

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146750.9 + phase: 0
(214 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
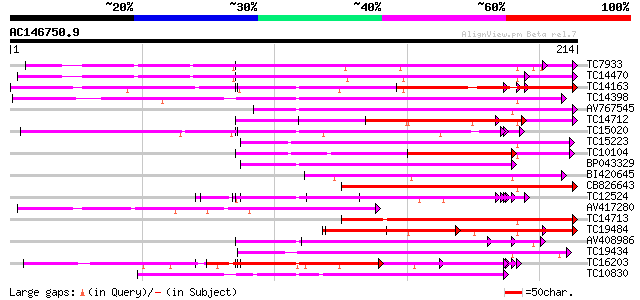
Sequences producing significant alignments: (bits) Value
TC7933 weakly similar to UP|AAQ56728 (AAQ56728) Polygalacturonas... 132 3e-32
TC14470 similar to UP|AAQ56728 (AAQ56728) Polygalacturonase inhi... 120 1e-28
TC14163 99 6e-22
TC14398 similar to UP|Q96477 (Q96477) LRR protein, partial (86%) 97 3e-21
AV767545 94 1e-20
TC14712 similar to UP|PGI2_PHAVU (P58822) Polygalacturonase inhi... 89 6e-19
TC15020 homologue to UP|Q9QX99 (Q9QX99) Transcription factor (T-... 87 3e-18
TC15223 similar to UP|CAE76632 (CAE76632) Leucine rich repeat pr... 84 2e-17
TC10104 similar to UP|Q8GTD6 (Q8GTD6) Polygalacturonase inhibito... 83 3e-17
BP043329 81 2e-16
BI420645 80 3e-16
CB826643 79 6e-16
TC12524 similar to UP|Q9LKZ6 (Q9LKZ6) Receptor-like protein kina... 79 8e-16
AV417280 77 3e-15
TC14713 similar to UP|PGI2_PHAVU (P58822) Polygalacturonase inhi... 76 4e-15
TC19484 similar to UP|BAC81651 (BAC81651) Polygalacturonase inhi... 75 9e-15
AV408986 74 1e-14
TC19434 similar to UP|Q94F63 (Q94F63) Somatic embryogenesis rece... 74 1e-14
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 74 2e-14
TC10830 weakly similar to UP|Q94JL9 (Q94JL9) At1g68400/T2E12_5, ... 72 7e-14
>TC7933 weakly similar to UP|AAQ56728 (AAQ56728) Polygalacturonase
inhibiting protein, partial (78%)
Length = 1320
Score = 132 bits (333), Expect = 3e-32
Identities = 84/204 (41%), Positives = 111/204 (54%), Gaps = 7/204 (3%)
Frame = +3
Query: 7 LTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVG 66
L +L S + FS C+ DK LL+I+ F P L WD T CCDW V
Sbjct: 36 LFMLLFSPVAFSGR-------CHPQDKKVLLQIKKDFNSPY-LLASWDPKTACCDWYCVE 191
Query: 67 CGRPYPGRVTVVTISRG---WGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ 123
C P R+T + IS LSG +P G+LPYL L ++PK+TGPI + +KL
Sbjct: 192 CD-PKTHRITSLIISSSIPDTNLSGPIPPSVGDLPYLEYLDFHKLPKLTGPIQPAIAKLT 368
Query: 124 RLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQL 183
+L++L + ++SGPIP FL +LK L + LS N LSG IPASL L +L N+ N+L
Sbjct: 369 KLKSLSITWTNVSGPIPDFLAQLKNLDSLYLSFNNLSGHIPASLSRLPNLLSLNLDRNKL 548
Query: 184 CGAIPAGLKKFNK---NVF-EHNK 203
G IPA F K N+F HN+
Sbjct: 549 TGPIPASFGFFKKPGPNLFLSHNQ 620
Score = 97.1 bits (240), Expect = 2e-21
Identities = 69/178 (38%), Positives = 86/178 (47%), Gaps = 49/178 (27%)
Frame = +3
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRL-QNLDLGSNSLSGPIPSFLG 144
LSG +PA LP L L+L + K+TGPIP SF ++ NL L N LSGPIPS LG
Sbjct: 474 LSGHIPASLSRLPNLLSLNL-DRNKLTGPIPASFGFFKKPGPNLFLSHNQLSGPIPSSLG 650
Query: 145 KL----------------------------------------------KRLKEVDLSNNK 158
+L K L +DL++NK
Sbjct: 651 QLDPERIDFSRNKLTGNASLLFAHNKKTQLLDLSRNLLSFDLSKVEFPKSLTWLDLNHNK 830
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
+ G+IP L +Q+L QFNVS+N L G IP G L KF+K + HNK LCG PL CK
Sbjct: 831 IHGSIPVGLSAVQNLQQFNVSYNLLSGKIPQGGELGKFDKFSYFHNKNLCGTPLPNCK 1004
>TC14470 similar to UP|AAQ56728 (AAQ56728) Polygalacturonase inhibiting
protein, partial (93%)
Length = 1195
Score = 120 bits (302), Expect = 1e-28
Identities = 76/196 (38%), Positives = 102/196 (51%), Gaps = 3/196 (1%)
Frame = +3
Query: 4 LLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWS 63
L L LL S + FS CN +DK LL+I+ P L W+ T CCDW
Sbjct: 15 LCFLLLLSFSHVAFSGR-------CNPEDKKVLLQIKKDLNNPY-LLASWNPETACCDWY 170
Query: 64 FVGCGRPYPGRVTVVTISRG---WGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFS 120
V C P R+T + IS SG +P G+LPYL +L ++PK+TGPI + +
Sbjct: 171 CVECD-PKTHRITSLIISSSVPETNFSGQIPPSVGDLPYLQVLEFHKLPKLTGPIQPAIA 347
Query: 121 KLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSF 180
KL L+ L L ++SGPIP FL +L L+ + LS N L+G IP+SL L +L +
Sbjct: 348 KLTNLRWLILTWTNISGPIPDFLSELPNLQWLHLSFNNLTGPIPSSLSKLPNLIDLRLDR 527
