

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146722.4 - phase: 1
(88 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
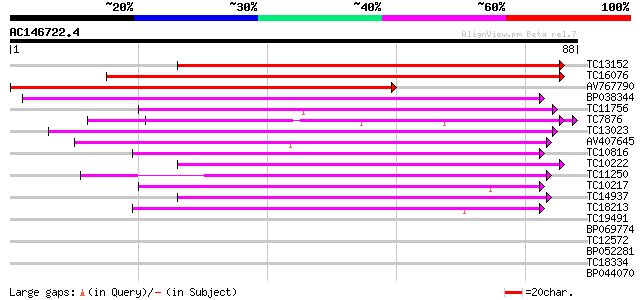
Score E
Sequences producing significant alignments: (bits) Value
TC13152 similar to UP|Q84XX6 (Q84XX6) Leunig (Fragment), partial... 118 2e-28
TC16076 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG... 92 2e-20
AV767790 84 4e-18
BP038344 56 1e-09
TC11756 weakly similar to GB|AAH01494.1|16306637|BC001494 U5 snR... 49 1e-07
TC7876 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-bi... 48 3e-07
TC13023 similar to UP|Q8L830 (Q8L830) WD40-repeat protein (Fragm... 44 5e-06
AV407645 44 5e-06
TC10816 homologue to UP|CAE45585 (CAE45585) Coatomer alpha subun... 40 6e-05
TC10222 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Ar... 40 9e-05
TC11250 homologue to UP|AAQ72359 (AAQ72359) B-type cell cycle sw... 39 2e-04
TC10217 homologue to UP|AAQ54906 (AAQ54906) Cell cycle switch pr... 37 6e-04
TC14937 similar to UP|Q9FN19 (Q9FN19) Genomic DNA, chromosome 5,... 37 8e-04
TC18213 weakly similar to UP|Q9FIV2 (Q9FIV2) Similarity to pre-m... 37 8e-04
TC19491 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Ar... 36 0.001
BP069774 36 0.001
TC12572 similar to PIR|T46032|T46032 WD-40 repeat regulatory pro... 35 0.002
BP052281 35 0.002
TC18334 weakly similar to UP|Q9LHN3 (Q9LHN3) Emb|CAB63739.1 (AT3... 35 0.003
BP044070 34 0.004
>TC13152 similar to UP|Q84XX6 (Q84XX6) Leunig (Fragment), partial (60%)
Length = 453
Score = 118 bits (295), Expect = 2e-28
Identities = 56/60 (93%), Positives = 58/60 (96%)
Frame = +2
Query: 27 LLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDVDN 86
LLASGGHDKK VLWYTDSLKQKATLEEHS LITDVRFSPSMPRLATSS+DKTV+VWDVDN
Sbjct: 2 LLASGGHDKKAVLWYTDSLKQKATLEEHSALITDVRFSPSMPRLATSSFDKTVKVWDVDN 181
>TC16076 weakly similar to GB|AAG32022.1|11141605|AF277458 LEUNIG
{Arabidopsis thaliana;} , partial (24%)
Length = 1048
Score = 92.0 bits (227), Expect = 2e-20
Identities = 42/71 (59%), Positives = 55/71 (77%)
Frame = +3
Query: 16 VVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSY 75
V C HFSSDGK+LAS G DKKVVLW D+L+ ++T E+H +I+DVRF P+ +LAT+S
Sbjct: 3 VTCCHFSSDGKMLASAGDDKKVVLWNMDTLQTESTPEQHKSVISDVRFRPNSSQLATASI 182
Query: 76 DKTVRVWDVDN 86
DK+VR+WD N
Sbjct: 183DKSVRLWDAAN 215
>AV767790
Length = 617
Score = 84.0 bits (206), Expect = 4e-18
Identities = 38/60 (63%), Positives = 50/60 (83%)
Frame = +3
Query: 1 FTFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITD 60
F+FSE+ S+R S +KVVC HFSSDGKLLAS GHD+KVVLW ++L+ ++T + HSL+ITD
