

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146721.7 - phase: 0
(188 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
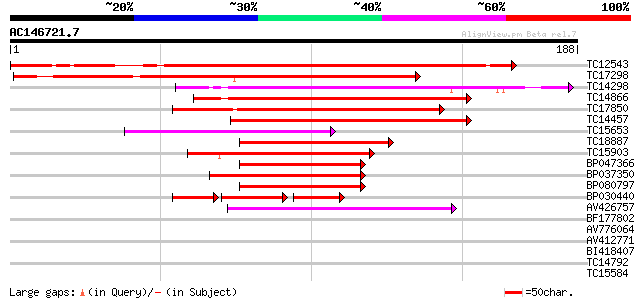
Score E
Sequences producing significant alignments: (bits) Value
TC12543 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor,... 238 4e-64
TC17298 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor,... 179 2e-46
TC14298 homologue to UP|Q8L5W2 (Q8L5W2) bZIP transcription facto... 82 6e-17
TC14866 similar to UP|Q8L5W2 (Q8L5W2) bZIP transcription factor ... 73 3e-14
TC17850 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor,... 67 2e-12
TC14457 similar to UP|Q8GTZ1 (Q8GTZ1) bZIP transcription factor,... 64 2e-11
TC15653 similar to UP|Q941L5 (Q941L5) bZIP transcription factor ... 52 5e-08
TC18887 similar to UP|Q94KA6 (Q94KA6) bZIP transcription factor ... 50 2e-07
TC15903 similar to UP|Q40724 (Q40724) Transcriptional activator ... 50 3e-07
BP047366 44 1e-05
BP037350 44 2e-05
BP080797 42 7e-05
BP030440 30 9e-05
AV426757 40 2e-04
BF177802 37 0.003
AV776064 36 0.005
AV412771 35 0.010
BI418407 32 0.089
TC14792 similar to UP|O73620 (O73620) Nuclear protein SDK2, part... 31 0.15
TC15584 28 0.75
>TC12543 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor, partial
(52%)
Length = 510
Score = 238 bits (608), Expect = 4e-64
Identities = 125/168 (74%), Positives = 141/168 (83%)
Frame = +3
Query: 1 MLPSNTSPYLQNYNNMIQNNNISSYQLQKFSNQIYGNYLNTPHQQFPDFNYSPQSSCISS 60
MLPSN PY NY MIQNN I ++Q Q+FSNQ++ Q PDF SPQSSCISS
Sbjct: 45 MLPSNPPPYPANYT-MIQNN-IPTFQFQRFSNQLF---------QVPDF--SPQSSCISS 185
Query: 61 NSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIE 120
NSTSDEADEQ LINERKHRRMISNRESARRSRMRKQ+HLDELWSQV+WLRNENHQL++
Sbjct: 186 NSTSDEADEQQQGLINERKHRRMISNRESARRSRMRKQRHLDELWSQVVWLRNENHQLLD 365
Query: 121 KLNHVSENHDQVVQENAQLKEEALELRQMIKDMQIHSPLIPSFSPLDD 168
KL+H SE+HDQVVQENAQLKEEA+ELRQM++DMQIHSP PSF+PL+D
Sbjct: 366 KLSHASESHDQVVQENAQLKEEAIELRQMLRDMQIHSP-CPSFAPLED 506
>TC17298 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor, partial
(39%)
Length = 484
Score = 179 bits (455), Expect = 2e-46
Identities = 95/138 (68%), Positives = 110/138 (78%), Gaps = 3/138 (2%)
Frame = +1
Query: 2 LPSNTSPYLQNYNNMIQNNNISSYQLQKFSNQIYGNYLNTPHQQFPDFNYSPQSSCISSN 61
LPS+ SP+ +MIQNNNI ++QLQKFSNQ +G N Q F DF+ SSCISSN
Sbjct: 91 LPSDPSPF-----SMIQNNNIPTFQLQKFSNQFFGYQNNL--QAFQDFSPQNSSSCISSN 249
Query: 62 STSDEADEQNLS---LINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQL 118
STSDEA++Q +INERKHRRMISNRESARRSRMRKQKHLDELWS V+ LRNENHQL
Sbjct: 250 STSDEAEDQQQQQSLIINERKHRRMISNRESARRSRMRKQKHLDELWSMVVCLRNENHQL 429
Query: 119 IEKLNHVSENHDQVVQEN 136
+EKLNH+SE+HD+VVQEN
Sbjct: 430 LEKLNHLSESHDRVVQEN 483
>TC14298 homologue to UP|Q8L5W2 (Q8L5W2) bZIP transcription factor ATB2,
