

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146751.11 - phase: 0
(399 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
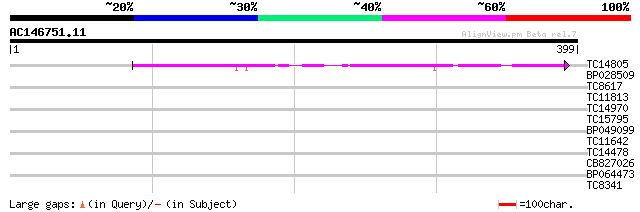
TC14805 similar to UP|RP3A_ARATH (Q39211) DNA-directed RNA polym... 140 4e-34
BP028509 29 1.2
TC8617 similar to UP|Q9SSZ4 (Q9SSZ4) Asparaginyl endopeptidase, ... 28 2.1
TC11813 similar to UP|Q8LJ68 (Q8LJ68) Protein phosphatase 2C-lik... 28 2.7
TC14970 similar to UP|O24294 (O24294) Legumin (Minor small) prec... 28 3.5
TC15795 weakly similar to UP|Q8H1T9 (Q8H1T9) Phospholipase D del... 28 3.5
BP049099 27 4.6
TC11642 similar to UP|Q9FHL3 (Q9FHL3) Genomic DNA, chromosome 5,... 27 6.0
TC14478 weakly similar to UP|Q94G86 (Q94G86) Beta-1,3-glucanase-... 27 7.8
CB827026 27 7.8
BP064473 27 7.8
TC8341 similar to UP|Q94FX1 (Q94FX1) Heme oxygenase 1 (Fragment)... 27 7.8
>TC14805 similar to UP|RP3A_ARATH (Q39211) DNA-directed RNA polymerase II 36
kDa polypeptide A (RNA polymerase II subunit 3) ,
complete
Length = 1273
Score = 140 bits (353), Expect = 4e-34
Identities = 104/317 (32%), Positives = 161/317 (49%), Gaps = 9/317 (2%)
Frame = +2
Query: 87 KVEVKRITDEEMEFDMIGIDPAVANAFRRILIAEVPTMAIERVYIANNTSLVQDEVLSHR 146
+V+++ + D+ F++ D ++ANA RR++I+EVPT+AI+ V I N+S++ DE ++HR
Sbjct: 71 RVKIRELKDDMARFELRDTDASIANALRRVMISEVPTIAIDLVEIEVNSSVLNDEFIAHR 250
Query: 147 LGLIPIDADPKL---FEYPDDA--GGENNEKNTIVFKQHVRCEKGQPRLTVKSDTLKWLP 201
LGLIP+ ++ + F DA G E ++ F V+C + L V S L
Sbjct: 251 LGLIPLTSERAMAMRFSRDCDACDGDGQCEFCSVEFHLRVKCMTDE-TLDVTSKDL---- 415
Query: 202 NGSELIAEGTKSAADPAPKTFTTFNQKSLPKFSKDPAPYNLDVIVAKLGPGQEIELEAHA 261
++ T D +P T S D A N +I+ KL GQE+ L A A
Sbjct: 416 ----YSSDHTVVPVDFSPDT------------SSDSA-VNKGIIIVKLRRGQELRLRAIA 544
Query: 262 VKGIGKTHAKWSPVATAWYRMLPEVVLTKDVKDELA----EELVSKCPAKVFDIEDIGRG 317
KGIGK HAKWSP AT + P++ + D+ + L +E V CP VFDI+
Sbjct: 545 RKGIGKDHAKWSPAATVSFMYKPQIHINDDLMETLTLKEKKEWVESCPTHVFDID---AN 715
Query: 318 RKKAVVKNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIESTGAIPPD 377
+K V A + E I++A+ + G V + +D FIF +E+TGA+
Sbjct: 716 TQKVYVHEAENYSYDDEVIKKAEAMGKPG-------LVEITAKQDSFIFIVETTGAVKAS 874
Query: 378 VLFTEAVKILEDKCERV 394
L A++IL+ K + V
Sbjct: 875 QLVLNAIEILKQKLDAV 925
>BP028509
Length = 551
Score = 29.3 bits (64), Expect = 1.2
Identities = 22/67 (32%), Positives = 29/67 (42%), Gaps = 10/67 (14%)
Frame = -3
Query: 108 AVANAFRRILIAEVPTMAIERV--------YIANNTSLVQDEVLSH--RLGLIPIDADPK 157
A NA+ RIL + PT I V NN L +E + RL L+P+ +
Sbjct: 339 AAMNAYTRILARKYPTFCINSVCXGYCKTDITVNNGFLTAEEGAASPVRLALLPLGSPSG 160
Query: 158 LFEYPDD 164
LF Y D
Sbjct: 159 LFYYKGD 139
>TC8617 similar to UP|Q9SSZ4 (Q9SSZ4) Asparaginyl endopeptidase, partial
(44%)
Length = 904
Score = 28.5 bits (62), Expect = 2.1
Identities = 15/40 (37%), Positives = 21/40 (52%), Gaps = 2/40 (5%)
Frame = -2
Query: 22 RGIPDYIMNLED--VPSTLPTHLELQKTRVFCNLDAPQHT 59
RG+P NL +P+ LP HL+ R C+L+ HT
Sbjct: 813 RGLPHQRYNLHSSHMPNYLPLHLQQWLHRCLCHLEEVLHT 694
>TC11813 similar to UP|Q8LJ68 (Q8LJ68) Protein phosphatase 2C-like protein,
partial (36%)
Length = 917
Score = 28.1 bits (61), Expect = 2.7
Identities = 24/63 (38%), Positives = 32/63 (50%), Gaps = 1/63 (1%)
Frame = +1
Query: 190 LTVKSDTLKWLPNGSELIAEGTKSAADPAPKTFTTF-NQKSLPKFSKDPAPYNLDVIVAK 248
L K L + NG EL +E + DPA + F + ++LPK SKD N+ VIV
Sbjct: 355 LARKRILLWYKKNGFELSSERGEGV-DPAAQAAAEFLSNRALPKGSKD----NITVIVVD 519
