

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146705.12 - phase: 0 /pseudo
(811 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
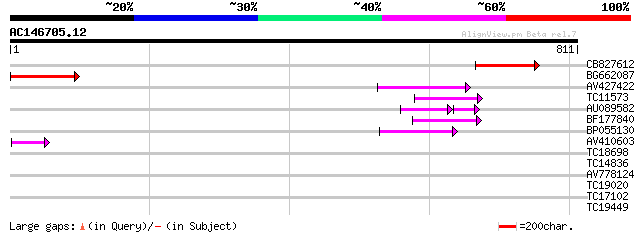
Score E
Sequences producing significant alignments: (bits) Value
CB827612 124 5e-29
BG662087 88 5e-18
AV427422 74 9e-14
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 64 1e-10
AU089582 47 5e-10
BF177840 59 4e-09
BP055130 49 2e-06
AV410603 47 2e-05
TC18698 37 0.010
TC14836 similar to UP|O24322 (O24322) Cysteine proteinase precur... 30 1.2
AV778124 29 2.6
TC19020 28 7.6
TC17102 UP|CAE45588 (CAE45588) Papain-like cysteine proteinase-l... 27 9.9
TC19449 similar to PIR|E86300|E86300 protein F3O9.30 [imported] ... 27 9.9
>CB827612
Length = 277
Score = 124 bits (312), Expect = 5e-29
Identities = 57/92 (61%), Positives = 73/92 (78%)
Frame = +1
Query: 667 SCHLPVELEHKAYWAIRNLNLDPNLAGDKRKLQLNELEELRMDAYENARIYKERTKTWHD 726
+CHLPVE+EH+AYWA+++ N + AG +RKLQL +LEELR+ AYE++RIYKE+TK HD
Sbjct: 1 ACHLPVEIEHRAYWAVKSCNXEMQQAGIERKLQLQQLEELRLGAYESSRIYKEKTKHXHD 180
Query: 727 KKIIKRHFKSGDLVLLFNSRLKLFPGKLRSRW 758
K I ++ F G VLLFNSRLKL GKLRS+W
Sbjct: 181 KMISRKEFSVGQQVLLFNSRLKLMAGKLRSKW 276
>BG662087
Length = 373
Score = 88.2 bits (217), Expect = 5e-18
Identities = 41/99 (41%), Positives = 63/99 (63%)
Frame = +1
Query: 1 NKATRKDHFPLPFIDQMLERLAKHSHFCYLDGYSGFFQIPIHPNDQEKTTFTCPFGTFAY 60
NKA KD +PLP ID++++ + + +D YSG+ QI +HP+D++KT F + Y
Sbjct: 52 NKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDEDKTAFMTARVNYCY 231
Query: 61 RRMPFGLCNAPATFQRCMMSIFSDFVEKIMEVFMDDFSV 99
+ +PFGL NA AT+Q M +F D V + MEV++D+ V
Sbjct: 232 QTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIV 348
>AV427422
Length = 417
Score = 73.9 bits (180), Expect = 9e-14
Identities = 41/135 (30%), Positives = 72/135 (52%), Gaps = 1/135 (0%)
Frame = +1
Query: 526 PSSFGNQYILVAVDYVSKWVEAIA-SPTNDAQVVIKMFKKVIFPRFGVPRVVISDGGSHF 584
P S G + +LV VD +SK+ + A+V+ +F + + GVP ++SD F
Sbjct: 13 PKSKGYEAVLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDPLF 192
Query: 585 ISRHFEKLLQKLGVRHKIATPYHPQTSGQVEVSNRQIKAILEKTVSTSRTDWSNKLDDAL 644
+S +++L + G + K++T YHP++ GQ EV NR ++ L ++ W++ + A
Sbjct: 193 MSNFWKELFKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPWAE 372
Query: 645 WAYRTAYKTPIGMTP 659
+ Y T+Y G TP
Sbjct: 373 YWYNTSYHVSTGQTP 417
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 63.9 bits (154), Expect = 1e-10
Identities = 30/98 (30%), Positives = 54/98 (54%)
Frame = +2
Query: 579 DGGSHFISRHFEKLLQKLGVRHKIATPYHPQTSGQVEVSNRQIKAILEKTVSTSRTDWSN 638
+ G F S+ + +G++ + ++ HPQT+GQ E +N+ I +++ + + W +
Sbjct: 8 ENGIQFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWID 187
Query: 639 KLDDALWAYRTAYKTPIGMTPFKLVYGKSCHLPVELEH 676
+L LW+Y T ++ I TPF L YG LPVE+ +
Sbjct: 188 ELPIVLWSYNTMPQSSIKETPF*LTYGADTMLPVEISN 301
>AU089582
Length = 383
Score = 47.0 bits (110), Expect(2) = 5e-10
Identities = 26/74 (35%), Positives = 39/74 (52%)
Frame = +2
Query: 560 KMFKKVIFPRFGVPRVVISDGGSHFISRHFEKLLQKLGVRHKIATPYHPQTSGQVEVSNR 619
K++ I GVP +ISD G+ F S + LG R K++T +HPQT GQ E + +
Sbjct: 26 KIYLDEIVSLHGVPVSIISDRGAQFTSHFWRSFQTALGTRLKMSTAFHPQTDGQSERTIQ 205
Query: 620 QIKAILEKTVSTSR 633
++ +L V R
Sbjct: 206 ILEDMLRACVXDLR 247
Score = 34.3 bits (77), Expect(2) = 5e-10
Identities = 13/37 (35%), Positives = 21/37 (56%)
Frame = +3
Query: 636 WSNKLDDALWAYRTAYKTPIGMTPFKLVYGKSCHLPV 672
W L +AY +Y++ I M PF+ +YG+ C P+
Sbjct: 255 WDQYLSLMEFAYNNSYRSSI*MAPFEALYGRRCRSPI 365
>BF177840
Length = 410
Score = 58.5 bits (140), Expect = 4e-09
Identities = 30/99 (30%), Positives = 50/99 (50%)
Frame = +2
Query: 576 VISDGGSHFISRHFEKLLQKLGVRHKIATPYHPQTSGQVEVSNRQIKAILEKTVSTSRTD 635
