

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146719.12 + phase: 0 /pseudo
(758 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
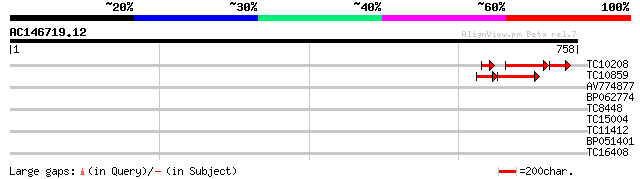
TC10208 weakly similar to UP|Q8TUQ7 (Q8TUQ7) TraB family protein... 65 3e-15
TC10859 52 4e-09
AV774877 32 0.29
BP062774 29 3.2
TC8448 weakly similar to UP|WSC4_YEAST (P38739) Cell wall integr... 28 5.4
TC15004 similar to UP|ATX1_ARATH (Q9LT02) Potential cation-trans... 28 7.1
TC11412 28 7.1
BP051401 27 9.3
TC16408 similar to GB|AAM47476.1|21360559|AY113173 AT5g04990/MUG... 27 9.3
>TC10208 weakly similar to UP|Q8TUQ7 (Q8TUQ7) TraB family protein, partial
(4%)
Length = 566
Score = 65.5 bits (158), Expect(3) = 3e-15
Identities = 35/57 (61%), Positives = 38/57 (66%)
Frame = +2
Query: 664 FKTWMGYVLSLKRK*LSG*KFF*AVKESFVKGTWVLVSLLSKLACSIKKQLHPCLAV 720
FKTW GY L LK L G*K F* VKESF KG W+LV LLSK+AC K L P + V
Sbjct: 245 FKTWKGYALPLKTNLLCG*KLF*PVKESFEKGLWILVFLLSKIACPPLKLLLPNVVV 415
Score = 29.6 bits (65), Expect(3) = 3e-15
Identities = 15/28 (53%), Positives = 21/28 (74%)
Frame = +1
Query: 722 ILLD*HRWNNGLL*IRKTYMTS*NPLHQ 749
I+L+*HR N GLL*IR + + *N L++
Sbjct: 424 IMLN*HRCNYGLL*IRNMHKSG*NSLYK 507
Score = 23.5 bits (49), Expect(3) = 3e-15
Identities = 12/18 (66%), Positives = 12/18 (66%)
Frame = +3
Query: 631 TFLLDCCTCSI*FVDV*C 648
TF LD CSI*FV V C
Sbjct: 141 TFFLDG*ICSI*FVGVWC 194
>TC10859
Length = 533
Score = 51.6 bits (122), Expect(2) = 4e-09
Identities = 33/58 (56%), Positives = 39/58 (66%), Gaps = 2/58 (3%)
Frame = +1
Query: 653 MLIS*P**IANFKTWMGYVLSLKRK*--LSG*KFF*AVKESFVKGTWVLVSLLSKLAC 708
M+I * *I FKTW Y ++L+ K LS *K F*AVKES KG W+L LLSK+AC
Sbjct: 244 MVIK*RE*IE*FKTWKSYAIALQLKINLLSR*KSF*AVKESSEKGLWILGFLLSKIAC 417
Score = 26.6 bits (57), Expect(2) = 4e-09
Identities = 14/29 (48%), Positives = 18/29 (61%)
Frame = +3
Query: 624 VSFQGCRTFLLDCCTCSI*FVDV*CWHSW 652
+ Q +TFLLDC SI*+V V C S+
Sbjct: 156 IGIQRNQTFLLDCYIHSI*YVGVRC*QSF 242
>AV774877
Length = 338
Score = 32.3 bits (72), Expect = 0.29
Identities = 19/51 (37%), Positives = 27/51 (52%), Gaps = 2/51 (3%)
Frame = -1
Query: 5 PSPLQPAMLMGSPSFPNALAWSHDNL--IAAASGHFVTILRPDLPNGPRGL 53
PSPL + L+ P + L + NL + S HF+ +R D P GPRG+
Sbjct: 338 PSPLPDSSLLFPPLDASHLLTVYQNLG*VDGVSVHFLHFIRVDGPPGPRGV 186
>BP062774
Length = 557
Score = 28.9 bits (63), Expect = 3.2
Identities = 17/38 (44%), Positives = 24/38 (62%), Gaps = 1/38 (2%)
Frame = -3
Query: 603 NGSHYHFWDLI*IFLLRKCYHVSFQGCRTFL-LDCCTC 639