Query: 249 LGP 251
L P
Sbjct: 520 LKP 528
>TC14970 similar to UP|O24294 (O24294) Legumin (Minor small) precursor,
partial (51%)
Length = 1566
Score = 27.7 bits (60), Expect = 3.5
Identities = 23/67 (34%), Positives = 27/67 (39%)
Frame = +2
Query: 310 DIEDIGRGRKKAVVKNARACTLCRECIREADEGEEGGEGEKWTDRVSLRRVKDHFIFTIE 369
D E IGRG+KK KN + GE GEGE + LR+ FT
Sbjct: 725 DEESIGRGKKKRKKKNHAG---------QFHLGERAGEGEDAELAMVLRKTSAP*RFTRT 877
Query: 370 STGAIPP 376
ST P
Sbjct: 878 STALHAP 898
>TC15795 weakly similar to UP|Q8H1T9 (Q8H1T9) Phospholipase D delta isoform
1a , partial (9%)
Length = 593
Score = 27.7 bits (60), Expect = 3.5
Identities = 16/45 (35%), Positives = 25/45 (55%), Gaps = 5/45 (11%)
Frame = +1
Query: 128 RVYIANNTSLVQDEVLSHRLGLIPIDADPKLFEYPD-----DAGG 167
R Y + + SL+Q +L + +P+D+D K+ PD DAGG
Sbjct: 67 RKYASEDFSLLQGHLLKYP---VPVDSDGKISSLPDCENFPDAGG 192
>BP049099
Length = 483
Score = 27.3 bits (59), Expect = 4.6
Identities = 12/25 (48%), Positives = 14/25 (56%)
Frame = -3
Query: 216 DPAPKTFTTFNQKSLPKFSKDPAPY 240
D APK F FN PK+ +DP Y
Sbjct: 166 DKAPKKFYLFNSYFSPKYLEDPLYY 92
>TC11642 similar to UP|Q9FHL3 (Q9FHL3) Genomic DNA, chromosome 5, TAC
clone:K19M13, partial (48%)
Length = 1127
Score = 26.9 bits (58), Expect = 6.0
Identities = 18/76 (23%), Positives = 34/76 (44%)
Frame = +3
Query: 182 RCEKGQPRLTVKSDTLKWLPNGSELIAEGTKSAADPAPKTFTTFNQKSLPKFSKDPAPYN 241
R +KG LT DT KW + + + + +DP P + PK+ +P ++
Sbjct: 774 RNDKGWTGLTTTHDTSKWGNTTTNDREDISSTLSDPGP------IWDAEPKWDAEPTNWD 935
Query: 242 LDVIVAKLGPGQEIEL 257
++ + GP + E+
Sbjct: 936 VENPIELPGPSDDAEV 983
>TC14478 weakly similar to UP|Q94G86 (Q94G86) Beta-1,3-glucanase-like
protein, partial (83%)
Length = 1784
Score = 26.6 bits (57), Expect = 7.8
Identities = 12/27 (44%), Positives = 17/27 (62%)
Frame = +2
Query: 213 SAADPAPKTFTTFNQKSLPKFSKDPAP 239
S+A K+ TTF+++S P S DP P
Sbjct: 143 SSASTTAKSPTTFHRQSPPSTSSDPPP 223
>CB827026
Length = 542
Score = 26.6 bits (57), Expect = 7.8
Identities = 11/21 (52%), Positives = 15/21 (71%)
Frame = +3
Query: 78 RLDNFYQNFKVEVKRITDEEM 98
+LDNFYQN + VK DE++
Sbjct: 465 QLDNFYQNHRRYVKSRNDEQL 527
>BP064473
Length = 471
Score = 26.6 bits (57), Expect = 7.8
Identities = 13/26 (50%), Positives = 15/26 (57%)
Frame = -1
Query: 337 READEGEEGGEGEKWTDRVSLRRVKD 362
RE + EEGG GEK + RR KD
Sbjct: 468 RERSDDEEGGTGEKKKRKGGKRRKKD 391
>TC8341 similar to UP|Q94FX1 (Q94FX1) Heme oxygenase 1 (Fragment),
partial (86%)
Length = 1286
Score = 26.6 bits (57), Expect = 7.8
Identities = 14/40 (35%), Positives = 19/40 (47%), Gaps = 11/40 (27%)
Frame = +1
Query: 34 VPSTLPTHLELQKTRVFC-----------NLDAPQHTDTI 62
+PS L +HL+ KT + N PQHTDT+
Sbjct: 22 IPSILHSHLQRNKTPIIITNLTKKTPIALNKPLPQHTDTV 141
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.134 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,004,801
Number of Sequences: 28460
Number of extensions: 76819
Number of successful extensions: 418
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 412
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 415
length of query: 399
length of database: 4,897,600
effective HSP length: 92
effective length of query: 307
effective length of database: 2,279,280
effective search space: 699738960
effective search space used: 699738960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.5 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146751.11