Sbjct: 438 FSFSEVGSIRKSNSKVVCCHFSSDGKLLASAGHDQKVVLWNMETLQTESTPDAHSLIITD 617
>BP038344
Length = 525
Score = 55.8 bits (133), Expect = 1e-09
Identities = 28/81 (34%), Positives = 46/81 (56%)
Frame = +2
Query: 3 FSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVR 62
+ L ++ A V C FS+DG LLAS DK +++W + +L L HS I+D+
Sbjct: 68 YRHLKTLTAHDAAVACVKFSNDGTLLASASLDKTLIIWSSATLSLLHRLTGHSEGISDLA 247
Query: 63 FSPSMPRLATSSYDKTVRVWD 83
+S + ++S D+T+R+WD
Sbjct: 248 WSSDSHYICSASDDRTLRIWD 310
Score = 28.9 bits (63), Expect = 0.17
Identities = 14/39 (35%), Positives = 21/39 (52%)
Frame = +2
Query: 50 TLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDVDNVS 88
TL H + V+FS LA++S DKT+ +W +S
Sbjct: 83 TLTAHDAAVACVKFSNDGTLLASASLDKTLIIWSSATLS 199
>TC11756 weakly similar to GB|AAH01494.1|16306637|BC001494 U5 snRNP-specific
40 kDa protein (hPrp8-binding) {Homo sapiens;} , partial
(43%)
Length = 719
Score = 49.3 bits (116), Expect = 1e-07
Identities = 23/66 (34%), Positives = 39/66 (58%), Gaps = 1/66 (1%)
Frame = +1
Query: 21 FSSDGKLLASGGHDKKVVLWYTDS-LKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTV 79
F+ G ++ASG HD+++ LW K L+ H + D+ ++ ++ ++S DKTV
Sbjct: 328 FNPAGTVIASGSHDREIFLWNVHGECKNFMVLKGHKNAVLDLHWTTDGTQIVSASPDKTV 507
Query: 80 RVWDVD 85
RVWDV+
Sbjct: 508 RVWDVE 525
Score = 26.6 bits (57), Expect = 0.83
Identities = 10/34 (29%), Positives = 22/34 (64%)
Frame = +1
Query: 51 LEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDV 84
L H +I ++F+P+ +A+ S+D+ + +W+V
Sbjct: 292 LTGHQSVIYTMKFNPAGTVIASGSHDREIFLWNV 393
>TC7876 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-binding
protein beta subunit-like protein, complete
Length = 1379
Score = 48.1 bits (113), Expect = 3e-07
Identities = 28/79 (35%), Positives = 47/79 (59%), Gaps = 5/79 (6%)
Frame = +3
Query: 13 TNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEE---HSLLITDVRFSPS--M 67
T V+ FS D + + S D+ + LW T + K T+++ HS ++ VRFSPS
Sbjct: 384 TKDVLSVAFSIDNRQIVSASRDRTIKLWNTLG-ECKYTIQDNDAHSDWVSCVRFSPSTLQ 560
Query: 68 PRLATSSYDKTVRVWDVDN 86
P + ++S+D+TV+VW++ N
Sbjct: 561 PTIVSASWDRTVKVWNLTN 617
Score = 37.0 bits (84), Expect = 6e-04
Identities = 22/67 (32%), Positives = 40/67 (58%)
Frame = +3
Query: 22 SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRV 81
S DG L ASGG D ++LW K+ +L+ S +I + FSP+ L ++ + ++++
Sbjct: 675 SPDGSLCASGGKDGVILLWDLAEGKRLYSLDAGS-IIHALCFSPNRYWLCAAT-ETSIKI 848
Query: 82 WDVDNVS 88
WD+++ S
Sbjct: 849 WDLESKS 869
Score = 34.7 bits (78), Expect = 0.003
Identities = 20/71 (28%), Positives = 35/71 (49%), Gaps = 6/71 (8%)