++SD + FIS + L K+G + +T HPQT GQ EV N+ + +L + +
Sbjct: 17 IVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNLKM 196
Query: 636 WSNKLDDALWAYRTAYKTPIGMTPFKLVYGKSCHLPVEL 674
W L +AY + +PF++VYG + P++L
Sbjct: 197 WETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDL 313
>BP055130
Length = 567
Score = 49.3 bits (116), Expect = 2e-06
Identities = 32/112 (28%), Positives = 51/112 (44%), Gaps = 1/112 (0%)
Frame = +2
Query: 530 GNQYILVAVDYVSKWVEAIASPTN-DAQVVIKMFKKVIFPRFGVPRVVISDGGSHFISRH 588
G I+V VD +SK+ N + V + F I G+PR ++SD F S
Sbjct: 179 GFTVIIVVVDRLSKYGHFAPHRANYTSSQVAETFVSTIVKLHGMPRAIVSDRDKAFTSAF 358
Query: 589 FEKLLQKLGVRHKIATPYHPQTSGQVEVSNRQIKAILEKTVSTSRTDWSNKL 640
++ + G +++ YHPQT GQ E N+ ++ L V + W + L
Sbjct: 359 WKHFFKLHGTTLNMSSSYHPQTDGQTEALNKCLELYLRCFVHETPRLWVSYL 514
>AV410603
Length = 162
Score = 46.6 bits (109), Expect = 2e-05
Identities = 22/54 (40%), Positives = 31/54 (56%)
Frame = +1
Query: 3 ATRKDHFPLPFIDQMLERLAKHSHFCYLDGYSGFFQIPIHPNDQEKTTFTCPFG 56
+T KD FP+P +D++L+ L F LD SG+ QI + P D+ KT F G
Sbjct: 1 STVKDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTHHG 162
>TC18698
Length = 808
Score = 37.4 bits (85), Expect = 0.010
Identities = 21/57 (36%), Positives = 31/57 (53%)
Frame = -2
Query: 46 QEKTTFTCPFGTFAYRRMPFGLCNAPATFQRCMMSIFSDFVEKIMEVFMDDFSVHGS 102
++KTT + Y+ MP GL N T+QR M IF + K +EV+++D V S
Sbjct: 807 KKKTTLKINRVNYYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKSS 637
>TC14836 similar to UP|O24322 (O24322) Cysteine proteinase precursor ,
partial (92%)
Length = 1416
Score = 30.4 bits (67), Expect = 1.2
Identities = 10/21 (47%), Positives = 16/21 (75%)
Frame = +2
Query: 664 YGKSCHLPVELEHKAYWAIRN 684
YGK+ + P+ ++KAYW I+N
Sbjct: 953 YGKAGYAPIRFKNKAYWIIKN 1015
>AV778124
Length = 515
Score = 29.3 bits (64), Expect = 2.6
Identities = 14/35 (40%), Positives = 19/35 (54%), Gaps = 1/35 (2%)
Frame = -3
Query: 403 NYQQKKKFFN-DLKHYYWDEPYLFRRGSDGIFRRC 436
N +Q KF N + HYYWDE +L + S + C
Sbjct: 117 NEKQLFKFININTDHYYWDENHLMSKLSSSLMLIC 13
>TC19020
Length = 438
Score = 27.7 bits (60), Expect = 7.6
Identities = 13/37 (35%), Positives = 19/37 (51%)
Frame = +3
Query: 412 NDLKHYYWDEPYLFRRGSDGIFRRCIPENEVSSILTH 448
+ L H+ W + +RR + RRC+PE E I H
Sbjct: 225 SSLSHFIWGQAQ-YRREWQWLLRRCLPEVEAHIIDIH 332
>TC17102 UP|CAE45588 (CAE45588) Papain-like cysteine proteinase-like protein
1, complete
Length = 1086
Score = 27.3 bits (59), Expect = 9.9
Identities = 8/21 (38%), Positives = 13/21 (61%)
Frame = +1
Query: 664 YGKSCHLPVELEHKAYWAIRN 684
YG + P+ ++ K YW I+N
Sbjct: 907 YGSESYAPIRMKQKPYWIIKN 969
>TC19449 similar to PIR|E86300|E86300 protein F3O9.30 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(24%)
Length = 600
Score = 27.3 bits (59), Expect = 9.9
Identities = 15/43 (34%), Positives = 21/43 (47%), Gaps = 2/43 (4%)
Frame = -3
Query: 467 ILHSGFWWPSLFKDVHLFISKCDKCQR--TGSITKRNEMPLNN 507
++ S WP+L + IS C +C R GSI N P +N
Sbjct: 154 MMPSSLKWPALSLLFQVLISPCRECTRFHNGSIRAMNMKPASN 26
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.140 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,716,354
Number of Sequences: 28460
Number of extensions: 211190
Number of successful extensions: 940
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 936
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 940
length of query: 811
length of database: 4,897,600
effective HSP length: 98
effective length of query: 713
effective length of database: 2,108,520
effective search space: 1503374760
effective search space used: 1503374760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146705.12


