

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146747.6 + phase: 0
(498 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
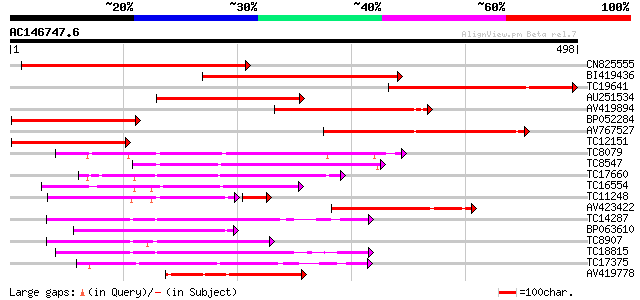
Score E
Sequences producing significant alignments: (bits) Value
CN825555 342 1e-94
BI419436 268 2e-72
TC19641 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {Ar... 236 5e-63
AU251534 236 5e-63
AV419894 226 5e-60
BP052284 222 1e-58
AV767527 217 3e-57
TC12151 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {Ar... 186 1e-47
TC8079 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, complete 168 2e-42
TC8547 homologue to UP|MMK1_MEDSA (Q07176) Mitogen-activated pro... 147 5e-36
TC17660 homologue to UP|Q9M6R8 (Q9M6R8) MAP kinase PsMAPK2, part... 144 3e-35
TC16554 homologue to UP|AAN71719 (AAN71719) Glycogen synthase ki... 140 4e-34
TC11248 homologue to UP|P93323 (P93323) Cdc2MsF protein, partial... 122 5e-34
AV423422 139 1e-33
TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein k... 123 8e-29
BP063610 121 2e-28
TC8907 similar to UP|Q8H0X3 (Q8H0X3) Serine/threonine protein ki... 118 2e-27
TC18815 similar to UP|Q8W1D5 (Q8W1D5) CBL-interacting protein ki... 112 1e-25
TC17375 mitogen-activated kinase kinase kinase alpha [Lotus corn... 112 1e-25
AV419778 102 1e-22
>CN825555
Length = 714
Score = 342 bits (876), Expect = 1e-94
Identities = 165/201 (82%), Positives = 182/201 (90%)
Frame = +1
Query: 11 GWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDN 70
GWPSWL VAGEAI TPRRA++FEKL KIGQGTYSNVYKA+D +TGKIVALKKVRFDN
Sbjct: 112 GWPSWLSKVAGEAINGLTPRRADTFEKLDKIGQGTYSNVYKARDTLTGKIVALKKVRFDN 291
Query: 71 LEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK 130
LEPESVKFMAREIL+LR+LDHPNV+KLEGLVTSRMSCSLYLVFEYM HDLAGL+ +K
Sbjct: 292 LEPESVKFMAREILILRRLDHPNVLKLEGLVTSRMSCSLYLVFEYMVHDLAGLATNPAIK 471
Query: 131 FTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQ 190
FTE QVKC+M QL +GLEHCH+R VLHRDIKGSNLLIDNEG+LKIADFGLA+F++P+ K
Sbjct: 472 FTESQVKCYMHQLFTGLEHCHNRHVLHRDIKGSNLLIDNEGVLKIADFGLASFFDPDHKH 651
Query: 191 SMTSRVVTLWYRPPELLLGAT 211
MTSRVVTLWYRPPELLLGAT
Sbjct: 652 PMTSRVVTLWYRPPELLLGAT 714
>BI419436
Length = 601
Score = 268 bits (684), Expect = 2e-72
Identities = 119/176 (67%), Positives = 148/176 (83%)
Frame = +1
Query: 170 EGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAEL 229
EG+LK+ADFGLA F + KQ +TSRVVTLWYRPPELLLGAT YG +DLWS GC+ AEL
Sbjct: 13 EGVLKVADFGLANFTSSGHKQPLTSRVVTLWYRPPELLLGATDYGPSVDLWSVGCVFAEL 192
Query: 230 LAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFP 289
L GKP++ GRTEVEQLHKIFKLCGSP E+YW+K +LP+AT+FKPQ PY C+ E+FKD P
Sbjct: 193 LVGKPVLQGRTEVEQLHKIFKLCGSPPEDYWKKTRLPHATLFKPQHPYDSCLRESFKDLP 372
Query: 290 PSSLPLIDSLLAIDPDRRGTASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMR 345