Frame = +3
Query: 19 SHF------SSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLAT 72
SHF SSDG+ SG D ++ LW + H+ + V FS ++ +
Sbjct: 258 SHFVQDVVLSSDGQFALSGSWDGELRLWDLAAGTSARRFVGHTKDVLSVAFSIDNRQIVS 437
Query: 73 SSYDKTVRVWD 83
+S D+T+++W+
Sbjct: 438 ASRDRTIKLWN 470
>TC13023 similar to UP|Q8L830 (Q8L830) WD40-repeat protein (Fragment),
partial (18%)
Length = 591
Score = 43.9 bits (102), Expect = 5e-06
Identities = 24/79 (30%), Positives = 39/79 (48%)
Frame = +1
Query: 7 NSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPS 66
++++ + V S D LL S GH +++ +W +LK + + H + + PS
Sbjct: 265 STLQGDSESVTALALSPDDNLLFSSGHSRQIRVWDLSTLKCVRSWKGHEGPVMCMSCHPS 444
Query: 67 MPRLATSSYDKTVRVWDVD 85
LAT D+ V VWDVD
Sbjct: 445 GGLLATGGADRKVLVWDVD 501
Score = 33.9 bits (76), Expect = 0.005
Identities = 19/64 (29%), Positives = 30/64 (46%)
Frame = +1
Query: 2 TFSELNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDV 61
T + S + V+C G LLA+GG D+KV++W D + H +++ V
Sbjct: 376 TLKCVRSWKGHEGPVMCMSCHPSGGLLATGGADRKVLVWDVDGGYCTHFFKGHGGVVSCV 555
Query: 62 RFSP 65
F P
Sbjct: 556 MFHP 567
>AV407645
Length = 413
Score = 43.9 bits (102), Expect = 5e-06
Identities = 23/76 (30%), Positives = 38/76 (49%), Gaps = 2/76 (2%)
Frame = +2
Query: 11 ASTNKVVCSHFSSDGKLLASGGHDKKVVLWYT--DSLKQKATLEEHSLLITDVRFSPSMP 68
A ++V FS +GK LAS +D+ ++W + L K L H + + +SP+
Sbjct: 2 AHDDEVWLVQFSHNGKYLASASNDRTAIIWEVGINGLSLKHRLSGHQKAFSSISWSPNDQ 181
Query: 69 RLATSSYDKTVRVWDV 84
L T ++ +R WDV
Sbjct: 182ELLTCGVEEAIRRWDV 229
>TC10816 homologue to UP|CAE45585 (CAE45585) Coatomer alpha subunit-like
protein, partial (20%)
Length = 979
Score = 40.4 bits (93), Expect = 6e-05
Identities = 21/64 (32%), Positives = 32/64 (49%)
Frame = +1
Query: 20 HFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTV 79
HF L SGG D K+ +W + TL H I V+F P + ++S D+T+
Sbjct: 412 HFHHSQPLFVSGGDDYKIKVWNYKLHRCLFTLLGHLDYIRTVQFHHESPWIVSASDDQTI 591
Query: 80 RVWD 83
R+W+
Sbjct: 592 RIWN 603
Score = 35.4 bits (80), Expect = 0.002
Identities = 17/67 (25%), Positives = 32/67 (47%)
Frame = +1
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVR 80
F + + S D+ + +W S + L H+ + F P + ++S D+TVR
Sbjct: 541 FHHESPWIVSASDDQTIRIWNWQSRTCISVLTGHNHYVMCALFHPREDLVVSASLDQTVR 720
Query: 81 VWDVDNV 87
VWD+ ++
Sbjct: 721 VWDIGSL 741
Score = 29.6 bits (65), Expect = 0.098
Identities = 21/98 (21%), Positives = 40/98 (40%), Gaps = 27/98 (27%)
Frame = +1
Query: 16 VVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQ---------------------------K 48
V+C+ F L+ S D+ V +W SLK+ K
