

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146706.4 + phase: 0
(478 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
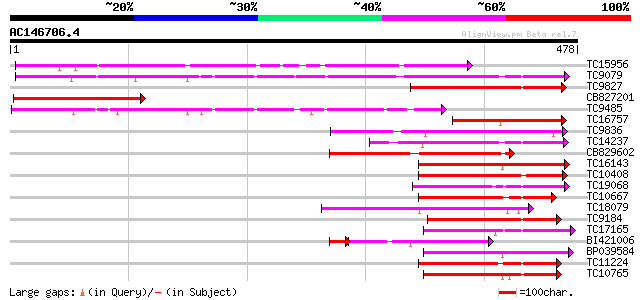
Score E
Sequences producing significant alignments: (bits) Value
TC15956 weakly similar to UP|LGT_CITUN (Q9MB73) Limonoid UDP-glu... 221 3e-58
TC9079 weakly similar to UP|Q8W4G1 (Q8W4G1) UDP-glucose glucosyl... 202 1e-52
TC9827 weakly similar to UP|Q8S9A2 (Q8S9A2) Glucosyltransferase-... 144 2e-35
CB827201 138 2e-33
TC9485 similar to UP|Q8S999 (Q8S999) Glucosyltransferase-10, par... 133 6e-32
TC16757 weakly similar to UP|Q9SBQ2 (Q9SBQ2) Anthocyanin 5-O-glu... 128 2e-30
TC9836 weakly similar to UP|Q9LXV0 (Q9LXV0) Glucosyltransferase-... 125 1e-29
TC14237 weakly similar to UP|AAR06913 (AAR06913) UDP-glycosyltra... 117 5e-27
CB829602 116 7e-27
TC16143 similar to UP|Q7XZD0 (Q7XZD0) Isoflavonoid glucosyltrans... 112 2e-25
TC10408 weakly similar to UP|Q9FUJ6 (Q9FUJ6) UDP-glucosyltransfe... 108 1e-24
TC19068 weakly similar to UP|AAR06913 (AAR06913) UDP-glycosyltra... 105 2e-23
TC10667 similar to UP|Q9M051 (Q9M051) Glucuronosyl transferase-l... 102 1e-22
TC18079 weakly similar to UP|UFO5_MANES (Q40287) Flavonol 3-O-gl... 100 7e-22
TC9184 weakly similar to UP|HQGT_RAUSE (Q9AR73) Hydroquinone glu... 99 1e-21
TC17165 similar to UP|Q9SMG6 (Q9SMG6) Betanidin-5-O-glucosyltran... 98 3e-21
BI421006 87 4e-21
BP039584 91 5e-19
TC11224 weakly similar to UP|Q8S996 (Q8S996) Glucosyltransferase... 89 1e-18
TC10765 homologue to UP|Q9ZWQ5 (Q9ZWQ5) UDP-glycose:flavonoid gl... 89 2e-18
>TC15956 weakly similar to UP|LGT_CITUN (Q9MB73) Limonoid
UDP-glucosyltransferase (Limonoid glucosyltransferase)
(Limonoid GTase) (LGTase) , partial (65%)
Length = 1169
Score = 221 bits (562), Expect = 3e-58
Identities = 142/397 (35%), Positives = 209/397 (51%), Gaps = 12/397 (3%)
Frame = +2
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATT------IHLYSRLINKPTIP----G 55
H L++++P QGHINP L+ K L + G+ VT ATT + + + +K P
Sbjct: 38 HILLVSFPAQGHINPLLRLAKCLAAKGSSVTLATTQTAGKSMRTATNITDKAPTPISDGS 217
Query: 56 LSFATFSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLIL 115
L F F DG S ++ Y+ + G + + +I N +C+I L
Sbjct: 218 LKFEFFDDGLGPDDDSIRG-NLSLYLQQLELVGRQQILKMIKEHADSNRQISCIINNPFL 394
Query: 116 SWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCL-ISLPGLSFSL 174
W VA E +PS LLWIQ++ VF +Y YFH + +T SK++ + LP S L
Sbjct: 395 PWVVDVAEEQGIPSALLWIQSSAVFTAYYSYFH---NLVTFPSKEDPLADVQLPSTSIVL 565
Query: 175 KSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKM 234
K ++P FL S TY F + EQ + LN+ +LV+T EE E D ++ + K +
Sbjct: 566 KHCEIPDFLHPSCTYPFLGTLILEQFKNLNKTFC--ILVDTYEELEHDFIDYLS--KTFL 733
Query: 235 I-PIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLS 293
I P+GPL S+ + GD ++ DD I+WL+SK + SVVYVSFG++ L