+ S + +W LI I +L+K YHV+ + CRT L D TC
Sbjct: 183 HNSLFPYWYLIYIAVLKK-YHVTLKFCRTCLAFDKVTC 73
>TC8448 weakly similar to UP|WSC4_YEAST (P38739) Cell wall integrity and
stress response component 4 precursor, partial (5%)
Length = 1049
Score = 28.1 bits (61), Expect = 5.4
Identities = 13/41 (31%), Positives = 23/41 (55%)
Frame = +3
Query: 256 YIRLLFLGLPCCM*HLNFIPILIEMPLSVFLLLVGSLVKFL 296
+ RL FLG PCC L ++ +++ L ++ + SL+ L
Sbjct: 360 HFRLPFLGTPCCWRFLGWVTLVLFKWLHLYF*CIQSLLDML 482
>TC15004 similar to UP|ATX1_ARATH (Q9LT02) Potential cation-transporting
ATPase , partial (9%)
Length = 830
Score = 27.7 bits (60), Expect = 7.1
Identities = 16/53 (30%), Positives = 27/53 (50%)
Frame = +2
Query: 451 KYQFFLTCPVAMIQPVYDYIKYAFFPCSLQEIHLNHVLA*LCPLEILSLQLFI 503
+Y+F+ TCP ++ Y + + F + IH H+L+ L + S Q FI
Sbjct: 578 RYEFYNTCPPNHLRCFY*HRSFVLFKFT---IHSTHLLSRELSLNVKSKQHFI 727
>TC11412
Length = 500
Score = 27.7 bits (60), Expect = 7.1
Identities = 15/36 (41%), Positives = 20/36 (54%)
Frame = +3
Query: 254 QCYIRLLFLGLPCCM*HLNFIPILIEMPLSVFLLLV 289
QCYIR FLG C M I ++ + LS+F L +
Sbjct: 279 QCYIRNFFLGYTCFM---MMIMVVYQNSLSIF*LAI 377
>BP051401
Length = 507
Score = 27.3 bits (59), Expect = 9.3
Identities = 13/23 (56%), Positives = 14/23 (60%), Gaps = 1/23 (4%)
Frame = -2
Query: 159 YTCIKA-TTTKKFPIRWGELHGS 180
Y C K+ K FPIRW ELH S
Sbjct: 419 YYCTKSHRLPKTFPIRWDELHES 351
>TC16408 similar to GB|AAM47476.1|21360559|AY113173 AT5g04990/MUG13_15
{Arabidopsis thaliana;}, partial (12%)
Length = 543
Score = 27.3 bits (59), Expect = 9.3
Identities = 16/39 (41%), Positives = 18/39 (46%)
Frame = +2
Query: 616 FLLRKCYHVSFQGCRTFLLDCCTCSI*FVDV*CWHSWML 654
FL C H S C F C CS+ FV +* SW L
Sbjct: 188 FL*WHCSHNSSFVCLEFYQCSCNCSVLFVQM*DRKSWSL 304
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.360 0.162 0.604
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,798,132
Number of Sequences: 28460
Number of extensions: 255388
Number of successful extensions: 2494
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 1832
Number of HSP's successfully gapped in prelim test: 60
Number of HSP's that attempted gapping in prelim test: 626
Number of HSP's gapped (non-prelim): 1945
length of query: 758
length of database: 4,897,600
effective HSP length: 97
effective length of query: 661
effective length of database: 2,136,980
effective search space: 1412543780
effective search space used: 1412543780
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146719.12


