

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146723.12 + phase: 0
(172 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
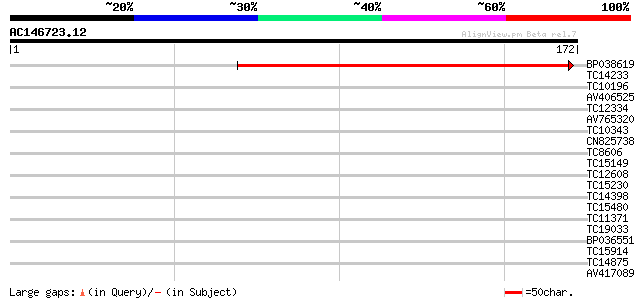
Score E
Sequences producing significant alignments: (bits) Value
BP038619 179 3e-46
TC14233 homologue to UP|Q41027 (Q41027) PsHSC71.0, partial (67%) 30 0.29
TC10196 homologue to UP|Q39804 (Q39804) BiP isoform B, partial (... 29 0.38
AV406525 28 0.85
TC12334 28 0.85
AV765320 28 0.85
TC10343 similar to UP|Q9SZZ1 (Q9SZZ1) Fibrillarin-like protein (... 28 0.85
CN825738 27 1.9
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 26 3.2
TC15149 similar to UP|Q9LVI6 (Q9LVI6) Probable receptor-like pro... 26 4.2
TC12608 26 4.2
TC15230 similar to GB|AAP75810.1|32189311|BT009660 At1g70180 {Ar... 26 4.2
TC14398 similar to UP|Q96477 (Q96477) LRR protein, partial (86%) 25 5.5
TC15480 similar to UP|V712_ARATH (Q9SIQ9) Vesicle-associated mem... 25 5.5
TC11371 similar to UP|Q8S9K1 (Q8S9K1) At1g23880/T23E23_8, partia... 25 5.5
TC19033 similar to UP|Q8H6X8 (Q8H6X8) GSK-3-like protein MsK4, p... 25 5.5
BP036551 25 7.2
TC15914 25 7.2
TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%) 25 9.4
AV417089 25 9.4
>BP038619
Length = 460
Score = 179 bits (453), Expect = 3e-46
Identities = 80/102 (78%), Positives = 92/102 (89%)
Frame = -3
Query: 70 PEFADSETEKFQNHLLNKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPGTLLVDPFI 129
PEFAD+ETEKF++HLLNKLSK+D++ +SV EVVGVCTEIF +FLHSEYGGPGTLLV+PF
Sbjct: 458 PEFADAETEKFKSHLLNKLSKRDIYGDSVGEVVGVCTEIFGSFLHSEYGGPGTLLVNPFA 279
Query: 130 DMADIVNERGLPGGPQAARAAINWAQAHVDKDWNEWTGGNSN 171
DMAD V+ERGLPGGPQAAR A+ WAQ HVDKDWNEWTGG +
Sbjct: 278 DMADAVSERGLPGGPQAARVAVKWAQNHVDKDWNEWTGGGDS 153
>TC14233 homologue to UP|Q41027 (Q41027) PsHSC71.0, partial (67%)
Length = 1570
Score = 29.6 bits (65), Expect = 0.29
Identities = 16/62 (25%), Positives = 26/62 (41%)
Frame = +1
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPGTLLVDPFIDMADIVNERGLPGGPQ 145
N+L++ D FE+ ++E+ +C I + GGP D+ E P G
Sbjct: 1132 NQLAEADEFEDKMKELESICNPIIAKMYQGGAGGP---------DVGGAAEEEYAPSGGS 1284
Query: 146 AA 147
A
Sbjct: 1285 GA 1290
>TC10196 homologue to UP|Q39804 (Q39804) BiP isoform B, partial (17%)
Length = 562
Score = 29.3 bits (64), Expect = 0.38
Identities = 12/36 (33%), Positives = 20/36 (55%)
Frame = +2
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPG 121
N+ ++K+ +EE ++EV VC I + G PG
Sbjct: 188 NQSAEKEEYEEKLKEVEAVCNPIITAVYQRSGGAPG 295
>AV406525
Length = 271
Score = 28.1 bits (61), Expect = 0.85
Identities = 16/40 (40%), Positives = 22/40 (55%)
Frame = +1
Query: 10 PFLIPSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLP 49
PFL+ + + +LCF ++ FSFPS RK ST P
Sbjct: 136 PFLV---AVIVTALCFFLALFSLLRPFSFPSSLRKKSTPP 246
>TC12334
Length = 490
Score = 28.1 bits (61), Expect = 0.85
Identities = 15/37 (40%), Positives = 18/37 (48%), Gaps = 2/37 (5%)
Frame = -1
Query: 3 SSTWCLVPFLIPSSSSSIPSL--CFSQTTTKCIQRFS 37