Sbjct: 734 IRPVGPLFKSS-----QAKSAAIRGDFMK---SDDCIEWLNSKPQSSVVYVSFGSIVYLP 889
Query: 294 KRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQ 353
+ Q++EIA L S SFLWV++ + V L E G++V+W Q
Sbjct: 890 QEQVDEIAFGLKKSQVSFLWVMKPPPEVTGLQPHV----LPYGFLEETGERGRVVQWSPQ 1057
Query: 354 VEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQW 390
EVL+H S+ CF+THCGWNS++E+L SGVP++ FP W
Sbjct: 1058EEVLAHPSVSCFLTHCGWNSSMEALTSGVPVLTFPAW 1168
>TC9079 weakly similar to UP|Q8W4G1 (Q8W4G1) UDP-glucose
glucosyltransferase, partial (65%)
Length = 1585
Score = 202 bits (513), Expect = 1e-52
Identities = 146/487 (29%), Positives = 233/487 (46%), Gaps = 20/487 (4%)
Frame = +1
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINK------PTIPGLSFA 59
H ++ PLQGHINP Q K L G +TF T + + RLI +P F
Sbjct: 19 HAVLTPSPLQGHINPLFQLAKLLHLRGFHITFVHTEYNHKRLIKSRGPNALDGLPDFVFE 198
Query: 60 TFSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENH-----PFTCLIYTLI 114
T DG +DG +D+ S + +++ + P TCL+ L+
Sbjct: 199 TIPDGLEDGDGEVS-QDLYSLCDSIRNKCHLPFQDLLAKLHRSASAGLVPPVTCLVSDLL 375
Query: 115 LSWAPKVAHELHLPSTLLWIQAATVFDIFYYYFH---------EHGDYITNKSKDETCLI 165
+++ + A +L LP LL +A+ F F ++ + Y+TN D T +
Sbjct: 376 MTFTIQAAQQLQLPILLLCPASASTFMSFLHFQTLLDKGVIPLKDESYLTNGYLD-TKVD 552
Query: 166 SLPGLSFSLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALN 225
+PG+ + + +DLP F+ ++ L + E ++ NT E E D L+
Sbjct: 553 WIPGMQ-NFRLKDLPDFIRTTDPNDIMLEFMVEVADRAKRA--SAIVFNTFNELERDVLS 723
Query: 226 KVDVGKIKMIPIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVS 285
+ + PIGP PS S G ++ + D+K +QWL+SK+ SVVYV+
Sbjct: 724 ALSTMLPSLYPIGPF-PSFLNQAPQNQLASLGSNLWKEDTK--CLQWLESKEPGSVVYVN 894
Query: 286 FGTLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNG 345
FG++ V+S Q+ E A L + FLW+IR + LS + E + G
Sbjct: 895 FGSITVMSPEQLLEFAWGLANGKKPFLWIIRPDLVADGSAI------LSPKFVDETSNRG 1056
Query: 346 KIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWK 405
I WC Q +VL+H S+G F+THCGWNST+ES+ +GVPM+ +P DQ TN + I + W
Sbjct: 1057LIASWCPQEKVLNHPSVGGFLTHCGWNSTIESICAGVPMLCWPFVADQPTNCRYICNEWD 1236
Query: 406 TGLRMEHDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNL 465
G+ ++ + VK EE+ + +M G+KG+++R+ + K A + GG S NL
Sbjct: 1237IGIEVDTN----VKREEVENLIHELM-VGDKGKKMRQRTMELKKKAEEDTRPGGCSYMNL 1401
Query: 466 RSYLNDI 472
+ ++
Sbjct: 1402DKLIKEV 1422
>TC9827 weakly similar to UP|Q8S9A2 (Q8S9A2) Glucosyltransferase-7
(Fragment), partial (42%)
Length = 611
Score = 144 bits (364), Expect = 2e-35
Identities = 65/131 (49%), Positives = 94/131 (71%)
Frame = +2
Query: 339 LENNMNGKIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAK 398
+E G IV WC Q+ VL H++LGCF+THCGWNSTLES+ GVP++A P WTD TN K
Sbjct: 8 VETTEKGLIVTWCPQLLVLKHQALGCFLTHCGWNSTLESVSLGVPVIAMPLWTDPVTNGK 187
Query: 399 LIEDVWKTGLRMEHDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEG 458
LI DVWKTG+R DE+ +V+ E I+ C++ ++ + EK E++ NA KW +LA+ ++ EG
Sbjct: 188 LIVDVWKTGVRAVADEKEIVRRETIKNCIKEII-ETEKINEIKSNAIKWMNLAKDSLAEG 364
Query: 459 GSSNRNLRSYL 469
G S++N+ ++
Sbjct: 365 GRSDKNIAEFV 397
>CB827201
Length = 374
Score = 138 bits (348), Expect = 2e-33
Identities = 67/112 (59%), Positives = 82/112 (72%), Gaps = 1/112 (0%)
Frame = +2
Query: 4 NHHFLIITYPLQGHINPALQFTKRLISLGAK-VTFATTIHLYSRLINKPTIPGLSFATFS 62
+H FL+I YP QGHINP LQF KRLIS GA+ VT TT+H Y R+ NKPT+P L+ FS
Sbjct: 38 HHRFLLIIYPAQGHINPTLQFAKRLISSGAEHVTLCTTLHTYRRISNKPTLPNLTLLPFS 217
Query: 63 DGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLI 114
DG+DDG K D D Y +E RRG+EF+T++ILS+ QE PFTCLIYTL+
Sbjct: 218 DGFDDGFKLTPDTDYSLYATELKRRGTEFVTDLILSAAQEGKPFTCLIYTLL 373
>TC9485 similar to UP|Q8S999 (Q8S999) Glucosyltransferase-10, partial (75%)
Length = 1195
Score = 133 bits (335), Expect = 6e-32
Identities = 119/389 (30%), Positives = 183/389 (46%), Gaps = 22/389 (5%)
Frame = +1
Query: 2 AQNHHFLIITYPLQGHINPALQFTKRL-ISLGAKVTFATTIHLYSRLINKPT------IP 54
A+ H + I YP QGHINP L+ K L G VTF T + + RL+ +P
Sbjct: 79 AKKPHVVCIPYPAQGHINPMLKLAKLLHFKGGFHVTFVNTEYNHKRLLKSRGPDSLNGLP 258
Query: 55 GLSFATFSDGYDDGQKSFGDEDIVSYMSEFTRRGS----EFLTNIILSSKQENHPFTCLI 110
F T DG + +DI S TR+ + L + + + P TC++
Sbjct: 259 SFRFETIPDGLPETDVDV-TQDIPSLCIS-TRKTCLPHFKKLLSKLNDVSSDVPPVTCIV 432
Query: 111 YTLILSWAPKVAHELHLPSTLLWIQAATVFDIFYYYFH--EHGDYITNKSKD------ET 162
+S+ A EL++P L W +A F + Y E G S D ET
Sbjct: 433 SDGCMSFTLDAAIELNIPEVLFWTTSACGFMGYVQYRELIEKGIIPLKDSSDITNGYLET 612
Query: 163 CLISLPGLSFSLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELD 222
+ LPG+ +++ +DLPSFL ++ L L + Q + +++NT + E D
Sbjct: 613 TIEWLPGMK-NIRLKDLPSFLRTTDPNDKMLDFLTGECQRALKA--SAIILNTFDALEHD 783
Query: 223 ALNKVDVGKIKMIPIGPLIPSAFLDGKDPTD---NSFGGDVVRVDSKDDYIQWLDSKDEK 279
L + IGPL L KD TD NS G ++ + DS + ++WLD+K+
Sbjct: 784 VLEAFSSILPPVYSIGPL----HLLIKDVTDKNLNSLGSNLWKEDS--ECLKWLDTKEPN 945
Query: 280 SVVYVSFGTLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREEL 339
SVVYV+FG++ V++ QM E A L +S +FLWVIR L K + ++ ++
Sbjct: 946 SVVYVNFGSITVMTSEQMVEFAWGLANSNKTFLWVIR-PDLVAGKHAVLPEEFVAA---- 1110
Query: 340 ENNMNGKIVKWCSQVEVLSHRSLGCFMTH 368
N G++ W Q +VL+H ++G F+TH
Sbjct: 1111-TNDRGRLSSWTPQEDVLTHPAIGGFLTH 1194
>TC16757 weakly similar to UP|Q9SBQ2 (Q9SBQ2) Anthocyanin
5-O-glucosyltransferase, partial (19%)
Length = 638
Score = 128 bits (321), Expect = 2e-30
Identities = 62/98 (63%), Positives = 81/98 (82%), Gaps = 2/98 (2%)
Frame = +2
Query: 374 TLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEH--DEEGMVKVEEIRKCLEVVM 431
T+ESL SGVPM AFP+++DQ TNAK++EDVWK G+RM+H +E+G+V+ EIRKCL+ VM
Sbjct: 2 TMESLXSGVPMXAFPRFSDQMTNAKMVEDVWKIGVRMDHSVNEDGVVEGGEIRKCLDAVM 181