Query: 181 NQLCGAIPAGLKKFNK 196
N+L G IP F K
Sbjct: 528 NRLTGPIPDSFGSFKK 575
Score = 88.2 bits (217), Expect = 1e-18
Identities = 58/178 (32%), Positives = 85/178 (47%), Gaps = 49/178 (27%)
Frame = +3
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQ-NLDLGSNSLSGPIPSFLG 144
L+G +P+ LP L L L + ++TGPIP+SF ++ +L L N LSGPIP+ LG
Sbjct: 462 LTGPIPSSLSKLPNLIDLRL-DRNRLTGPIPDSFGSFKKPGIDLTLSHNQLSGPIPTSLG 638
Query: 145 KLK----------------------------------------------RLKEVDLSNNK 158
+ L +DL++N+
Sbjct: 639 LIDPNRIDFSRNMLEGDASMLFGVNKTTEMIDLSRNLLTFDMSKVELPTSLISLDLNHNQ 818
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
+ G+IP L + L FNVS+N+LCG IP G L++F + + HN+CLCG+PL CK
Sbjct: 819 IYGSIPQELIKVDFLQAFNVSYNRLCGEIPQGGRLQRFQVDTYFHNRCLCGSPLPSCK 992
>TC14163
Length = 1712
Score = 99.0 bits (245), Expect = 6e-22
Identities = 64/199 (32%), Positives = 97/199 (48%), Gaps = 6/199 (3%)
Frame = +1
Query: 1 MFLLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHF-GGPKGRLDDWDNNTEC 59
++L L L + ++ K+N + CN DD+A L+ + P G L W T+C
Sbjct: 115 LYLSKLLILFAIFTLLHLKANGAK---CNVDDEAGLMGFKSGIKSDPSGILKSWIPGTDC 285
Query: 60 CDWSFVGCGRPYPGRVTVVTISRGWG-----LSGTLPAEFGNLPYLSMLSLAEMPKVTGP 114
C W V C RVT + +S SGT+ + L L + ++GP
Sbjct: 286 CTWQGVTCLFD-DKRVTSLYLSGNPENPKSFFSGTISPSLSKIKNLDGFYLLNLKNISGP 462
Query: 115 IPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLS 174
P KL +LQ + + +N LSG IP +G L RL + L+ N+ +GTIP+S+G L L+
Sbjct: 463 FPGFLFKLPKLQFIYIENNQLSGRIPENIGNLTRLDVLSLTGNRFTGTIPSSVGGLTHLT 642
Query: 175 QFNVSFNQLCGAIPAGLKK 193
Q + N L G IPA + +
Sbjct: 643 QLQLGNNSLTGTIPATIAR 699
Score = 77.0 bits (188), Expect = 2e-15
Identities = 45/103 (43%), Positives = 61/103 (58%), Gaps = 1/103 (0%)
Frame = +1
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLGK 145
SG +P F + L +L L+ K +G IP S S L +L+ L+LG N LSG IP FLGK
Sbjct: 742 SGAIPDFFSSFTDLGILRLSRN-KFSGKIPASISTLAPKLRYLELGHNQLSGKIPDFLGK 918
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
+ L +DLS+N+ SGT+PAS NL + N++ N L P
Sbjct: 919 FRALDTLDLSSNRFSGTVPASFKNLTKIFNLNLANNLLVDPFP 1047
Score = 71.2 bits (173), Expect = 1e-13
Identities = 43/112 (38%), Positives = 63/112 (55%), Gaps = 1/112 (0%)
Frame = +1
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+GT+PA L L+ LSL E + +G IP+ FS L L L N SG IP+ +
Sbjct: 667 LTGTIPATIARLKNLTYLSL-EGNQFSGAIPDFFSSFTDLGILRLSRNKFSGKIPASIST 843
Query: 146 LK-RLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
L +L+ ++L +N+LSG IP LG ++L ++S N+ G +PA K K
Sbjct: 844 LAPKLRYLELGHNQLSGKIPDFLGKFRALDTLDLSSNRFSGTVPASFKNLTK 999
Score = 65.9 bits (159), Expect = 5e-12
Identities = 34/68 (50%), Positives = 43/68 (63%)
Frame = +1
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFEHNKCLC 206
+R K +DLS+N + G +P S+ L+ L NVS+N LCG IP KF + F N CLC
Sbjct: 1351 ERFKYLDLSHNLVFGKVPKSVAGLEKL---NVSYNHLCGEIPK--TKFPASAFVGNDCLC 1515
Query: 207 GAPLAPCK 214
GAPL PCK
Sbjct: 1516 GAPLQPCK 1539
>TC14398 similar to UP|Q96477 (Q96477) LRR protein, partial (86%)
Length = 982
Score = 96.7 bits (239), Expect = 3e-21
Identities = 73/212 (34%), Positives = 99/212 (46%), Gaps = 3/212 (1%)
Frame = +1
Query: 2 FLLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNN-TECC 60
F + ++L+ + F+ +N+ D AL + P L WD C
Sbjct: 142 FFFCCVVSIYLTLVPFAVANSEGD---------ALYAFKQSLSDPDNVLQSWDATLVSPC 294
Query: 61 DWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFS 120
W V C RV + ++ LSG L + GNL L L L E + G IP
Sbjct: 295 TWFHVTCQDNSVTRVDLGNLN----LSGHLVPDLGNLHSLQYLELYEN-NIQGTIPEELG 459
Query: 121 KLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSF 180
LQ L +LDL N++SG IPS LG LK L+ + L+NN L+G IP SL L +L +VS
Sbjct: 460 NLQSLISLDLYHNNVSGSIPSSLGNLKNLRFLRLNNNHLTGQIPKSLSTLPNLKVLDVSN 639
Query: 181 NQLCGAIPAG--LKKFNKNVFEHNKCLCGAPL 210
N LCG IP + + FE+N L G L
Sbjct: 640 NNLCGPIPTSGPFEHIPLDNFENNPRLEGPEL 735