P+S+ L+ +LL+++P +RGTA++AL+ E+F T+PYACEPSSLP YPPSKE+D K R
Sbjct: 373 PASVNLLQTLLSVEPYKRGTATSALSSEYFRTKPYACEPSSLPTYPPSKEIDAKHR 540
>TC19641 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {Arabidopsis
thaliana;}, partial (15%)
Length = 823
Score = 236 bits (603), Expect = 5e-63
Identities = 122/166 (73%), Positives = 134/166 (80%)
Frame = +3
Query: 333 KYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRARERGRAIPAPEANAEIQTNLDR 392
KYPPSKELDVK+RDE ARR+KAL+GK NAVDG KRVR RERG AIPAPEANAEIQTNLDR
Sbjct: 3 KYPPSKELDVKLRDEAARRRKALSGKNNAVDGTKRVRTRERGCAIPAPEANAEIQTNLDR 182
Query: 393 WRVVTHANAKSKSEKFPPPHQDGAVGYPQDESSKGPVSFGASDTSFSSGTFNVKPSGPTR 452
RVVT ANAKSKSEKFPPPHQDGAVGYP D S+KG VSFGA++TSFSS N KPSG
Sbjct: 183 MRVVTRANAKSKSEKFPPPHQDGAVGYPLDASNKGAVSFGATETSFSSTISNSKPSGSVG 362
Query: 453 SHDGTGLHKGTKTKKEESQMASSWKFMRPFKPSTIGLSMDLLFRSK 498
S+ + ++G KT K S A SWK MR FKPST+GLS +LLFR K
Sbjct: 363 SY-ASSSYRGWKTNKAGSHSAPSWKLMRAFKPSTVGLSFNLLFRRK 497
>AU251534
Length = 391
Score = 236 bits (603), Expect = 5e-63
Identities = 110/130 (84%), Positives = 121/130 (92%)
Frame = +2
Query: 130 KFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKK 189
+FTEPQVKC+M+QLLSGLEHCH R VLHRDIKGSNLLIDNEG+LKIADFGLA+ ++PN K
Sbjct: 2 RFTEPQVKCYMRQLLSGLEHCHHRRVLHRDIKGSNLLIDNEGVLKIADFGLASVFDPNHK 181
Query: 190 QSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIF 249
MTSRVVTLWYRPPELLLGAT Y VG+DLWSAGCIL ELLAGKPIMPGRTEVEQLHKI+
Sbjct: 182 HPMTSRVVTLWYRPPELLLGATDYDVGVDLWSAGCILGELLAGKPIMPGRTEVEQLHKIY 361
Query: 250 KLCGSPAEEY 259
KLCGSP++EY
Sbjct: 362 KLCGSPSDEY 391
>AV419894
Length = 413
Score = 226 bits (577), Expect = 5e-60
Identities = 110/139 (79%), Positives = 125/139 (89%)
Frame = +1
Query: 233 KPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSS 292
+PIMPGRTEVEQLHKIFKLCGSP+EEYW+K KLP+ATIFKPQQ YKRCI+ETFK+FPPSS
Sbjct: 1 RPIMPGRTEVEQLHKIFKLCGSPSEEYWKKSKLPHATIFKPQQSYKRCIAETFKNFPPSS 180
Query: 293 LPLIDSLLAIDPDRRGTASAALNHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQ 352
LPLI++LL IDPD R TA+AAL+ EFFTT+PYACEPSSLPKYPPSKE+D K+RDEEARR
Sbjct: 181 LPLIETLLTIDPDERLTATAALHSEFFTTKPYACEPSSLPKYPPSKEMDAKLRDEEARRL 360
Query: 353 KALNGKANAVDGAKRVRAR 371
+A GKANA DG K+ R R
Sbjct: 361 RAA-GKANA-DGVKKSRPR 411
>BP052284
Length = 550
Score = 222 bits (565), Expect = 1e-58
Identities = 108/114 (94%), Positives = 113/114 (98%)
Frame = +1
Query: 2 LDPLSVNQQGWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIV 61
LDPL+V QQGWP+WLMAVAGEAIGDWTPRRA+SFEKLAKIGQGTYSNVYKAKDL+TGKIV
Sbjct: 208 LDPLAVTQQGWPAWLMAVAGEAIGDWTPRRASSFEKLAKIGQGTYSNVYKAKDLLTGKIV 387
Query: 62 ALKKVRFDNLEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEY 115
ALKKVRFDNLEPESVKFMAREILVLR+LDHPNVVKLEGLVTSRMSCSLYLVFEY
Sbjct: 388 ALKKVRFDNLEPESVKFMAREILVLRRLDHPNVVKLEGLVTSRMSCSLYLVFEY 549
>AV767527
Length = 573