Query: 432 GKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYL 469
G GEK EE+RRNA+K+K LA A KEGGSS +NL ++L
Sbjct: 182 GNGEKAEEVRRNAEKFKGLAMEAGKEGGSSEKNLMAFL 295
>TC9836 weakly similar to UP|Q9LXV0 (Q9LXV0) Glucosyltransferase-like
protein, partial (55%)
Length = 1286
Score = 125 bits (315), Expect = 1e-29
Identities = 71/211 (33%), Positives = 119/211 (55%), Gaps = 11/211 (5%)
Frame = +1
Query: 271 QWLDSKDEKSVVYVSFGTLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDD 330
+WLD+K SV++VSFG++ +S QM ++A AL SG +F+WV+R +
Sbjct: 385 KWLDTKPSNSVLFVSFGSMNTISASQMMQLATALDRSGRNFIWVVRPPIGFDI------N 546
Query: 331 DELSCREELENNMNGKIVK-------WCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVP 383
E E L G++ K W QVE+LSH ++ F++HCGWNS LESL GVP
Sbjct: 547 SEFRAEEWLPEGFLGRVAKKGLVVHDWAPQVEILSHGAVSAFLSHCGWNSVLESLSHGVP 726
Query: 384 MVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVEEIRKCLEVVMGKGEKGEELRRN 443
++ +P +Q N K++E+ + + + V+ E++ + +E+VM + E G ++R+N
Sbjct: 727 ILGWPMAAEQFFNCKMLEEELGVCVEVARGKRCEVRHEDLVEKIELVMNEAESGVKIRKN 906
Query: 444 AKKWKDLARAAVKE----GGSSNRNLRSYLN 470
A +++ R AV++ GSS R + +L+
Sbjct: 907 AGNIREMIRDAVRDEDGYKGSSVRAIDEFLS 999
>TC14237 weakly similar to UP|AAR06913 (AAR06913) UDP-glycosyltransferase
85A8, partial (27%)
Length = 999
Score = 117 bits (292), Expect = 5e-27
Identities = 59/170 (34%), Positives = 99/170 (57%), Gaps = 2/170 (1%)
Frame = +2
Query: 304 LLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGK--IVKWCSQVEVLSHRS 361
+ +S FLW++R + + +D ++ +E + + G+ I WC Q EVLSH S
Sbjct: 197 IANSKVPFLWILRPDVV-------MGEDSINVPQEFLDEIKGRGYIASWCFQEEVLSHPS 355
Query: 362 LGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVE 421
+G F+THCGWNST+E + +G+P++ +P +++Q TN++ W+ G+ + HD VK +
Sbjct: 356 IGAFLTHCGWNSTIEGISAGLPLICWPFFSEQHTNSRYACTTWEIGMEVNHD----VKRD 523
Query: 422 EIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLND 471
EI + +M KGEKG+E+R+ +WK A A GGSS + + +
Sbjct: 524 EITTLVNEMM-KGEKGKEMRKKGLEWKKKAIEATGLGGSSYNDFHKLIKE 670
>CB829602
Length = 542
Score = 116 bits (291), Expect = 7e-27
Identities = 56/156 (35%), Positives = 95/156 (60%)
Frame = +1
Query: 270 IQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVD 329
++WLDS++ SV+YV+FG++ ++ +Q+ E+A + +S F+WVIR ++ + +
Sbjct: 97 LKWLDSQEPSSVLYVNFGSVINMTPQQLVELAWGIANSKKKFIWVIRPDLVEGEASIVLP 276
Query: 330 DDELSCREELENNMNGKIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQ 389
+ ++ G ++ WC Q +VL H +LG F+THCGWNST+ES+ SGVP++ P
Sbjct: 277 EIVAETKDR------GIMLSWCPQEQVLKHSALGGFLTHCGWNSTIESISSGVPLICSPF 438
Query: 390 WTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVEEIRK 425
+ DQ N++ I W G+ M D+ VK +E+ K
Sbjct: 439 FNDQFVNSRYICSEWDFGMEMNSDD---VKRDEVEK 537
>TC16143 similar to UP|Q7XZD0 (Q7XZD0) Isoflavonoid glucosyltransferase,
partial (30%)
Length = 604