>AV767545
Length = 562
Score = 94.4 bits (233), Expect = 1e-20
Identities = 53/124 (42%), Positives = 72/124 (57%), Gaps = 2/124 (1%)
Frame = -1
Query: 93 EFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEV 152
EF L +L+L ++GPIP S S L L+ LD+ N + G IPS LG+L L+ +
Sbjct: 556 EFAEGSSLKVLNLGSN-NISGPIPVSISNLIDLERLDISRNHILGAIPSSLGQLLELQWL 380
Query: 153 DLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP--AGLKKFNKNVFEHNKCLCGAPL 210
D+S N L+G+IP+SL + +L + N+LCG IP L F + HN CLCG PL
Sbjct: 379 DVSINSLAGSIPSSLSQITNLKHASFRANRLCGEIPQTRPLNIFPAAAYAHNLCLCGKPL 200
Query: 211 APCK 214
PCK
Sbjct: 199 QPCK 188
>TC14712 similar to UP|PGI2_PHAVU (P58822) Polygalacturonase inhibitor 2
precursor (Polygalacturonase-inhibiting protein)
(PGIP-2), partial (51%)
Length = 824
Score = 89.0 bits (219), Expect = 6e-19
Identities = 55/154 (35%), Positives = 74/154 (47%), Gaps = 25/154 (16%)
Frame = +1
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG +P +G+ L +++G IP S SKL L +DL N L G F G
Sbjct: 88 LSGAIPDSYGSFSNLFTSLTLNRNQLSGKIPASLSKLN-LAFVDLSMNMLEGDASVFFGS 264
Query: 146 LKR-----------------------LKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQ 182
K L +DL NN++ G +P L L+ L + NVS+N
Sbjct: 265 KKNTQKIILARNSLAFDLGKVGLSSNLNTIDLRNNRVYGKLPQELTGLKFLKKLNVSYNS 444
Query: 183 LCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
LCG IP G L++F+ + HNKCLCG+PL CK
Sbjct: 445 LCGQIPQGGNLQRFDVYSYAHNKCLCGSPLPACK 546
Score = 53.5 bits (127), Expect = 3e-08
Identities = 32/77 (41%), Positives = 44/77 (56%), Gaps = 1/77 (1%)
Frame = +1
Query: 110 KVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRL-KEVDLSNNKLSGTIPASLG 168
++TGP+P+S S L L + N LSG IP G L + L+ N+LSG IPASL
Sbjct: 13 RLTGPLPSSISTLPNLVGITANDNKLSGAIPDSYGSFSNLFTSLTLNRNQLSGKIPASLS 192
Query: 169 NLQSLSQFNVSFNQLCG 185
L +L+ ++S N L G
Sbjct: 193 KL-NLAFVDLSMNMLEG 240
Score = 51.6 bits (122), Expect = 1e-07
Identities = 27/62 (43%), Positives = 38/62 (60%), Gaps = 1/62 (1%)
Frame = +1
Query: 135 LSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSL-SQFNVSFNQLCGAIPAGLKK 193
L+GP+PS + L L + ++NKLSG IP S G+ +L + ++ NQL G IPA L K
Sbjct: 16 LTGPLPSSISTLPNLVGITANDNKLSGAIPDSYGSFSNLFTSLTLNRNQLSGKIPASLSK 195
Query: 194 FN 195
N
Sbjct: 196 LN 201
>TC15020 homologue to UP|Q9QX99 (Q9QX99) Transcription factor (T-cell
leukemia, homeobox 1), partial (5%)
Length = 1304
Score = 86.7 bits (213), Expect = 3e-18
Identities = 67/192 (34%), Positives = 94/192 (48%), Gaps = 8/192 (4%)
Frame = +1
Query: 5 LHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWS- 63
L LT++FL ++F A A KA I+ P L W+ + C
Sbjct: 1 LCLTIIFLPLLHFHHVQALTSASDIAALKAFKASIKPSSIAPWSCLASWNFTIDPCSLPR 180
Query: 64 ---FVGCGRPYPGRVTVVTISR----GWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIP 116
F+ CG T I++ G SG+L L L L LAE +GPIP
Sbjct: 181 RTHFI-CGLTCTATTTTNNINQITLDPAGYSGSLTPLISQLTQLVTLDLAEN-SFSGPIP 354
Query: 117 NSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQF 176
+S S L L+ L L SNS SG IP + LK L +D S+N LSG++P S+ +L +L +
Sbjct: 355 SSLSSLSNLKTLTLRSNSFSGSIPPSITTLKSLYSLDFSHNSLSGSLPNSMNSLINLRRI 534
Query: 177 NVSFNQLCGAIP 188
++SFN+L G+IP
Sbjct: 535 DLSFNKLTGSIP 570
Score = 62.4 bits (150), Expect = 6e-11
Identities = 39/101 (38%), Positives = 56/101 (54%)
Frame = +1
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
SG +P+ +L L L+L +G IP S + L+ L +LD NSLSG +P+ + L
Sbjct: 340 SGPIPSSLSSLSNLKTLTLRSN-SFSGSIPPSITTLKSLYSLDFSHNSLSGSLPNSMNSL 516
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
L+ +DLS NKL+G+IP NL L+ V N L G +
Sbjct: 517 INLRRIDLSFNKLTGSIPKLPPNLLELA---VKANSLSGPL 630
Score = 49.3 bits (116), Expect = 5e-07
Identities = 40/134 (29%), Positives = 63/134 (46%), Gaps = 25/134 (18%)
Frame = +1
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPN----------------------SFSKLQ 123
LSG+LP +L L + L+ K+TG IP +F L