partial (62%)
Length = 1585
Score = 82.0 bits (201), Expect = 6e-17
Identities = 61/149 (40%), Positives = 86/149 (56%), Gaps = 17/149 (11%)
Frame = +3
Query: 56 SCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNEN 115
S + NS S+E D Q +++++RK +RMISNRESARRSRMRKQKHLD+L SQV LR EN
Sbjct: 642 SSLLQNSGSEE-DLQ--AVMDQRKRKRMISNRESARRSRMRKQKHLDDLASQVTQLRKEN 812
Query: 116 HQLIEKLNHVSENHDQVVQENAQLKEEALE-------LRQMIKDMQIHSPLI------PS 162
Q+I +N ++ + V EN+ L+ + E L ++I+ + +S PS
Sbjct: 813 QQIITSVNITTQQYLSVEAENSVLRAQMGELSNRLESLNEIIEVLNANSAAFGTTFTEPS 992
Query: 163 ----FSPLDDTYLRDDSSNNSISSSMDLL 187
F+PL +YL N I S D+L
Sbjct: 993 NGFFFNPLSMSYL-----NQQIMPSADML 1064
>TC14866 similar to UP|Q8L5W2 (Q8L5W2) bZIP transcription factor ATB2,
partial (60%)
Length = 948
Score = 72.8 bits (177), Expect = 3e-14
Identities = 39/92 (42%), Positives = 60/92 (64%)
Frame = +3
Query: 62 STSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEK 121
S+ E D Q ++++RK++R SNRESARRSRMRKQ HL++L SQV L EN +++
Sbjct: 15 SSGSEGDLQ--MVMDQRKNKRKQSNRESARRSRMRKQSHLEDLTSQVTQLTKENGEILTN 188
Query: 122 LNHVSENHDQVVQENAQLKEEALELRQMIKDM 153
+N S+ + V EN+ L+ + EL Q ++ +
Sbjct: 189 INITSQQYQNVETENSILRAQMGELSQRLQSL 284
>TC17850 similar to UP|Q9AT29 (Q9AT29) bZIP transcription factor, partial
(23%)
Length = 508
Score = 67.0 bits (162), Expect = 2e-12
Identities = 37/90 (41%), Positives = 55/90 (61%)
Frame = -1
Query: 55 SSCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNE 114
SSC+ +N T EA+E + + + A RSRM+KQKHL+ LWS+V+ L E
Sbjct: 307 SSCLDNNGTYAEANELQFR*-SAKGSTGE*CLTQFAHRSRMKKQKHLNGLWSKVVRLGIE 131
Query: 115 NHQLIEKLNHVSENHDQVVQENAQLKEEAL 144
NH L+ KLNHVSE+HD+ + L+++ +
Sbjct: 130 NHNLVLKLNHVSESHDRALNRRHALRKKLM 41
Score = 26.6 bits (57), Expect = 2.9
Identities = 14/25 (56%), Positives = 16/25 (64%)
Frame = -3
Query: 131 QVVQENAQLKEEALELRQMIKDMQI 155
Q Q A LKEE +L QM+ DMQI
Sbjct: 83 QSPQ*KACLKEETYDLPQMVADMQI 9
>TC14457 similar to UP|Q8GTZ1 (Q8GTZ1) bZIP transcription factor, partial
(50%)
Length = 645
Score = 63.9 bits (154), Expect = 2e-11
Identities = 32/80 (40%), Positives = 54/80 (67%)
Frame = +1
Query: 74 LINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQVV 133
+++++K +RM SNRESARRSRMRKQ+HL+ + +QV L+ EN+Q+ + ++ + V
Sbjct: 7 VMDQKKRKRMQSNRESARRSRMRKQEHLEGMSAQVEQLKKENNQISTNIGVTTQMYLNVE 186
Query: 134 QENAQLKEEALELRQMIKDM 153
ENA L+ + EL ++ +
Sbjct: 187 AENAILRVQMAELSNRLQSL 246
>TC15653 similar to UP|Q941L5 (Q941L5) bZIP transcription factor BZI-4,
partial (25%)
Length = 511
Score = 52.4 bits (124), Expect = 5e-08
Identities = 28/70 (40%), Positives = 40/70 (57%)
Frame = +2
Query: 39 LNTPHQQFPDFNYSPQSSCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQ 98
L+ P+ F + S S E I++RK +RM+SNRESARRSRMRKQ
Sbjct: 287 LDLPNPNFTKNKKNSPEGMASVQRGSSSGSEGGDLAIDDRKRKRMVSNRESARRSRMRKQ 466