Score = 112 bits (279), Expect = 2e-25
Identities = 56/138 (40%), Positives = 91/138 (65%), Gaps = 10/138 (7%)
Frame = +1
Query: 345 GKIVK-WCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDV 403
G I++ W QV +L HR++G F+THCGWNST+E++ +GVPM+ +P +Q N KLI +V
Sbjct: 10 GMIIRGWVPQVVILGHRAVGAFVTHCGWNSTVEAVSAGVPMITWPVVGEQFYNEKLITEV 189
Query: 404 WKTGLRMEHDE---------EGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAA 454
G+ + +E E +V E I K + +M G++GE++RR A+++ + AR A
Sbjct: 190 RGIGVEVGAEECCLMGFDKREKLVSRESIEKAVRRLMDGGDEGEQIRRRAQEYGEKARQA 369
Query: 455 VKEGGSSNRNLRSYLNDI 472
V+EGGSS++NL + ++D+
Sbjct: 370 VEEGGSSHKNLTALIDDL 423
>TC10408 weakly similar to UP|Q9FUJ6 (Q9FUJ6) UDP-glucosyltransferase HRA25,
partial (19%)
Length = 571
Score = 108 bits (271), Expect = 1e-24
Identities = 51/126 (40%), Positives = 80/126 (63%)
Frame = +1
Query: 345 GKIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVW 404
GK+VKW Q ++L+H ++ CF++HCGWNST+E + + VP + +P TDQ N I DVW
Sbjct: 16 GKVVKWGPQKKILNHPAIACFISHCGWNSTIEGVHASVPFLCWPFSTDQFLNKSFICDVW 195
Query: 405 KTGLRMEHDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRN 464
K GL ++ DE G++ EIR +E V+ E+L+ + K K++ + EGG S+ N
Sbjct: 196 KIGLGLDKDENGIIPKGEIRTKVEQVI----VNEDLKARSLKLKEITINNLAEGGQSSNN 363
Query: 465 LRSYLN 470
L+ ++N
Sbjct: 364 LKKFIN 381
>TC19068 weakly similar to UP|AAR06913 (AAR06913) UDP-glycosyltransferase
85A8, partial (26%)
Length = 565
Score = 105 bits (261), Expect = 2e-23
Identities = 55/133 (41%), Positives = 80/133 (59%)
Frame = +1
Query: 340 ENNMNGKIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKL 399
E + G I WCSQ VL+H S+G F+THCGWNST+ES+ +GVPM+ +P + DQ TN +
Sbjct: 37 EISSRGLIASWCSQEHVLNHPSIGGFLTHCGWNSTIESISAGVPMLCWPFFADQLTNCRS 216
Query: 400 IEDVWKTGLRMEHDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGG 459
I + W G M+ D G K EE+ K + +M GEKG+++R+ + K A + GG
Sbjct: 217 ICNEWDIG--MQIDTNG--KREEVEKLINELM-VGEKGKKMRQKTMELKKRAEEDTRPGG 381
Query: 460 SSNRNLRSYLNDI 472
S NL + ++
Sbjct: 382 CSYMNLDQVIKEV 420
>TC10667 similar to UP|Q9M051 (Q9M051) Glucuronosyl transferase-like
protein, partial (20%)
Length = 824
Score = 102 bits (255), Expect = 1e-22
Identities = 52/117 (44%), Positives = 72/117 (61%)
Frame = +1
Query: 345 GKIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVW 404
G VKW Q +VL H ++G F THCGWNSTLES+ GVPM+ P + DQ NAK + DVW
Sbjct: 7 GTSVKWAPQEQVLKHPAIGAFWTHCGWNSTLESVCEGVPMICSPCFGDQKVNAKYVSDVW 186
Query: 405 KTGLRMEHDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSS 461
K G+++++ EI K + ++M G++ E+R N K K+ A + EGGSS
Sbjct: 187 KVGVQLQNKPGR----GEIEKTISLLM-LGDEAYEIRGNILKLKEKANVCLSEGGSS 342
>TC18079 weakly similar to UP|UFO5_MANES (Q40287) Flavonol
3-O-glucosyltransferase 5 (UDP-glucose flavonoid
3-O-glucosyltransferase 5) , partial (33%)