Sbjct: 481 LSGSLPNSMNSLINLRRIDLS-FNKLTGSIPKLPPNLLELAVKANSLSGPLQKATFEGLN 657
Query: 124 RLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSG---TIPASLGNLQSLSQFNVSF 180
+L+ ++L N+L+G + + L L++VDLSNN +G + PA G +L N+ F
Sbjct: 658 QLEVVELSENTLAGTVEPWFFLLPSLQQVDLSNNSFTGVQISRPAR-GEGSNLVAVNLGF 834
Query: 181 NQLCGAIPAGLKKF 194
N++ G PA L +
Sbjct: 835 NRIQGYAPANLAAY 876
>TC15223 similar to UP|CAE76632 (CAE76632) Leucine rich repeat protein
precursor, partial (35%)
Length = 876
Score = 83.6 bits (205), Expect = 2e-17
Identities = 47/128 (36%), Positives = 67/128 (51%), Gaps = 2/128 (1%)
Frame = +3
Query: 88 GTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLK 147
G++PAEFG + LS L+L + ++G IP++ + L+L N L G IP G
Sbjct: 3 GSVPAEFGRMRVLSTLNL-DSNSLSGQIPSTLLSNSGMGILNLSRNGLEGTIPDVFGPNS 179
Query: 148 RLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCL 205
+DLS N L G +P SL + + + ++S N LCG+IP G + F N CL
Sbjct: 180 YFMALDLSYNGLQGRVPGSLCSAKYIGHLDLSHNHLCGSIPMGEPFDHLEASSFSSNDCL 359
Query: 206 CGAPLAPC 213
CG PL C
Sbjct: 360 CGNPLKTC 383
>TC10104 similar to UP|Q8GTD6 (Q8GTD6) Polygalacturonase inhibitor-like
protein (Fragment), partial (42%)
Length = 563
Score = 83.2 bits (204), Expect = 3e-17
Identities = 46/130 (35%), Positives = 70/130 (53%), Gaps = 2/130 (1%)
Frame = +2
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
+SG +P G + LS L+L +M +++G IP S + + +L++ ++L G IP G+
Sbjct: 26 VSGPIPESLGKMAVLSTLNL-DMNRLSGHIPKSLF-VSGISDLNISHHALEGAIPDAFGE 199
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNK 203
+DLS N L G IP S+ + + ++S N LCGAIP G + F +N
Sbjct: 200 RSYFTALDLSYNNLKGAIPKSISSAAYIGHLDLSHNHLCGAIPIGSPFDHLEASSFVYND 379
Query: 204 CLCGAPLAPC 213
CLCG PL C
Sbjct: 380 CLCGKPLKAC 409
Score = 42.7 bits (99), Expect = 5e-05
Identities = 20/41 (48%), Positives = 27/41 (65%)
Frame = +2
Query: 151 EVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
++DLS N++SG IP SLG + LS N+ N+L G IP L
Sbjct: 2 DLDLSRNQVSGPIPESLGKMAVLSTLNLDMNRLSGHIPKSL 124
>BP043329
Length = 484
Score = 80.9 bits (198), Expect = 2e-16
Identities = 44/104 (42%), Positives = 61/104 (58%)
Frame = +3
Query: 88 GTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLK 147
G +P E GNL L ++ L + + G IP+S L++L++LDL N L G IPS L L
Sbjct: 75 GRIPPEIGNLTNLEVVWLTQC-NLVGVIPDSIGNLKKLKDLDLALNDLYGSIPSSLTGLT 251
Query: 148 RLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
L++++L NN LSG +P +GNL L + S N L G IP L
Sbjct: 252 SLRQIELYNNSLSGELPRGMGNLTELRLLDASMNHLTGRIPEEL 383
Score = 37.4 bits (85), Expect = 0.002
Identities = 25/67 (37%), Positives = 34/67 (50%)
Frame = +3
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG LP GNL L +L A M +TG IP L L++L+L N G +P+ +
Sbjct: 285 LSGELPRGMGNLTELRLLD-ASMNHLTGRIPEELCSLP-LESLNLYENRFEGELPASIAD 458
Query: 146 LKRLKEV 152
L E+
Sbjct: 459 SPNLYEL 479
Score = 26.9 bits (58), Expect = 2.7
Identities = 14/28 (50%), Positives = 17/28 (60%), Gaps = 1/28 (3%)
Frame = +3
Query: 162 TIPASLGNLQSLSQFNVSFNQLC-GAIP 188
TIP SLG L +L N+S+N G IP
Sbjct: 3 TIPPSLGTLTTLKMLNLSYNPFYPGRIP 86
>BI420645
Length = 551
Score = 79.7 bits (195), Expect = 3e-16
Identities = 49/103 (47%), Positives = 58/103 (55%), Gaps = 4/103 (3%)
Frame = +3
Query: 112 TGPIPNSFSK-LQRLQNLDLGSNSLSGPIPSFLGKLKRLK-EVDLSNNKLSGTIPASLGN 169
TGP+P+ F L L+ LDL N SG IPS +GKL L+ VDLS+N SG IPASLGN
Sbjct: 6 TGPLPDGFGGGLSLLEKLDLSFNQFSGSIPSDMGKLSSLQGNVDLSHNHFSGLIPASLGN 185
Query: 170 LQSLSQFNVSFNQLCGAIP--AGLKKFNKNVFEHNKCLCGAPL 210
L ++S+N L G IP L F N LCG PL
Sbjct: 186 LPEKVYIDLSYNNLSGPIPQTGALMNRGPTAFIGNSGLCGPPL 314
Score = 34.3 bits (77), Expect = 0.017
Identities = 18/52 (34%), Positives = 29/52 (55%)
Frame = +3
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGP 138
SG +PA GNLP + L+ ++GPIP + + + R +G++ L GP
Sbjct: 156 SGLIPASLGNLPEKVYIDLS-YNNLSGPIPQTGALMNRGPTAFIGNSGLCGP 308