S W PF PSS SS P+ FS T + R+S
Sbjct: 121 SQPWMRTPFRPPSSKSSAPAAFRSFSSTANSIVLRYS 11
>AV765320
Length = 572
Score = 28.1 bits (61), Expect = 0.85
Identities = 17/42 (40%), Positives = 23/42 (54%)
Frame = +1
Query: 8 LVPFLIPSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLP 49
L PFL SSSS+ S S +T++ + P RKV T+P
Sbjct: 367 LTPFLAAISSSSLASSILSSSTSE--EPVDSPRAFRKVKTMP 486
>TC10343 similar to UP|Q9SZZ1 (Q9SZZ1) Fibrillarin-like protein (Fibrillarin
2) (AtFib2), partial (85%)
Length = 1116
Score = 28.1 bits (61), Expect = 0.85
Identities = 16/42 (38%), Positives = 23/42 (54%)
Frame = -1
Query: 15 SSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLPFSSIKRS 56
+SSS+ P+ S T T C++R + S + T P SSI S
Sbjct: 201 ASSSATPASSSSSTATSCLKRGTSTSSSSSSITSPSSSITPS 76
>CN825738
Length = 335
Score = 26.9 bits (58), Expect = 1.9
Identities = 22/62 (35%), Positives = 29/62 (46%), Gaps = 7/62 (11%)
Frame = -1
Query: 8 LVPFLIPSS-------SSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLPFSSIKRSITRA 60
L FLIPSS SSS + FS T+TK ST+PFS + + ++
Sbjct: 326 LTSFLIPSSTKPLPLTSSSASVIPFSPTSTK--------------STIPFSPLPPTSSKG 189
Query: 61 AE 62
AE
Sbjct: 188 AE 183
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 26.2 bits (56), Expect = 3.2
Identities = 18/51 (35%), Positives = 26/51 (50%)
Frame = +1
Query: 2 ASSTWCLVPFLIPSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLPFSS 52
+SS+ L PSSSSS P S +++ S+ S + S+ PFSS
Sbjct: 292 SSSSSSLSSSSTPSSSSSSPPSSSSSSSSSSSSSSSWSSPSSSSSSSPFSS 444
>TC15149 similar to UP|Q9LVI6 (Q9LVI6) Probable receptor-like protein kinase
protein (AT3g17840/MEB5_6), partial (45%)
Length = 1489
Score = 25.8 bits (55), Expect = 4.2
Identities = 14/59 (23%), Positives = 26/59 (43%), Gaps = 7/59 (11%)
Frame = -1
Query: 19 SIPSLCFSQTTTKCIQRFS-------FPSINRKVSTLPFSSIKRSITRAAEYKFPDPIP 70
++P + F T+ + + + S P ++ P +S + AAEY P P+P
Sbjct: 319 AVPKVPFPNTSAEALSKSSRSNPLAELPKNTSFLAPTPLASTLLMVAAAAEYPLPLPLP 143
>TC12608
Length = 735
Score = 25.8 bits (55), Expect = 4.2
Identities = 10/19 (52%), Positives = 12/19 (62%)
Frame = -1
Query: 10 PFLIPSSSSSIPSLCFSQT 28
PFLIP S +P LC S +
Sbjct: 249 PFLIPRKHSELPLLCISNS 193
>TC15230 similar to GB|AAP75810.1|32189311|BT009660 At1g70180 {Arabidopsis
thaliana;}, partial (11%)
Length = 611
Score = 25.8 bits (55), Expect = 4.2
Identities = 17/44 (38%), Positives = 25/44 (56%), Gaps = 3/44 (6%)
Frame = -3
Query: 2 ASSTWCLVPFLIPSSSSSIPSLCFSQ--TTTKCIQRFS-FPSIN 42
AS + L P IPSS S +CF+ +T+ ++R + FPS N
Sbjct: 255 ASRIFFLGPIGIPSSLRSFSPICFNAVISTSSALKRMTYFPSPN 124
>TC14398 similar to UP|Q96477 (Q96477) LRR protein, partial (86%)
Length = 982
Score = 25.4 bits (54), Expect = 5.5
Identities = 15/42 (35%), Positives = 19/42 (44%), Gaps = 12/42 (28%)
Frame = -3
Query: 14 PSSSSSIPSLCFSQTT------------TKCIQRFSFPSINR 43
PSSS +P + FS ++ TKC RF FP R
Sbjct: 458 PSSSGIVP*MLFSYSSRYCRECRFPRSGTKCPDRFKFPRSTR 333
>TC15480 similar to UP|V712_ARATH (Q9SIQ9) Vesicle-associated membrane