Sbjct: 652 VMCALFHPREDLVVSASLDQTVRVWDIGSLKRKNASPADDILRLSQMNTDLFGGVDAVVK 831
Query: 49 ATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDVDN 86
LE H + F P++P + +++ D+ V++W +++
Sbjct: 832 YVLEGHDRGVNWASFHPALPLIVSAADDRQVKLWRMND 945
Score = 23.9 bits (50), Expect = 5.4
Identities = 16/78 (20%), Positives = 27/78 (34%)
Frame = +1
Query: 6 LNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSP 65
L +N+V F + + H + LW +EH + V F
Sbjct: 244 LTKFETKSNRVKGLSFHPKRPWILASLHSGVIQLWDYRMGTLIDKFDEHDGPVRGVHFHH 423
Query: 66 SMPRLATSSYDKTVRVWD 83
S P + D ++VW+
Sbjct: 424 SQPLFVSGGDDYKIKVWN 477
>TC10222 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Arabidopsis
thaliana;}, partial (55%)
Length = 557
Score = 39.7 bits (91), Expect = 9e-05
Identities = 24/60 (40%), Positives = 32/60 (53%)
Frame = +1
Query: 27 LLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDVDN 86
LL +G D+ V LW +D L + T H L + V P A+SS D VRV+DVD+
Sbjct: 106 LLLTGSLDETVRLWRSDELVLERTNTGHCLGVASVAAHPLGSIAASSSLDSFVRVFDVDS 285
Score = 27.3 bits (59), Expect = 0.48
Identities = 19/60 (31%), Positives = 28/60 (46%), Gaps = 1/60 (1%)
Frame = +1
Query: 25 GKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATS-SYDKTVRVWD 83
G + AS D V ++ DS ATLE + +RF P LA + +V++WD
Sbjct: 226 GSIAASSSLDSFVRVFDVDSNATLATLEAPPSEVWQMRFDPKGALLAVAGGGSASVKLWD 405
Score = 25.4 bits (54), Expect = 1.8
Identities = 24/86 (27%), Positives = 34/86 (38%), Gaps = 15/86 (17%)
Frame = +1
Query: 6 LNSVRASTNKVVCSHFSSDGKLLA-SGGHDKKVVLWYTDSLKQKATL------------- 51
L ++ A ++V F G LLA +GG V LW T + + ATL
Sbjct: 295 LATLEAPPSEVWQMRFDPKGALLAVAGGGSASVKLWDTSTWELVATLSIPRPEGGSKPTD 474
Query: 52 -EEHSLLITDVRFSPSMPRLATSSYD 76
+ V +SP R+A S D
Sbjct: 475 KSGSKKFVLSVAWSPDGKRIACGSMD 552
>TC11250 homologue to UP|AAQ72359 (AAQ72359) B-type cell cycle switch
protein ccs52B, partial (28%)
Length = 564
Score = 38.5 bits (88), Expect = 2e-04
Identities = 20/73 (27%), Positives = 37/73 (50%)
Frame = +2
Query: 12 STNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLA 71
+ N++V +H G ++++W SL + ATL HS+ + + SP +
Sbjct: 176 NVNEIVSTH----------GYSQNQIMVWKYPSLSKVATLTGHSMRVLYLAMSPDGQTIV 325
Query: 72 TSSYDKTVRVWDV 84
T + D+T+R W+V
Sbjct: 326 TGAGDETLRFWNV 364
>TC10217 homologue to UP|AAQ54906 (AAQ54906) Cell cycle switch protein
CCS52a, partial (45%)
Length = 640
Score = 37.0 bits (84), Expect = 6e-04
Identities = 19/66 (28%), Positives = 35/66 (52%), Gaps = 3/66 (4%)
Frame = +2
Query: 21 FSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATS---SYDK 77
+S D + LASGG+D ++ +W S + EH+ + + +SP + L S + D+