>CB826643
Length = 346
Score = 79.0 bits (193), Expect = 6e-16
Identities = 43/91 (47%), Positives = 55/91 (60%), Gaps = 2/91 (2%)
Frame = +3
Query: 126 QNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCG 185
Q LDL N S K L +DL++N + G+IPA L +++L QFNVS+NQLCG
Sbjct: 3 QILDLSRNLFSFDFSKVGFPKKSLIWLDLNHNNIFGSIPAGLTTVENLQQFNVSYNQLCG 182
Query: 186 AIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
IP G L +F+K + HNKCLC PL CK
Sbjct: 183 KIPQGGELGRFDKYSYLHNKCLCDPPLPNCK 275
>TC12524 similar to UP|Q9LKZ6 (Q9LKZ6) Receptor-like protein kinase 1,
partial (27%)
Length = 834
Score = 78.6 bits (192), Expect = 8e-16
Identities = 47/101 (46%), Positives = 59/101 (57%)
Frame = +2
Query: 88 GTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLK 147
G +P E GNL L A ++G IP KLQ+L L L N LSG + LG LK
Sbjct: 35 GGIPPEIGNLTQLLRFDAAYCG-LSGEIPAELGKLQKLDTLFLQVNVLSGSLTPELGHLK 211
Query: 148 RLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
LK +DLSNN LSG +PAS L++L+ N+ N+L GAIP
Sbjct: 212 SLKSMDLSNNMLSGQVPASFAELKNLTLLNLFRNRLHGAIP 334
Score = 70.5 bits (171), Expect = 2e-13
Identities = 40/112 (35%), Positives = 62/112 (54%)
Frame = +2
Query: 85 GLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLG 144
GLSG +PAE G L L L L ++ ++G + L+ L+++DL +N LSG +P+
Sbjct: 98 GLSGEIPAELGKLQKLDTLFL-QVNVLSGSLTPELGHLKSLKSMDLSNNMLSGQVPASFA 274
Query: 145 KLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
+LK L ++L N+L G IP +G + +L + N G+IP L K K
Sbjct: 275 ELKNLTLLNLFRNRLHGAIPEFVGEMPALEVLQLWENNFTGSIPQSLGKNGK 430
Score = 70.1 bits (170), Expect = 3e-13
Identities = 40/111 (36%), Positives = 65/111 (58%)
Frame = +2
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG+L E G+L L + L+ ++G +P SF++L+ L L+L N L G IP F+G+
Sbjct: 173 LSGSLTPELGHLKSLKSMDLSNN-MLSGQVPASFAELKNLTLLNLFRNRLHGAIPEFVGE 349
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
+ L+ + L N +G+IP SLG L+ ++S N+L G +P + N+
Sbjct: 350 MPALEVLQLWENNFTGSIPQSLGKNGKLTLVDLSSNKLTGTLPPHMCSGNR 502
Score = 68.2 bits (165), Expect = 1e-12
Identities = 47/130 (36%), Positives = 66/130 (50%), Gaps = 24/130 (18%)
Frame = +2
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG +PA F L L++L+L ++ G IP ++ L+ L L N+ +G IP LGK
Sbjct: 245 LSGQVPASFAELKNLTLLNLFRN-RLHGAIPEFVGEMPALEVLQLWENNFTGSIPQSLGK 421
Query: 146 LKRLKEVDLSNNKLSGT------------------------IPASLGNLQSLSQFNVSFN 181
+L VDLS+NKL+GT IP SLG +SL++ + N
Sbjct: 422 NGKLTLVDLSSNKLTGTLPPHMCSGNRLQTLIALGNFLFGPIPESLGKCESLTRIRMGQN 601
Query: 182 QLCGAIPAGL 191
L G+IP GL
Sbjct: 602 FLNGSIPKGL 631
Score = 60.5 bits (145), Expect = 2e-10
Identities = 38/120 (31%), Positives = 58/120 (47%), Gaps = 2/120 (1%)
Frame = +2
Query: 71 YPGRVTVVTISRGW--GLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNL 128
+ G + + + + W +G++P G L+++ L+ K+TG +P RLQ L
Sbjct: 338 FVGEMPALEVLQLWENNFTGSIPQSLGKNGKLTLVDLSSN-KLTGTLPPHMCSGNRLQTL 514
Query: 129 DLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
N L GPIP LGK + L + + N L+G+IP L L L+Q N L G P
Sbjct: 515 IALGNFLFGPIPESLGKCESLTRIRMGQNFLNGSIPKGLFGLPKLTQVEFQDNLLSGEFP 694
Score = 57.8 bits (138), Expect = 1e-09
Identities = 32/77 (41%), Positives = 44/77 (56%)
Frame = +2
Query: 113 GPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQS 172
G IP L +L D LSG IP+ LGKL++L + L N LSG++ LG+L+S
Sbjct: 35 GGIPPEIGNLTQLLRFDAAYCGLSGEIPAELGKLQKLDTLFLQVNVLSGSLTPELGHLKS 214
Query: 173 LSQFNVSFNQLCGAIPA 189
L ++S N L G +PA
Sbjct: 215 LKSMDLSNNMLSGQVPA 265
Score = 57.0 bits (136), Expect = 2e-09
Identities = 43/137 (31%), Positives = 66/137 (47%), Gaps = 24/137 (17%)
Frame = +2
Query: 73 GRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGS 132
G++T+V +S L+GTLP + L L +A + GPIP S K + L + +G
Sbjct: 425 GKLTLVDLSSN-KLTGTLPPHMCSGNRLQTL-IALGNFLFGPIPESLGKCESLTRIRMGQ 598
Query: 133 NSLSGPIPSFLGKLKRLKEVD------------------------LSNNKLSGTIPASLG 168