Score = 217 bits (553), Expect = 3e-57
Identities = 113/182 (62%), Positives = 134/182 (73%), Gaps = 1/182 (0%)
Frame = +3
Query: 276 PYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFFTTEPYACEPSSLPKYP 335
PYKRCI E FKDFPPS+LPLID+LLAIDP R TAS AL EFFTTEPYAC+PSSLPKYP
Sbjct: 3 PYKRCIREIFKDFPPSALPLIDTLLAIDPVERRTASDALRSEFFTTEPYACDPSSLPKYP 182
Query: 336 PSKELDVKMRDEEARRQKALNGKANAVDGAKRVRARERG-RAIPAPEANAEIQTNLDRWR 394
PSKE+D K RD+E RRQ+A GKA A K R R+R +A APEANAE+Q+N+DR R
Sbjct: 183 PSKEMDAKRRDDEMRRQRAA-GKAQADGSKKHHRTRDRAVKAFAAPEANAELQSNIDRRR 359
Query: 395 VVTHANAKSKSEKFPPPHQDGAVGYPQDESSKGPVSFGASDTSFSSGTFNVKPSGPTRSH 454
++THANAKSKSEKFPPPHQDG +G+P S +D SF+S ++N P P ++
Sbjct: 360 LITHANAKSKSEKFPPPHQDGQLGFPLGASHHIDPDTVPTDVSFTSVSYNF-PKEPFQAW 536
Query: 455 DG 456
G
Sbjct: 537 SG 542
>TC12151 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {Arabidopsis
thaliana;}, partial (17%)
Length = 708
Score = 186 bits (471), Expect = 1e-47
Identities = 91/105 (86%), Positives = 97/105 (91%)
Frame = +3
Query: 2 LDPLSVNQQGWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIV 61
LD + QGWP WLMAVAG+AI DWTPRRAN+FEKLAKIGQGTYSNVYKA+DLVTGKIV
Sbjct: 390 LDSFTATHQGWPPWLMAVAGDAIRDWTPRRANTFEKLAKIGQGTYSNVYKARDLVTGKIV 569
Query: 62 ALKKVRFDNLEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMS 106
ALKKVRFDNLE ESVKFMAREILVLR+LDHPNVVKLEGLVTSR+S
Sbjct: 570 ALKKVRFDNLEAESVKFMAREILVLRRLDHPNVVKLEGLVTSRIS 704
>TC8079 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, complete
Length = 1550
Score = 168 bits (426), Expect = 2e-42
Identities = 116/322 (36%), Positives = 169/322 (52%), Gaps = 14/322 (4%)
Frame = +3
Query: 41 IGQGTYSNVYKAKDLVTGKIVALKKVR--FDNLEPESVKFMAREILVLRKLDHPNVVKLE 98
IG+G Y V + T ++VA+KK+ FDN K REI +LR LDH NV+ L
Sbjct: 309 IGRGAYGIVCSLLNTETNELVAVKKIANAFDN--HMDAKRTLREIKLLRHLDHENVIALR 482
Query: 99 GLVTS---RMSCSLYLVFEYMEHDLAGL-SAGQGVKFTEPQVKCFMKQLLSGLEHCHSRG 154
++ R +Y+ E M+ DL + + QG+ +E + F+ Q+L GL++ HS
Sbjct: 483 DVIPPPLRREFTDVYITTELMDTDLHQIIRSNQGL--SEEHCQYFLYQVLRGLKYIHSAN 656
Query: 155 VLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGATFYG 214
++HRD+K SNLL+++ LKI DFGLA N MT VVT WYR PELLL ++ Y
Sbjct: 657 IIHRDLKPSNLLLNSNCDLKIIDFGLARPTVEN--DFMTEYVVTRWYRAPELLLNSSDYT 830
Query: 215 VGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQ 274
ID+WS GCI EL+ KP+ PG+ V QL + +L G+P E K + + Q
Sbjct: 831 SAIDVWSVGCIFMELMNKKPLFPGKDHVHQLRLLTELLGTPTEADLGLVKNDDVRRYIRQ 1010
Query: 275 QPY--KRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFF------TTEPYAC 326
P ++ +++ F P ++ L+D +L IDP +R T AL H + EP
Sbjct: 1011LPQYPRQPLTKVFPHVHPMAMDLVDKMLTIDPTKRITVEQALAHPYLEKLHDVADEPVCT 1190
Query: 327 EPSSLPKYPPSKELDVKMRDEE 348
+P S E + + DEE
Sbjct: 1191KPFSF-------EFEQQQLDEE 1235
>TC8547 homologue to UP|MMK1_MEDSA (Q07176) Mitogen-activated protein