Sbjct: 140 WSYDNRELASGGNDNRLFVWNQHSTQPVLKYCEHTAAVKAIAWSPHLHGLLASGGGTADR 319
Query: 78 TVRVWD 83
+R W+
Sbjct: 320 CIRFWN 337
Score = 32.3 bits (72), Expect = 0.015
Identities = 14/60 (23%), Positives = 35/60 (58%), Gaps = 1/60 (1%)
Frame = +2
Query: 26 KLLASGGHDK-KVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDV 84
+L+++ G+ + ++++W ++ + ATL H+ + + SP + T + D+T+ W+V
Sbjct: 416 ELVSTHGYSQNQIIVWRYPTMSKLATLTGHTYRVLYLAISPDGQTIVTGAGDETLXFWNV 595
>TC14937 similar to UP|Q9FN19 (Q9FN19) Genomic DNA, chromosome 5, TAC
clone:K8K14 (AT5g67320/K8K14_4), partial (19%)
Length = 1089
Score = 36.6 bits (83), Expect = 8e-04
Identities = 20/58 (34%), Positives = 30/58 (51%)
Frame = +1
Query: 27 LLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWDV 84
+LAS D V LW + K +L H + V FSP+ LA+ S DK++ +W +
Sbjct: 52 VLASASFDSTVKLWDVEVGKLIYSLNGHRDGVYSVAFSPNGEYLASGSPDKSIHIWSL 225
Score = 28.1 bits (61), Expect = 0.28
Identities = 13/34 (38%), Positives = 22/34 (64%), Gaps = 9/34 (26%)
Frame = +1
Query: 61 VRFSPSMPR---------LATSSYDKTVRVWDVD 85
+R+SP+ P LA++S+D TV++WDV+
Sbjct: 1 IRWSPTGPGTSNPNKKLVLASASFDSTVKLWDVE 102
>TC18213 weakly similar to UP|Q9FIV2 (Q9FIV2) Similarity to pre-mRNA
splicing factor, partial (25%)
Length = 662
Score = 36.6 bits (83), Expect = 8e-04
Identities = 23/65 (35%), Positives = 34/65 (51%), Gaps = 1/65 (1%)
Frame = +1
Query: 20 HFSSDGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPR-LATSSYDKT 78
+FS DGK LASG D V L+ S K ++ DV F P +P +A+ S+D
Sbjct: 112 NFSRDGKKLASGSSDGSVYLYDYHSSKVVKKIKAFDQACIDVSFHPVIPNVIASCSWDGG 291
Query: 79 VRVWD 83
+ V++
Sbjct: 292 ILVFE 306
>TC19491 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Arabidopsis
thaliana;}, partial (39%)
Length = 527
Score = 36.2 bits (82), Expect = 0.001
Identities = 19/61 (31%), Positives = 31/61 (50%)
Frame = +1
Query: 24 DGKLLASGGHDKKVVLWYTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSYDKTVRVWD 83
D +LL + D V ++ + T+ H+ + V SP +AT S D+TVR+WD
Sbjct: 52 DPRLLFTASDDGNVHMYDAEGKALVGTMSGHASWVLCVDVSPDGGAIATGSSDRTVRLWD 231
Query: 84 V 84
+
Sbjct: 232 L 234
>BP069774
Length = 446
Score = 35.8 bits (81), Expect = 0.001
Identities = 22/69 (31%), Positives = 37/69 (52%), Gaps = 2/69 (2%)
Frame = +2
Query: 18 CSHFSSDGKLLASGGHDKKVVLW--YTDSLKQKATLEEHSLLITDVRFSPSMPRLATSSY 75
C F+ G LLA+G +D V+W T + + + + S IT + +S R+ S+
Sbjct: 113 CIAFNRRGTLLAAGCNDGSCVIWDFITRGIAKVLSDDGCSSPITSICWSKFGHRILVSAA 292
Query: 76 DKTVRVWDV 84
DK++ +WDV
Sbjct: 293 DKSLTLWDV 319
>TC12572 similar to PIR|T46032|T46032 WD-40 repeat regulatory protein tup1
homolog - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (51%)