Length = 889
Score = 100 bits (248), Expect = 7e-22
Identities = 65/191 (34%), Positives = 104/191 (54%), Gaps = 13/191 (6%)
Frame = +3
Query: 264 DSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQ 323
+ +D+ ++WLD + SV+YVSFG+ + + QM EIA L S F+WV+R
Sbjct: 45 EDQDEILRWLDKQPAGSVIYVSFGSGGTMPQGQMTEIALGLELSQHRFVWVVRPPNEADA 224
Query: 324 KEEEVD-DDELSCREELENNMN-----GKIVK-WCSQVEVLSHRSLGCFMTHCGWNSTLE 376
+E++ E MN G +V W Q E+L H + G F+T CGWNS LE
Sbjct: 225 SATFFGFGNEMALNYLPEGFMNRTREFGVVVPMWAPQAEILGHPATGGFVTPCGWNSVLE 404
Query: 377 SLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMV----KVEEIRKCL--EVV 430
S+ +GVPMVA+P +++Q NA ++ + +R++ E G+V V+ +R+ + E
Sbjct: 405 SVLNGVPMVAWPLYSEQKMNAYMLSEELGVAVRVKVAEGGVVCRDQVVDMVRRVMVSEEG 584
Query: 431 MGKGEKGEELR 441
MG ++ EL+
Sbjct: 585 MGMKDRVRELK 617
>TC9184 weakly similar to UP|HQGT_RAUSE (Q9AR73) Hydroquinone
glucosyltransferase (Arbutin synthase) , partial (26%)
Length = 643
Score = 99.4 bits (246), Expect = 1e-21
Identities = 49/113 (43%), Positives = 75/113 (66%)
Frame = +3
Query: 353 QVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEH 412
Q +VLSH S+G F++HCGWNS LES+ +GVP+VA+P + +Q NA L+ + K LR +
Sbjct: 9 QAKVLSHASIGGFLSHCGWNSVLESVANGVPLVAWPLYAEQRMNAVLVTEDVKVALRPKV 188
Query: 413 DEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNL 465
E G+V++ EI + + +M +GE+G++LR K K+ A +KE GSS +
Sbjct: 189 GENGLVELVEIARVVRTLM-QGEEGKKLRCKMKDLKEAAATTLKENGSSTNQI 344
>TC17165 similar to UP|Q9SMG6 (Q9SMG6) Betanidin-5-O-glucosyltransferase,
partial (24%)
Length = 546
Score = 97.8 bits (242), Expect = 3e-21
Identities = 53/136 (38%), Positives = 82/136 (59%), Gaps = 8/136 (5%)
Frame = +1
Query: 350 WCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGL- 408
W QV +L H ++G F+THCGWNSTLE + +GVPMV +P +Q N KL+ +V K G+
Sbjct: 13 WAPQVLILEHEAIGAFVTHCGWNSTLEGVAAGVPMVTWPIAAEQFYNEKLVTEVLKIGVP 192
Query: 409 -------RMEHDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSS 461
RM + +G VK + + K ++ +M +GE+ EE+R K A+ AV+EGGSS
Sbjct: 193 VGAKKWGRMVLEGDG-VKSDAVEKGVKRIM-EGEEAEEMRNRVKVLSQQAQWAVEEGGSS 366
Query: 462 NRNLRSYLNDIACITQ 477
+L + + ++ + Q
Sbjct: 367 YSDLNALIEELGMLRQ 414
>BI421006
Length = 519
Score = 87.0 bits (214), Expect(2) = 4e-21
Identities = 49/129 (37%), Positives = 69/129 (52%), Gaps = 6/129 (4%)
Frame = +2
Query: 286 FGTLAVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCR------EEL 339
F + + Q+E+IA L S F+WV+RD K + D D++ R E+
Sbjct: 101 FWFFSTFIEEQIEQIANGLEQSKQKFIWVLRDA----DKGDIFDGDKVKERGLPKGFEKR 268
Query: 340 ENNMNGKIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKL 399
M + W Q+E+LSH S G FM+HCGWNS +ES+ GVP+ A+P +DQ N L
Sbjct: 269 VEGMGLVVRDWAPQLEILSHPSTGGFMSHCGWNSCIESMSMGVPIAAWPMHSDQPRNTVL 448
Query: 400 IEDVWKTGL 408
I +V K L
Sbjct: 449 ITEVLKVAL 475