kinase homolog MMK1 (MAP kinase MSK7) (MAP kinase ERK1)
, partial (71%)
Length = 1098
Score = 147 bits (370), Expect = 5e-36
Identities = 88/230 (38%), Positives = 126/230 (54%), Gaps = 8/230 (3%)
Frame = +1
Query: 109 LYLVFEYMEHDLAGL-SAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLI 167
+Y+ +E M+ DL + + QG+ +E + F+ Q+L GL++ HS VLHRD+K SNLL+
Sbjct: 64 VYIAYELMDTDLHQIIRSNQGL--SEEHCQYFLYQILRGLKYIHSANVLHRDLKPSNLLL 237
Query: 168 DNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILA 227
+ LKI DFGLA ++ MT VVT WYR PELLL ++ Y ID+WS GCI
Sbjct: 238 NANCDLKICDFGLARV--TSETDFMTEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFM 411
Query: 228 ELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQQPYKR-CISETFK 286
EL+ KP+ PGR V QL + +L G+P+E+ + PY+R E F
Sbjct: 412 ELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFLNENAKRYIRQLPPYRRQSFQEKFP 591
Query: 287 DFPPSSLPLIDSLLAIDPDRRGTASAALNHEFFTT------EPYACEPSS 330
P ++ L++ +L DP +R T AL H + T+ EP P S
Sbjct: 592 QVHPEAIDLVEKMLTFDPRKRITVEEALAHPYLTSLHDISDEPVCMTPFS 741
>TC17660 homologue to UP|Q9M6R8 (Q9M6R8) MAP kinase PsMAPK2, partial (64%)
Length = 717
Score = 144 bits (364), Expect = 3e-35
Identities = 98/240 (40%), Positives = 132/240 (54%), Gaps = 5/240 (2%)
Frame = +2
Query: 61 VALKKVR--FDNLEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCS---LYLVFEY 115
VA+KK++ F+N ++++ + RE+ +LR L H NV+ L+ ++ S +YLV+E
Sbjct: 8 VAIKKIQNAFEN-RVDALRTL-RELKLLRHLHHDNVIALKDIMMPVHRSSFKDVYLVYEL 181
Query: 116 MEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKI 175
M+ DL + + + F+ QLL GL++ HS +LHRD+K NLLI+ LKI
Sbjct: 182 MDTDLHQIIKSSQ-SLSNDHCQYFLFQLLRGLKYLHSANILHRDLKPGNLLINANCDLKI 358
Query: 176 ADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPI 235
DFGLA N +K Q MT VVT WYR PELLL YG ID+WS GCI AELL KPI
Sbjct: 359 CDFGLARI-NCSKNQFMTEYVVTRWYRAPELLLCCDNYGTSIDVWSVGCIFAELLGRKPI 535
Query: 236 MPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPL 295
PG + QL I + GS EE P A + PY I F P++ PL
Sbjct: 536 FPGSECLNQLKLIINILGSQREEDIEFIDNPKAKKYIKSLPYS--IGAPFSRLYPNAHPL 709
>TC16554 homologue to UP|AAN71719 (AAN71719) Glycogen synthase kinase 3 beta
protein kinase DWARF12 , partial (71%)
Length = 927
Score = 140 bits (354), Expect = 4e-34
Identities = 90/240 (37%), Positives = 135/240 (55%), Gaps = 10/240 (4%)
Frame = +1
Query: 29 PRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRK 88
P++ S+ +G G++ V++AK L +G+ VA+KKV D ++ RE+ ++R
Sbjct: 220 PKQTISYMAERVVGTGSFGIVFQAKCLESGEAVAIKKVLQDR------RYKNRELQLMRV 381
Query: 89 LDHPNVVKLEGLVTSRMSCS---LYLVFEYMEHDLAGL-----SAGQGVKFTEPQVKCFM 140
+DHPNVV L+ S S L LV EY+ + + +A Q + VK +M
Sbjct: 382 MDHPNVVSLKHCFFSTTSTDELFLNLVMEYVPESMYRVIKHYSNANQRMPII--YVKLYM 555
Query: 141 KQLLSGLEHCHS-RGVLHRDIKGSNLLIDN-EGILKIADFGLATFYNPNKKQSMTSRVVT 198