N L+G IP L L +L +V+ LSNNKLSG +P+++G
Sbjct: 599 NFLNGSIPKGLFGLPKLTQVEFQDNLLSGEFPETGSVSHNIGQITLSNNKLSGPLPSTIG 778
Query: 169 NLQSLSQFNVSFNQLCG 185
N S+ + + N+ G
Sbjct: 779 NFTSMQKLLLDGNKFSG 829
Score = 40.0 bits (92), Expect = 3e-04
Identities = 22/55 (40%), Positives = 29/55 (52%)
Frame = +2
Query: 133 NSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
N+ G IP +G L +L D + LSG IPA LG LQ L + N L G++
Sbjct: 23 NNYQGGIPPEIGNLTQLLRFDAAYCGLSGEIPAELGKLQKLDTLFLQVNVLSGSL 187
Score = 34.7 bits (78), Expect = 0.013
Identities = 17/40 (42%), Positives = 22/40 (54%)
Frame = +2
Query: 157 NKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
N G IP +GNL L +F+ ++ L G IPA L K K
Sbjct: 23 NNYQGGIPPEIGNLTQLLRFDAAYCGLSGEIPAELGKLQK 142
>AV417280
Length = 427
Score = 76.6 bits (187), Expect = 3e-15
Identities = 49/142 (34%), Positives = 75/142 (52%), Gaps = 5/142 (3%)
Frame = +3
Query: 4 LLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCD-- 61
+L L LL + +F+ S + CN DK ALL+ + G P +L W+ T+CCD
Sbjct: 12 VLLLLLLLTVTCFFTPSLSEK---CNPQDKTALLQFKKELGNP-AKLSSWNATTDCCDPA 179
Query: 62 WSFVGCGRPYPG-RVTVVTISRGWGLSGT--LPAEFGNLPYLSMLSLAEMPKVTGPIPNS 118
W V C RV + +S G+ L +P G+LP+L++LSL +P + GPIP++
Sbjct: 180 WEGVSCDTDTKTYRVNDLDLS-GFSLPSPHPIPPSVGDLPHLNILSLRNIPNLIGPIPSA 356
Query: 119 FSKLQRLQNLDLGSNSLSGPIP 140
+KL L + + +SG IP
Sbjct: 357 ITKLTSLHYIYISQTGISGNIP 422
>TC14713 similar to UP|PGI2_PHAVU (P58822) Polygalacturonase inhibitor 2
precursor (Polygalacturonase-inhibiting protein)
(PGIP-2), partial (26%)
Length = 533
Score = 76.3 bits (186), Expect = 4e-15
Identities = 39/91 (42%), Positives = 56/91 (60%), Gaps = 2/91 (2%)
Frame = +2
Query: 126 QNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCG 185
Q + L NSL+ + +G L +DL N+++ GT+P L L+ L + NVS+N LCG
Sbjct: 2 QKIILARNSLAFDLGK-VGLSSNLNTIDLRNHRVYGTLPQELTGLKFLKKLNVSYNSLCG 178
Query: 186 AIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
IP G L++F+ + HNKCLCG+PL CK
Sbjct: 179 QIPQGGNLQRFDVYSYAHNKCLCGSPLPACK 271
>TC19484 similar to UP|BAC81651 (BAC81651) Polygalacturonase inhibiting
protein (Fragment), partial (89%)
Length = 595
Score = 75.1 bits (183), Expect = 9e-15
Identities = 42/98 (42%), Positives = 60/98 (60%), Gaps = 2/98 (2%)
Frame = +3
Query: 119 FSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNV 178
F ++++ LDL N LS + K L +DL++N + G+IP L +++L QFNV
Sbjct: 213 FGPNKKIRILDLSKNLLSFDFSNVEIPTKSLIWLDLNHNMIHGSIPIGLTAVENLQQFNV 392
Query: 179 SFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
S+NQL G IP G L +F+K + HNK LCG+PL CK
Sbjct: 393 SYNQLSGKIPKGGELGRFDKYSYLHNKDLCGSPLPMCK 506
Score = 49.3 bits (116), Expect = 5e-07
Identities = 28/52 (53%), Positives = 36/52 (68%), Gaps = 1/52 (1%)
Frame = +1
Query: 120 SKLQRLQNLDLGSNSLSGPIPSFLGKLKRL-KEVDLSNNKLSGTIPASLGNL 170
SK L +L L N L+GPIPS LG L++ ++ LS+N+LSG IPASLG L
Sbjct: 4 SK*LNLSSLHLDQNQLTGPIPSSLGSLQKPGPDIVLSHNQLSGPIPASLGQL 159
Score = 38.9 bits (89), Expect = 7e-04
Identities = 23/62 (37%), Positives = 35/62 (56%), Gaps = 1/62 (1%)
Frame = +1
Query: 143 LGKLKRLKEVDLSNNKLSGTIPASLGNLQSLS-QFNVSFNQLCGAIPAGLKKFNKNVFEH 201
L K L + L N+L+G IP+SLG+LQ +S NQL G IPA L + + + +
Sbjct: 1 LSK*LNLSSLHLDQNQLTGPIPSSLGSLQKPGPDIVLSHNQLSGPIPASLGQLDPDRIDF 180
Query: 202 NK 203
++
Sbjct: 181 SR 186
>AV408986
Length = 434
Score = 74.3 bits (181), Expect = 1e-14
Identities = 43/106 (40%), Positives = 59/106 (55%)
Frame = +3
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG +P E GN+ L +L+L +G IP KL L+ L + +N L+G IP+ LG
Sbjct: 3 LSGEIPPEIGNISSLELLAL-HQNSFSGAIPKELGKLSGLKRLYVYTNQLNGTIPTELGN 179
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
E+DLS N+L G IP LG + +LS ++ N L G IP L
Sbjct: 180 CTNAIEIDLSENRLIGIIPKELGQISNLSLLHLFENNLQGHIPREL 317
Score = 65.5 bits (158), Expect = 7e-12
Identities = 37/97 (38%), Positives = 54/97 (55%)
Frame = +3