Score = 31.2 bits (69), Expect(2) = 4e-21
Identities = 11/18 (61%), Positives = 17/18 (94%)
Frame = +3
Query: 270 IQWLDSKDEKSVVYVSFG 287
++WLD +++KSV+YVSFG
Sbjct: 54 MEWLDRQEQKSVLYVSFG 107
>BP039584
Length = 545
Score = 90.5 bits (223), Expect = 5e-19
Identities = 44/134 (32%), Positives = 77/134 (56%), Gaps = 8/134 (5%)
Frame = -3
Query: 350 WCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLR 409
W Q+ +L H ++G +THCGWN+TLES+ +G+PM P + +Q N KL+ DV G+
Sbjct: 537 WAPQLLILEHPAIGGVVTHCGWNTTLESVIAGLPMATMPLFAEQFYNEKLLVDVLGVGVS 358
Query: 410 MEHDE--------EGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSS 461
+ + + +VK E I K + ++MG GE+ E+RR ++ D A+ ++ GGSS
Sbjct: 357 IGMKKWRNWNVVGDEIVKRENIVKAISLLMGGGEEALEMRRRVRELSDAAKKTIQPGGSS 178
Query: 462 NRNLRSYLNDIACI 475
++ +++ +
Sbjct: 177 YNKVKGLFDELKAL 136
>TC11224 weakly similar to UP|Q8S996 (Q8S996) Glucosyltransferase-13
(Fragment), partial (19%)
Length = 596
Score = 89.4 bits (220), Expect = 1e-18
Identities = 47/121 (38%), Positives = 74/121 (60%)
Frame = +3
Query: 345 GKIVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVW 404
GK+V W Q+E+L H S+G F+TH GWNS LE + GVPM+ P + DQ N ++ +
Sbjct: 33 GKVVSWAPQMEILKHASVGVFLTHGGWNSVLECIVGGVPMIGRPFFGDQRINIWMLAKLL 212
Query: 405 KTGLRMEHDEEGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRN 464
G+ +++ G++ E I K L+ M GE+G+ +R+ + K++A AV+ GSS +N
Sbjct: 213 GVGVGLQN---GVLAKETILKTLKSTM-TGEEGKVMRQKMAELKEMAWKAVEPDGSSTKN 380
Query: 465 L 465
L
Sbjct: 381 L 383
>TC10765 homologue to UP|Q9ZWQ5 (Q9ZWQ5) UDP-glycose:flavonoid
glycosyltransferase, partial (19%)
Length = 542
Score = 88.6 bits (218), Expect = 2e-18
Identities = 50/125 (40%), Positives = 76/125 (60%), Gaps = 9/125 (7%)
Frame = +2
Query: 350 WCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLR 409
W Q +L+H + G F+THCGWN+ +E++ +GVPM+ P ++DQ N KLI +V G+
Sbjct: 8 WAPQPLILNHLATGGFLTHCGWNAVVEAISAGVPMITMPGFSDQYYNEKLITEVHGFGVE 187
Query: 410 MEHDE------EGMVKV---EEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGS 460
+ E +G KV E I K ++ +M GE+G E+R AK+ +D A AV++GGS
Sbjct: 188 VGAAEWSISPYDGKKKVVSGERIEKAVKSLM--GEEGAEIRSKAKEVQDKAWKAVQQGGS 361
Query: 461 SNRNL 465
S +L
Sbjct: 362 SYNSL 376
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,406,199
Number of Sequences: 28460
Number of extensions: 118623
Number of successful extensions: 784
Number of sequences better than 10.0: 95
Number of HSP's better than 10.0 without gapping: 753
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 758
length of query: 478
length of database: 4,897,600
effective HSP length: 94
effective length of query: 384
effective length of database: 2,222,360
effective search space: 853386240
effective search space used: 853386240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146706.4