Query: 99 KHLDELWSQV 108
K L++L +++
Sbjct: 467 KQLEDLTAEM 496
>TC18887 similar to UP|Q94KA6 (Q94KA6) bZIP transcription factor 6, partial
(64%)
Length = 944
Score = 50.1 bits (118), Expect = 2e-07
Identities = 26/51 (50%), Positives = 33/51 (63%)
Frame = +3
Query: 77 ERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSE 127
ER+ RRMI NRESA RSR RKQ + EL +V L+ EN +L +K + E
Sbjct: 780 ERRQRRMIKNRESAARSRARKQAYTMELEQEVAKLKEENEELQKKQEEIME 932
>TC15903 similar to UP|Q40724 (Q40724) Transcriptional activator protein
(RITA-1 protein), partial (21%)
Length = 603
Score = 49.7 bits (117), Expect = 3e-07
Identities = 32/68 (47%), Positives = 43/68 (63%), Gaps = 6/68 (8%)
Frame = +1
Query: 60 SNSTSDEAD------EQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRN 113
S+ SDE D EQ+ + + ++ RR +SNRESARRSR RKQ HL EL +QV L+
Sbjct: 394 SSDRSDEDDDEAGPCEQSNNPQDVKRLRRKVSNRESARRSRRRKQAHLAELETQVEKLKL 573
Query: 114 ENHQLIEK 121
EN L ++
Sbjct: 574 ENATLYKQ 597
>BP047366
Length = 535
Score = 44.3 bits (103), Expect = 1e-05
Identities = 23/42 (54%), Positives = 28/42 (65%)
Frame = -3
Query: 77 ERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQL 118
ER+ +RMI NRESA RSR RKQ + EL +V L EN +L
Sbjct: 488 ERRQKRMIKNRESAARSRARKQAYTQELEIKVSRLEEENERL 363
>BP037350
Length = 495
Score = 43.9 bits (102), Expect = 2e-05
Identities = 24/52 (46%), Positives = 32/52 (61%)
Frame = -2
Query: 67 ADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQL 118
A E + ER+ +RMI NRESA RSR RK + +EL ++V L EN +L
Sbjct: 407 APEDMVGKTVERRQKRMIKNRESAARSRARKHAYTNEL*NKVSRLEEENERL 252
>BP080797
Length = 498
Score = 42.0 bits (97), Expect = 7e-05
Identities = 23/42 (54%), Positives = 29/42 (68%)
Frame = -2
Query: 77 ERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQL 118
ER+ RR I NRESA RSR RKQ + +EL S+V L +N +L
Sbjct: 449 ERRLRRKIKNRESAARSRARKQAYHNELVSKVTRLEQDNIKL 324
>BP030440
Length = 536
Score = 29.6 bits (65), Expect(3) = 9e-05
Identities = 12/22 (54%), Positives = 18/22 (81%)
Frame = +1
Query: 71 NLSLINERKHRRMISNRESARR 92
++ +I++RKHRRM+SN E A R
Sbjct: 277 SVQIIDKRKHRRMMSNTEFAHR 342
Score = 27.3 bits (59), Expect(3) = 9e-05
Identities = 11/17 (64%), Positives = 15/17 (87%)
Frame = +3
Query: 95 MRKQKHLDELWSQVLWL 111
M+KQKHL+ LWS+V+ L
Sbjct: 345 MKKQKHLNGLWSKVVRL 395
Score = 22.3 bits (46), Expect(3) = 9e-05
Identities = 9/15 (60%), Positives = 12/15 (80%)
Frame = +2
Query: 55 SSCISSNSTSDEADE 69
SSC+ +N T DEA+E
Sbjct: 230 SSCLGNN*TYDEANE 274
>AV426757
Length = 415
Score = 40.4 bits (93), Expect = 2e-04
Identities = 21/76 (27%), Positives = 45/76 (58%)
Frame = +1
Query: 73 SLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQV 132
++++ ++ +R+++NR+SA+RSR+RK +++ EL V L+ E L ++ + +
Sbjct: 31 TVVDPKRVKRILANRQSAQRSRVRKLQYISELERSVTTLQTEVSALSPRVAFLDHQRLIL 210