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+GT+P E GN + L+E ++ G IP ++ L L L N+L G IP LG
Sbjct: 147 LNGTIPTELGNCTNAIEIDLSEN-RLIGIIPKELGQISNLSLLHLFENNLQGHIPRELGS 323
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQ 182
L++LK++DLS N L+GTIP NL + + N+
Sbjct: 324 LRQLKKLDLSLNNLTGTIPLEFQNLTYIEDLQLFDNK 434
Score = 65.1 bits (157), Expect = 9e-12
Identities = 37/96 (38%), Positives = 55/96 (56%), Gaps = 4/96 (4%)
Frame = +3
Query: 111 VTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNL 170
++G IP + L+ L L NS SG IP LGKL LK + + N+L+GTIP LGN
Sbjct: 3 LSGEIPPEIGNISSLELLALHQNSFSGAIPKELGKLSGLKRLYVYTNQLNGTIPTELGNC 182
Query: 171 QSLSQFNVSFNQLCGAIPAGLKKFNK----NVFEHN 202
+ + ++S N+L G IP L + + ++FE+N
Sbjct: 183 TNAIEIDLSENRLIGIIPKELGQISNLSLLHLFENN 290
>TC19434 similar to UP|Q94F63 (Q94F63) Somatic embryogenesis receptor-like
kinase 2, partial (26%)
Length = 550
Score = 74.3 bits (181), Expect = 1e-14
Identities = 50/134 (37%), Positives = 71/134 (52%), Gaps = 8/134 (5%)
Frame = +2
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
+G + NL YL + S +TG IP+ L L +LDL N+L+G IPS LG L
Sbjct: 20 AGPQLGQLSNLQYLELYS----NNMTGKIPDELGNLTNLVSLDLYLNNLTGNIPSTLGNL 187
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP--AGLKKFNKNVFEHNKC 204
+L+ + L+NN LSG IP +L N+ SL ++S NQL G +P F +++N
Sbjct: 188 GKLRFLRLNNNTLSGGIPMTLTNVSSLQVLDLSNNQLKGVVPVNGSFSLFTPISYQNNPG 367
Query: 205 LC------GAPLAP 212
L APL+P
Sbjct: 368 LIQPKNPPSAPLSP 409
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation
aberrant root formation protein), complete
Length = 3308
Score = 73.9 bits (180), Expect = 2e-14
Identities = 65/219 (29%), Positives = 98/219 (44%), Gaps = 31/219 (14%)
Frame = +2
Query: 6 HLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGR---LDDWDNNTEC--- 59
+L +L + I+F + F D ALLK+++ G K + L+DW +T
Sbjct: 128 YLLVLCFTLIWFRWTVVYSSF----SDLDALLKLKESMKGAKAKHHALEDWKFSTSLSAH 295
Query: 60 CDWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSF 119
C +S V C + RV + ++ L G LP E G L L L+++ M +T +P+
Sbjct: 296 CSFSGVTCDQNL--RVVALNVTLV-PLFGHLPPEIGLLEKLENLTIS-MNNLTDQLPSDL 463
Query: 120 SKLQ-------------------------RLQNLDLGSNSLSGPIPSFLGKLKRLKEVDL 154
+ L L+ LD NS SGP+P + KL++LK + L
Sbjct: 464 ASLTSLKVLNISHNLFSGQFPGNITVGMTELEALDAYDNSFSGPLPEEIVKLEKLKYLHL 643
Query: 155 SNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKK 193
+ N SGTIP S QSL ++ N L G +P L K
Sbjct: 644 AGNYFSGTIPESYSEFQSLEFLGLNANSLTGRVPESLAK 760
Score = 68.9 bits (167), Expect = 6e-13
Identities = 40/108 (37%), Positives = 59/108 (54%)
Frame = +2
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+GT+P E ++ L L L+ + +TG IP SFSKL+ L ++ N G +PSF+G
Sbjct: 947 LTGTIPPELSSMMSLMSLDLS-INDLTGEIPESFSKLKNLTLMNFFQNKFRGSLPSFIGD 1123
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKK 193
L L+ + + N S +P +LG F+V+ N L G IP L K
Sbjct: 1124 LPNLETLQVWENNFSFVLPHNLGGNGRFLYFDVTKNHLTGLIPPDLCK 1267
Score = 68.9 bits (167), Expect = 6e-13
Identities = 53/152 (34%), Positives = 68/152 (43%), Gaps = 31/152 (20%)
Frame = +2
Query: 71 YPGRVTV------VTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQR 124
+PG +TV + SG LP E L L L LA +G IP S+S+ Q
Sbjct: 521 FPGNITVGMTELEALDAYDNSFSGPLPEEIVKLEKLKYLHLAGN-YFSGTIPESYSEFQS 697
Query: 125 LQNLDLGSNSLSGPIPSFLGKLKRLKE-------------------------VDLSNNKL 159
L+ L L +NSL+G +P L KLK LKE ++++N L
Sbjct: 698 LEFLGLNANSLTGRVPESLAKLKTLKELHLGYSNAYEGGIPPAFGSMENLRLLEMANCNL 877
Query: 160 SGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
+G IP SLGNL L V N L G IP L
Sbjct: 878 TGEIPPSLGNLTKLHSLFVQMNNLTGTIPPEL 973
Score = 68.6 bits (166), Expect = 8e-13
Identities = 37/102 (36%), Positives = 57/102 (55%)
Frame = +2
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
+G +PA NL L LSL + + G IP ++ L +++ N+L+GPIP+ +
Sbjct: 1523 TGKIPAAMKNLRALQSLSL-DANEFIGEIPGGVFEIPMLTKVNISGNNLTGPIPTTITHR 1699