Q+ GL + H+ GV HRD+K N+L+D +K+ DFG A K ++ S + +
Sbjct: 556 YQIFRGLAYIHTVPGVCHRDLKPQNILVDPLTHQVKLCDFGSAKMLV--KGEANISYICS 729
Query: 199 LWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEE 258
+YR PEL+ GAT Y ID+WSAGC+LAELL G+P+ PG V+QL I K+ G+P E
Sbjct: 730 RFYRAPELIFGATEYSTSIDIWSAGCVLAELLLGQPLFPGENAVDQLVHIIKVLGTPTRE 909
>TC11248 homologue to UP|P93323 (P93323) Cdc2MsF protein, partial (72%)
Length = 747
Score = 122 bits (307), Expect(2) = 5e-34
Identities = 81/180 (45%), Positives = 102/180 (56%), Gaps = 11/180 (6%)
Frame = +1
Query: 34 SFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRKLDH-P 92
+FEKL K+G+GTY VY+A++ TGKIVALKK R + RE+ +LR L P
Sbjct: 46 AFEKLEKVGEGTYGKVYRAREKATGKIVALKKTRLHEDDEGVPPTTLREVSILRMLSRDP 225
Query: 93 NVVKLEGLVTSRM---SCSLYLVFEYMEHDLAGL-----SAGQGVKFTEPQVKCFMKQLL 144
+VV+L + + LYLVFEYM+ DL GQ V VK M QL
Sbjct: 226 HVVRLMDVKQGQSKEGKTVLYLVFEYMDTDLKKFIRTFRQTGQNV--PPKTVKSLMYQLR 399
Query: 145 SGLEHCHSRGVLHRDIKGSNLLIDNE-GILKIADFGLA-TFYNPNKKQSMTSRVVTLWYR 202
G+ CH G+LHRD+K NLL+D E +LKIAD GLA F P KK T ++TLWYR
Sbjct: 400 KGVAFCHGHGILHRDLKPHNLLMDRETNMLKIADLGLARAFTVPIKK--YTHEILTLWYR 573
Score = 38.9 bits (89), Expect(2) = 5e-34
Identities = 15/26 (57%), Positives = 20/26 (76%)
Frame = +2
Query: 205 ELLLGATFYGVGIDLWSAGCILAELL 230
E+LLGAT Y + +D+WS CI AEL+
Sbjct: 581 EVLLGATHYSMAVDMWSVACIFAELV 658
>AV423422
Length = 375
Score = 139 bits (350), Expect = 1e-33
Identities = 72/128 (56%), Positives = 95/128 (73%)
Frame = +1
Query: 283 ETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFFTTEPYACEPSSLPKYPPSKELDV 342
ETFKDFP ++ LI++LL+IDP RGT+++AL EFF+T+P C+PSSLPKYPPSKE D
Sbjct: 1 ETFKDFPAPAIELIETLLSIDPTDRGTSASALKSEFFSTKPLPCDPSSLPKYPPSKEFDA 180
Query: 343 KMRDEEARRQKALNGKANAVDGAKRVRARERGRAIPAPEANAEIQTNLDRWRVVTHANAK 402
K+R+EEARRQ A KA D +RVR RAIPAPEANAE+ ++ + R +T +++
Sbjct: 181 KVREEEARRQGAAGSKAQRHDPERRVR---ESRAIPAPEANAELVLSMQKRRGLT--SSQ 345
Query: 403 SKSEKFPP 410
S+SEKF P
Sbjct: 346 SRSEKFNP 369
>TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein kinase-like
protein (CBL-interacting protein kinase 5), partial
(75%)
Length = 1999
Score = 123 bits (308), Expect = 8e-29
Identities = 85/289 (29%), Positives = 146/289 (50%), Gaps = 2/289 (0%)
Frame = +3
Query: 33 NSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPES-VKFMAREILVLRKLDH 91
N +E +GQG ++ VY ++L T + VA+K ++ + L+ + VK + RE+ V+R + H
Sbjct: 426 NKYEMGRVLGQGNFAKVYHGRNLATNENVAIKVIKKEKLKKDRLVKQIKREVSVMRLVRH 605
Query: 92 PNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEHCH 151
P++V+L+ ++ ++ ++LV EY++ +G K E + + +QL+S ++ CH
Sbjct: 606 PHIVELKEVMATKGK--IFLVMEYVKGGELFTKVNKG-KLNEDDARKYFQQLISAVDFCH 776
Query: 152 SRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSM-TSRVVTLWYRPPELLLGA 210
SRGV HRD+K NLL+D LK++DFGL+ + M + T Y PE+L
