

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146719.6 - phase: 0 /pseudo
(312 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
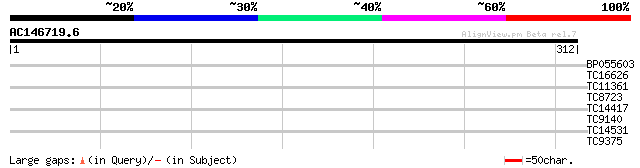
BP055603 29 0.90
TC16626 similar to UP|Q9LX30 (Q9LX30) Protein kinase like, parti... 28 1.5
TC11361 27 3.4
TC8723 homologue to UP|TBB1_LUPAL (P37392) Tubulin beta-1 chain,... 27 4.4
TC14417 homologue to UP|AAR23806 (AAR23806) Initiation factor eI... 27 5.8
TC9140 homologue to UP|TBB3_PEA (P29502) Tubulin beta-3 chain (F... 26 7.6
TC14531 26 7.6
TC9375 homologue to PIR|JA0049|JA0049 Tubulin beta-2 chain - soy... 26 7.6
>BP055603
Length = 467
Score = 29.3 bits (64), Expect = 0.90
Identities = 17/47 (36%), Positives = 24/47 (50%), Gaps = 6/47 (12%)
Frame = -1
Query: 224 LCLSVLQMHLVCLWST*IAFFMHFWT---VSWLF---LLMTF*FTPR 264
LC S++Q+ ++ WST I WT V WL LL+ +PR
Sbjct: 233 LCYSIVQIMMLVSWSTTIVTLTRXWTPFVVDWLLSIVLLLAVCVSPR 93
>TC16626 similar to UP|Q9LX30 (Q9LX30) Protein kinase like, partial (6%)
Length = 625
Score = 28.5 bits (62), Expect = 1.5
Identities = 15/35 (42%), Positives = 23/35 (64%)
Frame = +2
Query: 223 KLCLSVLQMHLVCLWST*IAFFMHFWTVSWLFLLM 257
+LC+SVLQ++L+ S I F F+ +WL L+M
Sbjct: 407 RLCISVLQLYLIPPLS--ILLFSVFFFSTWLVLMM 505
>TC11361
Length = 549
Score = 27.3 bits (59), Expect = 3.4
Identities = 13/29 (44%), Positives = 15/29 (50%)
Frame = -2
Query: 208 CRRQLLERGMHIMSIKLCLSVLQMHLVCL 236
CRR + R + KLCL LQ HL L
Sbjct: 290 CRRSIWSRIWELCMRKLCLKHLQQHLALL 204
>TC8723 homologue to UP|TBB1_LUPAL (P37392) Tubulin beta-1 chain, complete
Length = 1689
Score = 26.9 bits (58), Expect = 4.4
Identities = 12/20 (60%), Positives = 14/20 (70%)
Frame = -2
Query: 275 WFCKCSKRRSFMLNYLSVNS 294
W+C S+ RSFML SVNS
Sbjct: 1391 WYCWYSETRSFMLLSASVNS 1332
>TC14417 homologue to UP|AAR23806 (AAR23806) Initiation factor eIF4A-15,
complete
Length = 1563
Score = 26.6 bits (57), Expect = 5.8
Identities = 14/37 (37%), Positives = 22/37 (58%)
Frame = +3
Query: 254 FLLMTF*FTPRLRKNMLSI*RWFCKCSKRRSFMLNYL 290
FLLM +P + K+M+++ +W CK +FML L
Sbjct: 159 FLLMGKISSPHMMKSMIALTQWDCKKIF*EAFMLMVL 269
>TC9140 homologue to UP|TBB3_PEA (P29502) Tubulin beta-3 chain (Fragment),
partial (34%)
Length = 797
Score = 26.2 bits (56), Expect = 7.6
Identities = 12/20 (60%), Positives = 13/20 (65%)
Frame = -3
Query: 275 WFCKCSKRRSFMLNYLSVNS 294
W+C S RSFML SVNS
Sbjct: 396 WYCWYSDTRSFMLLSASVNS 337
>TC14531
Length = 535
Score = 26.2 bits (56), Expect = 7.6
Identities = 7/20 (35%), Positives = 14/20 (70%)
Frame = -1
Query: 274 RWFCKCSKRRSFMLNYLSVN 293
+W+C+CSK+ S + N ++
Sbjct: 457 KWYCQCSKQTSILANITHIS 398
>TC9375 homologue to PIR|JA0049|JA0049 Tubulin beta-2 chain - soybean
{Glycine max;} , partial (15%)
Length = 699
Score = 26.2 bits (56), Expect = 7.6
Identities = 12/20 (60%), Positives = 13/20 (65%)
Frame = -1
Query: 275 WFCKCSKRRSFMLNYLSVNS 294
W+C S RSFML SVNS
Sbjct: 153 WYCWYSATRSFMLLSASVNS 94
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.378 0.167 0.656
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,090,959
Number of Sequences: 28460
Number of extensions: 95853
Number of successful extensions: 1967
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1468
Number of HSP's successfully gapped in prelim test: 49
Number of HSP's that attempted gapping in prelim test: 474
Number of HSP's gapped (non-prelim): 1561
length of query: 312
length of database: 4,897,600
effective HSP length: 90
effective length of query: 222
effective length of database: 2,336,200
effective search space: 518636400
effective search space used: 518636400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146719.6