Query: 133 VQENAQLKEEALELRQ 148
+N+ LK+ L Q
Sbjct: 211 NVDNSTLKQRIAALAQ 258
>BF177802
Length = 481
Score = 36.6 bits (83), Expect = 0.003
Identities = 20/70 (28%), Positives = 37/70 (52%)
Frame = +1
Query: 31 SNQIYGNYLNTPHQQFPDFNYSPQSSCISSNSTSDEADEQNLSLINERKHRRMISNRESA 90
++ ++ + H FN S +S S+S A + + ++++ +R++ NRESA
Sbjct: 277 ASSVFQKQSHQHHSMLSSFNTSFDAS---SSSGEKIAQQDRVYAFSDQRRKRILKNRESA 447
Query: 91 RRSRMRKQKH 100
RSR RKQ +
Sbjct: 448 LRSRARKQAY 477
>AV776064
Length = 418
Score = 35.8 bits (81), Expect = 0.005
Identities = 18/34 (52%), Positives = 22/34 (63%)
Frame = -1
Query: 67 ADEQNLSLINERKHRRMISNRESARRSRMRKQKH 100
A E + ER+ +RMI NRESA RSR RKQ +
Sbjct: 385 APEDMVEKTVERRQKRMIKNRESAARSRARKQAY 284
>AV412771
Length = 405
Score = 34.7 bits (78), Expect = 0.010
Identities = 17/36 (47%), Positives = 25/36 (69%)
Frame = +3
Query: 55 SSCISSNSTSDEADEQNLSLINERKHRRMISNRESA 90
+S +SS S S+ +L + NERK +R++SNRESA
Sbjct: 297 ASIMSSGSASEGGGGADLHVDNERKRKRILSNRESA 404
>BI418407
Length = 422
Score = 31.6 bits (70), Expect = 0.089
Identities = 28/92 (30%), Positives = 40/92 (43%)
Frame = -3
Query: 22 ISSYQLQKFSNQIYGNYLNTPHQQFPDFNYSPQSSCISSNSTSDEADEQNLSLINERKHR 81
IS L FS ++ G+ NT FP + S SSN S + L L E R
Sbjct: 276 ISIKSLNGFSCKMKGSTRNTLFS-FPTTTITRLSEYSSSNIPSSV---RGL*LSREPMER 109
Query: 82 RMISNRESARRSRMRKQKHLDELWSQVLWLRN 113
R I+ ++S +D+ WS +WL+N
Sbjct: 108 RRINETGITKQSMNIMNWKIDDRWSLCVWLKN 13
>TC14792 similar to UP|O73620 (O73620) Nuclear protein SDK2, partial (5%)
Length = 492
Score = 30.8 bits (68), Expect = 0.15
Identities = 22/70 (31%), Positives = 40/70 (56%), Gaps = 3/70 (4%)
Frame = +3
Query: 91 RRSRMRKQKHLDELWSQVLWLRN--ENHQLIEKLNHVSENHDQVVQENAQLKEEALE-LR 147
+++++RK K L S W +N + HQ + LNH + N ++ +E+ +L+EE +E L
Sbjct: 246 QKNKIRKPKI--SLKSTTPWKKNKTQKHQQQQSLNH-NNNEEEEEEEDLELEEEPVERLL 416
Query: 148 QMIKDMQIHS 157
+ Q+HS
Sbjct: 417 EPFTREQLHS 446
>TC15584
Length = 797
Score = 28.5 bits (62), Expect = 0.75
Identities = 14/40 (35%), Positives = 21/40 (52%)
Frame = -2
Query: 4 SNTSPYLQNYNNMIQNNNISSYQLQKFSNQIYGNYLNTPH 43
S+ +PYL M+++ N + K SN+ NY TPH
Sbjct: 742 SSKAPYLATTLKMMRHTNDKKQVMMKTSNKPLINYKYTPH 623
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.126 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,379,655
Number of Sequences: 28460
Number of extensions: 45203
Number of successful extensions: 299
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 290
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 294
length of query: 188
length of database: 4,897,600
effective HSP length: 85
effective length of query: 103
effective length of database: 2,478,500
effective search space: 255285500
effective search space used: 255285500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146721.7