Sbjct: 777 SRGVTHRDLKPENLLLDENEDLKVSDFGLSALPEQRRDDGMLVTPCGTPAYVAPEVLKKK 956
Query: 211 TFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATI 270
+ G D+WS G IL LL+G G E + +I++
Sbjct: 957 GYDGSKADIWSCGVILYALLSGYLPFQG----ENVMRIYR------------------KA 1070
Query: 271 FKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFF 319
FK + + IS P + LI +LL DP++R + ++ +F
Sbjct: 1071FKAEYEFPEWIS-------PQAKNLISNLLVADPEKRYSIPEIISDPWF 1196
>BP063610
Length = 461
Score = 121 bits (304), Expect = 2e-28
Identities = 67/145 (46%), Positives = 87/145 (59%)
Frame = -3
Query: 57 TGKIVALKKVRFDNLEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYM 116
TG+IVALKKV + + REI +L HP +V ++ +V S+++V EYM
Sbjct: 459 TGEIVALKKVXMEKEKEGFPLTSLREINILLSFHHPYIVDVKEVVVGSSLDSIFMVMEYM 280
Query: 117 EHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIA 176
EHDL GL F++ +VKC M QLL G+++ H VLHRD+K SNLL++N G LKI
Sbjct: 279 EHDLKGLMESMKQPFSQSEVKCLMIQLLEGVKYLHDNWVLHRDLKTSNLLLNNRGELKIC 100
Query: 177 DFGLATFYNPNKKQSMTSRVVTLWY 201
DFGLA Y K T VVTLWY
Sbjct: 99 DFGLARXYGSPLK-PYTHLVVTLWY 28
>TC8907 similar to UP|Q8H0X3 (Q8H0X3) Serine/threonine protein kinase-like
protein, partial (44%)
Length = 941
Score = 118 bits (296), Expect = 2e-27
Identities = 71/204 (34%), Positives = 120/204 (58%), Gaps = 4/204 (1%)
Frame = +3
Query: 33 NSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPES-VKFMAREILVLRKLDH 91
N +E +GQG ++ VY +++ T + VA+K ++ + L+ E VK + RE+ V+R + H
Sbjct: 336 NKYEIGKILGQGNFAKVYHGRNMETNESVAIKVIKKERLKKERLVKQIKREVSVMRLVRH 515
Query: 92 PNVVKLEGLVTSRMSCSLYLVFEYMEHD--LAGLSAGQGVKFTEPQVKCFMKQLLSGLEH 149
P++V+L+ ++ ++ +++V EY++ A L+ G K TE + + +QL+S ++
Sbjct: 516 PHIVELKEVMATKTK--IFMVVEYVKGGELFAKLTKG---KMTEVAARKYFQQLISAVDF 680
Query: 150 CHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSM-TSRVVTLWYRPPELLL 208
CHSRGV HRD+K NLL+D+ LK++DFGL++ + M + T Y PE+L
Sbjct: 681 CHSRGVTHRDLKPENLLLDDNEDLKVSDFGLSSLPEQRRSDGMLLTPCGTPAYVAPEVLK 860
Query: 209 GATFYGVGIDLWSAGCILAELLAG 232
+ G D+WS G IL LL G
Sbjct: 861 KKGYDGSKADIWSCGVILYALLCG 932
>TC18815 similar to UP|Q8W1D5 (Q8W1D5) CBL-interacting protein kinase
CIPK25, partial (58%)
Length = 979
Score = 112 bits (280), Expect = 1e-25
Identities = 80/281 (28%), Positives = 139/281 (48%), Gaps = 2/281 (0%)
Frame = +2
Query: 41 IGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKF-MAREILVLRKLDHPNVVKLEG 99
+G+GT + VY AK++ +G+ VA+K + ++ E + + REI ++R + HPN+V L+
Sbjct: 56 LGKGTLAKVYFAKEITSGEGVAIKVMSKARIKKEGMMDQIKREISIMRLVRHPNIVNLKE 235
Query: 100 LVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGVLHRD 159
++ ++ ++ + EY+ +G K + + + +QL+S +++CHSRGV HRD
Sbjct: 236 VMATKTK--IFFIMEYIRGGELFAKVAKG-KLKDDLARRYFQQLISAVDYCHSRGVSHRD 406
Query: 160 IKGSNLLIDNEGILKIADFGLATFYNPNKKQSMT-SRVVTLWYRPPELLLGATFYGVGID 218
+K NLL+D LK++DFGL+ ++ + ++ T Y PE+L + G D