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
L VDLS N L+G +P + NL LS N+S N++ G +P
Sbjct: 1700 ASLTAVDLSRNNLAGEVPKGMKNLMDLSILNLSRNEISGPVP 1825
Score = 67.4 bits (163), Expect = 2e-12
Identities = 38/106 (35%), Positives = 54/106 (50%)
Frame = +2
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+G +P GNL L L +M +TG IP S + L +LDL N L+G IP K
Sbjct: 875 LTGEIPPSLGNLTKLHSL-FVQMNNLTGTIPPELSSMMSLMSLDLSINDLTGEIPESFSK 1051
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
LK L ++ NK G++P+ +G+L +L V N +P L
Sbjct: 1052LKNLTLMNFFQNKFRGSLPSFIGDLPNLETLQVWENNFSFVLPHNL 1189
Score = 55.1 bits (131), Expect = 9e-09
Identities = 32/104 (30%), Positives = 56/104 (53%)
Frame = +2
Query: 88 GTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLK 147
G +P G L+ + +A + GP+P +L + +L +N L+G +PS + +
Sbjct: 1313 GPIPKGIGECRSLTKIRVANN-FLDGPVPPGVFQLPSVTITELSNNRLNGELPSVISG-E 1486
Query: 148 RLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
L + LSNN +G IPA++ NL++L ++ N+ G IP G+
Sbjct: 1487 SLGTLTLSNNLFTGKIPAAMKNLRALQSLSLDANEFIGEIPGGV 1618
Score = 52.4 bits (124), Expect = 6e-08
Identities = 36/125 (28%), Positives = 59/125 (46%), Gaps = 23/125 (18%)
Frame = +2
Query: 88 GTLPAEFGNLPYLSMLSLAE------MPK-----------------VTGPIPNSFSKLQR 124
G+LP+ G+LP L L + E +P +TG IP K R
Sbjct: 1097 GSLPSFIGDLPNLETLQVWENNFSFVLPHNLGGNGRFLYFDVTKNHLTGLIPPDLCKSGR 1276
Query: 125 LQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLC 184
L+ + N GPIP +G+ + L ++ ++NN L G +P + L S++ +S N+L
Sbjct: 1277 LKTFIITDNFFRGPIPKGIGECRSLTKIRVANNFLDGPVPPGVFQLPSVTITELSNNRLN 1456
Query: 185 GAIPA 189
G +P+
Sbjct: 1457 GELPS 1471
Score = 49.3 bits (116), Expect = 5e-07
Identities = 29/90 (32%), Positives = 47/90 (52%)
Frame = +2
Query: 75 VTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNS 134
+T V IS G L+G +P + L+ + L+ + G +P L L L+L N
Sbjct: 1634 LTKVNIS-GNNLTGPIPTTITHRASLTAVDLSRN-NLAGEVPKGMKNLMDLSILNLSRNE 1807
Query: 135 LSGPIPSFLGKLKRLKEVDLSNNKLSGTIP 164
+SGP+P + + L +DLS+N +GT+P
Sbjct: 1808 ISGPVPDEIRFMTSLTTLDLSSNNFTGTVP 1897
Score = 43.5 bits (101), Expect = 3e-05
Identities = 24/67 (35%), Positives = 41/67 (60%)
Frame = +2
Query: 75 VTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNS 134
+T V +SR L+G +P NL LS+L+L+ +++GP+P+ + L LDL SN+
Sbjct: 1706 LTAVDLSRN-NLAGEVPKGMKNLMDLSILNLSRN-EISGPVPDEIRFMTSLTTLDLSSNN 1879
Query: 135 LSGPIPS 141
+G +P+
Sbjct: 1880 FTGTVPT 1900
>TC10830 weakly similar to UP|Q94JL9 (Q94JL9) At1g68400/T2E12_5, partial
(24%)
Length = 702
Score = 72.0 bits (175), Expect = 7e-14
Identities = 52/140 (37%), Positives = 70/140 (49%)
Frame = +2
Query: 49 RLDDWDNNTECCDWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEM 108
+L W++ T C W V C R R+ + + L G+L +L L +LSL
Sbjct: 254 KLTTWNSTTHLCTWHGVTCLRNRVSRLVLENLD----LHGSLHP-LTSLTQLRVLSLKNN 418
Query: 109 PKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLG 168
+ GPIPN S L ++ L L N+ SG P L L RL +DLS+N LSG IPA++
Sbjct: 419 -RFHGPIPN-LSNLTGMRLLFLSHNNFSGDFPVTLTSLTRLYRLDLSHNSLSGEIPAAVN 592
Query: 169 NLQSLSQFNVSFNQLCGAIP 188
N L + NQL G IP
Sbjct: 593 NFTRLLTLRLDGNQLHGRIP 652
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.139 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,176,824
Number of Sequences: 28460
Number of extensions: 57991
Number of successful extensions: 533
Number of sequences better than 10.0: 108
Number of HSP's better than 10.0 without gapping: 360
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 447
length of query: 214
length of database: 4,897,600
effective HSP length: 86
effective length of query: 128
effective length of database: 2,450,040
effective search space: 313605120
effective search space used: 313605120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 53 (25.0 bits)
Medicago: description of AC146750.9