Length = 770
Score = 35.0 bits (79), Expect = 0.002
Identities = 18/79 (22%), Positives = 39/79 (48%), Gaps = 1/79 (1%)
Frame = +3
Query: 6 LNSVRASTNKVVCSHFSSDGKLLASGGHDKKVVLWYTDSLK-QKATLEEHSLLITDVRFS 64
L + A ++ V F+ DG L+ S +D +W + K +++ + ++ V+FS
Sbjct: 18 LKVLPAHSDPVTAVDFNRDGSLIVSSSYDGLCRIWDASTGHCMKTLIDDENPPVSFVKFS 197
Query: 65 PSMPRLATSSYDKTVRVWD 83
P+ + + D +R+W+
Sbjct: 198PNAKFILVGTLDNNLRLWN 254
>BP052281
Length = 472
Score = 35.0 bits (79), Expect = 0.002
Identities = 23/66 (34%), Positives = 31/66 (46%), Gaps = 9/66 (13%)
Frame = +2
Query: 27 LLASGGHDKKVVLWYTD------SLKQKATLEEHSLLITDVRFSPSM---PRLATSSYDK 77
LL SG D ++LW D L K +L+ H + V S M P+L T DK
Sbjct: 23 LLVSGASDGLLILWSADHGQDSRELVPKLSLKAHDGGVVAVELSRVMGGTPQLITIGADK 202
Query: 78 TVRVWD 83
T+ +WD
Sbjct: 203TLAIWD 220
>TC18334 weakly similar to UP|Q9LHN3 (Q9LHN3) Emb|CAB63739.1
(AT3g18860/MCB22_3), partial (21%)
Length = 595
Score = 34.7 bits (78), Expect = 0.003
Identities = 24/83 (28%), Positives = 40/83 (47%), Gaps = 13/83 (15%)
Frame = +2
Query: 13 TNKVVCSHFSSDGKLL-------------ASGGHDKKVVLWYTDSLKQKATLEEHSLLIT 59
++KV+ H S G L+ ASGG D V++W + ++ TL+ H L +T
Sbjct: 254 SSKVLVGHSSFVGPLVWIPPNPELPQGGVASGGMDTLVLVWDLSTGEKAHTLKGHQLQVT 433
Query: 60 DVRFSPSMPRLATSSYDKTVRVW 82
+ F + ++S D T+R W
Sbjct: 434 SIAFDDG--DVVSASVDCTLRRW 496
Score = 26.9 bits (58), Expect = 0.63
Identities = 11/17 (64%), Positives = 13/17 (75%)
Frame = +2
Query: 70 LATSSYDKTVRVWDVDN 86
+ATSS DKTVR W +N
Sbjct: 191 IATSSRDKTVRFWSPEN 241
>BP044070
Length = 289
Score = 34.3 bits (77), Expect = 0.004
Identities = 24/65 (36%), Positives = 35/65 (52%), Gaps = 5/65 (7%)
Frame = +2
Query: 21 FSSDGKLLASGGHDKKVVLW-YTDS----LKQKATLEEHSLLITDVRFSPSMPRLATSSY 75
FSS+G+LLAS DK + + +T S L E H ++D+ FS L ++S
Sbjct: 95 FSSNGRLLASSSADKTLRTYGFTTSDSVTLSPMQLYEGHDXGVSDLAFSSDSRFLVSASD 274
Query: 76 DKTVR 80
DKT+R
Sbjct: 275 DKTLR 289
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.128 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,505,205
Number of Sequences: 28460
Number of extensions: 16005
Number of successful extensions: 156
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 130
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 154
length of query: 88
length of database: 4,897,600
effective HSP length: 64
effective length of query: 24
effective length of database: 3,076,160
effective search space: 73827840
effective search space used: 73827840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 48 (23.1 bits)
Medicago: description of AC146722.4