Sbjct: 407 LKPENLLLDENENLKVSDFGLSGLPEQLRQDGLLHTQCGTPAYVAPEVLRKKGYDGFKTD 586
Query: 219 LWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQQPYK 278
WS G IL LLAG + +K+ + + + P P+
Sbjct: 587 TWSCGVILYALLAGCLPFQHENLMTMYNKVLR----------AEFQFP---------PW- 706
Query: 279 RCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFF 319
F P S LI +L DP+RR T S+ + +F
Sbjct: 707 ---------FSPESKKLISKILVADPNRRITISSIMRVSWF 802
>TC17375 mitogen-activated kinase kinase kinase alpha [Lotus corniculatus
var. japonicus]
Length = 1422
Score = 112 bits (280), Expect = 1e-25
Identities = 80/263 (30%), Positives = 130/263 (49%), Gaps = 3/263 (1%)
Frame = +3
Query: 59 KIVALKKVRF---DNLEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEY 115
++ A+K+V+ D E +K + +EI +L + HPN+V+ G S S+YL EY
Sbjct: 3 QMCAIKEVKVFSDDKTSKECLKQLNQEINLLNQFSHPNIVQYYGSELGEESLSVYL--EY 176
Query: 116 MEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKI 175
+ + F EP ++ + +Q++SGL + HSR +HRDIKG+N+L+D G +K+
Sbjct: 177 VSGGSIHKLLQEYGAFKEPVIQNYTRQIVSGLAYLHSRNTVHRDIKGANILVDPNGEIKL 356
Query: 176 ADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPI 235
ADFG++ + N SM S + ++ PE+++ YG+ +D+ S GC + E+ KP
Sbjct: 357 ADFGMSK--HINSAASMLSFKGSPYWMAPEVVMNTNGYGLPVDISSLGCTILEMATSKPP 530
Query: 236 MPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATIFKPQQPYKRCISETFKDFPPSSLPL 295
++ E + IFK+ S K +S+ K+F
Sbjct: 531 W---SQFEGVAAIFKIGNS-----------------KDMPEIPEHLSDDAKNF------- 629
Query: 296 IDSLLAIDPDRRGTASAALNHEF 318
I L DP R TA + LNH F
Sbjct: 630 IKQCLQRDPLARPTAQSLLNHPF 698
>AV419778
Length = 411
Score = 102 bits (255), Expect = 1e-22
Identities = 51/123 (41%), Positives = 75/123 (60%)
Frame = +1
Query: 138 CFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVV 197
CF Q+ GL + H RG HRD+K NLL+ + I+K++DFGLA + + T V
Sbjct: 10 CF--QVFQGLAYMHQRGYFHRDLKPENLLVTKD-IIKVSDFGLAREISSHPPY--TEYVS 174
Query: 198 TLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAE 257
T WYR PE+LL + Y +D+W+ G I+AEL +P+ PG +E ++++KI + GSP
Sbjct: 175 TRWYRAPEVLLQSYLYSSKVDMWAMGAIMAELFTLRPLFPGTSEADEIYKICSVIGSPTT 354
Query: 258 EYW 260
E W
Sbjct: 355 ESW 363
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.134 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,003,921
Number of Sequences: 28460
Number of extensions: 127325
Number of successful extensions: 952
Number of sequences better than 10.0: 206
Number of HSP's better than 10.0 without gapping: 871
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 880
length of query: 498
length of database: 4,897,600
effective HSP length: 94
effective length of query: 404
effective length of database: 2,222,360
effective search space: 897833440
effective search space used: 897833440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146747.6


