRIRELTSVV KRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY
Sbjct: 258 EKGRRIRELTSVVSKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 437
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 120
GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG
Sbjct: 438 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 617
Query: 121 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDIEVPV 174
VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIH+PKEE+EYI+PAAV A+DIEVPV
Sbjct: 618 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHSPKEEDEYIQPAAVSASDIEVPV 779
>TC14149 homologue to PIR|T45927|T45927 ribosomal protein S3a homolog -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(87%)
Length = 1191
Score = 335 bits (860), Expect = 2e-93
Identities = 169/174 (97%), Positives = 172/174 (98%)
Frame = +1
Query: 1 EKGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 60
EKGRRIRELTS+VQKRFKFPEN+VELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY
Sbjct: 310 EKGRRIRELTSIVQKRFKFPENNVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACY 489
Query: 61 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 120
GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG
Sbjct: 490 GVLRFVMESGAKGCEVIVSGKLRAQRAKSMKFKDGYMISSGQPVKDYIDSAVRHVLLRQG 669
Query: 121 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAVVANDIEVPV 174
VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEY RP AV+ANDIEVPV
Sbjct: 670 VLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEY-RPVAVLANDIEVPV 828
>BP076331
Length = 358
Score = 48.1 bits (113), Expect(2) = 3e-08
Identities = 24/42 (57%), Positives = 29/42 (68%)
Frame = -1
Query: 102 QPVKDYIDSAVRHVLLRQGVLGIKVKIMLDWDPKGKQGPKTP 143
QP+ +Y +S VRH+LLRQ IMLD DPKGK GP+TP
Sbjct: 358 QPLNNYNESVVRHILLRQ--------IMLD*DPKGKHGPQTP 257
Score = 24.3 bits (51), Expect(2) = 3e-08
Identities = 10/12 (83%), Positives = 11/12 (91%)
Frame = -3
Query: 143 PLPDIVTIHTPK 154
PL +IVTIHTPK
Sbjct: 260 PLLNIVTIHTPK 225
>TC10017 UP|RR3_LOTJA (Q9BBP8) Chloroplast 30S ribosomal protein S3,
complete
Length = 1185
Score = 43.1 bits (100), Expect = 3e-05
Identities = 34/132 (25%), Positives = 65/132 (48%), Gaps = 2/132 (1%)
Frame = +1
Query: 2 KGRRIRELTSVVQKRFKFPENSVELYAEKVNNRGLCAIAQAESLRYKLLGGLAVRRACYG 61
K RRI EL VQK+ + + + ++ N AE + +L ++ R+A
Sbjct: 262 KPRRIEELQINVQKKMNYVNRKLNIAITRIANAYRDPNILAEFIAGQLKNRVSFRKAMKK 441
Query: 62 VLRFVMESGAKGCEVIVSGKLRAQRAKSMKF-KDGYM-ISSGQPVKDYIDSAVRHVLLRQ 119
+ ++G KG +V ++G++ + +++ ++G + + + + DY VR +
Sbjct: 442 AIELTEQAGTKGVQVQIAGRIDGKEIARVEWIREGRVPLQTIRAKIDYCSYTVRTI---Y 612
Query: 120 GVLGIKVKIMLD 131
GVLGIKV I L+
Sbjct: 613 GVLGIKVWIFLN 648
>TC8811 similar to UP|AAP04330 (AAP04330) G-protein-coupled receptor GPR34
type 2a (Fragment), partial (6%)
Length = 675
Score = 30.0 bits (66), Expect = 0.24
Identities = 15/39 (38%), Positives = 21/39 (53%)
Frame = -3
Query: 140 PKTPLPDIVTIHTPKEEEEYIRPAAVVANDIEVPVPVPV 178
P+ P+P + +TP E + VV + VPVPVPV
Sbjct: 172 PEGPMPVPIPPNTPGVELNGVVEPVVVEEGVPVPVPVPV 56
>TC11139
Length = 659
Score = 25.8 bits (55), Expect = 4.5
Identities = 11/25 (44%), Positives = 15/25 (60%)
Frame = +3
Query: 59 CYGVLRFVMESGAKGCEVIVSGKLR 83
C+G+ + E + GC VIVS LR
Sbjct: 270 CFGIEPSIPEGKSYGCAVIVSSDLR 344
>TC9015 similar to UP|Q9SS90 (Q9SS90) F4P13.19 protein, partial (12%)
Length = 639
Score = 25.0 bits (53), Expect = 7.7
Identities = 13/35 (37%), Positives = 20/35 (57%), Gaps = 6/35 (17%)
Frame = -2
Query: 117 LRQGVLGIKV------KIMLDWDPKGKQGPKTPLP 145
L++G LG K+ ++ D +PKG +GPK P
Sbjct: 146 LKEGTLGSKLVA*EGFEVPKDSEPKGFEGPKDAEP 42
>AV427555
Length = 346
Score = 25.0 bits (53), Expect = 7.7
Identities = 19/69 (27%), Positives = 31/69 (44%)
Frame = -2
Query: 106 DYIDSAVRHVLLRQGVLGIKVKIMLDWDPKGKQGPKTPLPDIVTIHTPKEEEEYIRPAAV 165
D+I VRH+LL + LG+K+ P+ +H + E YIR + V
Sbjct: 213 DFICVHVRHLLLHRRELGVKL--------------VHPITKHRGVHRWRLERGYIRLSVV 76
Query: 166 VANDIEVPV 174
+ N + + V
Sbjct: 75 LRNRLHLVV 49
>AV428223
Length = 340
Score = 25.0 bits (53), Expect = 7.7
Identities = 12/28 (42%), Positives = 16/28 (56%)
Frame = +3
Query: 68 ESGAKGCEVIVSGKLRAQRAKSMKFKDG 95
ESG G EVIV +R+ + +F DG
Sbjct: 183 ESGVSGGEVIVRADVRSLESVGRRFIDG 266
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.138 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,447,083
Number of Sequences: 28460
Number of extensions: 27782
Number of successful extensions: 140
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 137
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 137
length of query: 178
length of database: 4,897,600
effective HSP length: 84
effective length of query: 94
effective length of database: 2,506,960
effective search space: 235654240
effective search space used: 235654240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146759.10