protein 712 (AtVAMP712), partial (53%)
Length = 482
Score = 25.4 bits (54), Expect = 5.5
Identities = 26/91 (28%), Positives = 41/91 (44%), Gaps = 1/91 (1%)
Frame = +1
Query: 3 SSTWCLVPFLIPSSSSSIPSLCFSQTTTKCIQRFSFPSINR-KVSTLPFSSIKRSITRAA 61
SS+ PFL+PSSSSS S+ S + C + + R V+ FS + + A
Sbjct: 40 SSSSSSSPFLLPSSSSS--SIHPSIHRSSCRMGILYALVARGSVALAEFSGTATNASTIA 213
Query: 62 EYKFPDPIPEFADSETEKFQNHLLNKLSKKD 92
+ + D IP D+ Q+ + L + D
Sbjct: 214 K-QILDKIPGNNDTHVSYSQDRFIFHLKRTD 303
>TC11371 similar to UP|Q8S9K1 (Q8S9K1) At1g23880/T23E23_8, partial (41%)
Length = 1023
Score = 25.4 bits (54), Expect = 5.5
Identities = 21/49 (42%), Positives = 25/49 (50%), Gaps = 9/49 (18%)
Frame = -1
Query: 16 SSSSIPSLCFSQTTT-------KCIQRFSFP--SINRKVSTLPFSSIKR 55
S SS+ S+ TT K I++FSF SIN ST PFSS R
Sbjct: 783 SDSSVSSIYNE*RTTASNINHIKVIRKFSFFTWSINMAPSTAPFSSSNR 637
>TC19033 similar to UP|Q8H6X8 (Q8H6X8) GSK-3-like protein MsK4, partial
(18%)
Length = 665
Score = 25.4 bits (54), Expect = 5.5
Identities = 22/70 (31%), Positives = 30/70 (42%)
Frame = -3
Query: 13 IPSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEF 72
+PSS SS SL + S PS + +V + +T E P P P
Sbjct: 474 VPSSGSSTLSLNLXSS-------ISLPSQSTEVEVEVEEEEEPPLTTDLELPTP-PFPRL 319
Query: 73 ADSETEKFQN 82
AD+ T KFQ+
Sbjct: 318 ADAITFKFQS 289
>BP036551
Length = 428
Score = 25.0 bits (53), Expect = 7.2
Identities = 13/35 (37%), Positives = 18/35 (51%)
Frame = +3
Query: 7 CLVPFLIPSSSSSIPSLCFSQTTTKCIQRFSFPSI 41
C PF+I SS S S F + T C F+F ++
Sbjct: 261 CCNPFMISSSIIS*YSTVFRVSATFCAASFAFTAL 365
>TC15914
Length = 498
Score = 25.0 bits (53), Expect = 7.2
Identities = 12/33 (36%), Positives = 20/33 (60%)
Frame = -1
Query: 14 PSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVS 46
PSS +++ S S +T + QR+S +NR +S
Sbjct: 108 PSSFTTVLSRVTSTSTKRIPQRYSESPVNRSIS 10
>TC14875 similar to UP|Q9VNX6 (Q9VNX6) CG7421-PA, partial (4%)
Length = 1240
Score = 24.6 bits (52), Expect = 9.4
Identities = 6/15 (40%), Positives = 11/15 (73%)
Frame = +1
Query: 158 VDKDWNEWTGGNSNE 172
++++W EW GGN +
Sbjct: 760 IEEEWVEWNGGNGGD 804
>AV417089
Length = 407
Score = 24.6 bits (52), Expect = 9.4
Identities = 12/27 (44%), Positives = 17/27 (62%)
Frame = +3
Query: 1 MASSTWCLVPFLIPSSSSSIPSLCFSQ 27
M S +C +PF +P+ SSI L FS+
Sbjct: 204 MTSKRYCFLPFFLPTLLSSI--LIFSR 278
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.132 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,533,127
Number of Sequences: 28460
Number of extensions: 51477
Number of successful extensions: 381
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 378
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 379
length of query: 172
length of database: 4,897,600
effective HSP length: 84
effective length of query: 88
effective length of database: 2,506,960
effective search space: 220612480
effective search space used: 220612480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146723.12


