

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146705.10 - phase: 0
(146 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
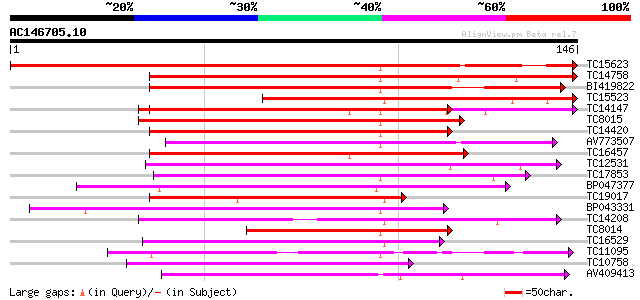
Score E
Sequences producing significant alignments: (bits) Value
TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding p... 209 2e-55
TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding p... 133 1e-32
BI419822 119 2e-28
TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-bindi... 103 1e-23
TC14147 similar to UP|ROC1_NICSY (Q08935) 29 kDa ribonucleoprote... 83 2e-17
TC8015 similar to UP|ROC4_NICSY (P19683) 31 kDa ribonucleoprotei... 81 6e-17
TC14420 similar to UP|ROC3_NICSY (P19682) 28 kDa ribonucleoprote... 80 1e-16
AV773507 76 2e-15
TC16457 similar to UP|Q41124 (Q41124) Chloroplast RNA binding pr... 74 8e-15
TC12531 similar to UP|Q9ASQ8 (Q9ASQ8) At1g22910/F19G10_13, parti... 74 8e-15
TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, ... 71 8e-14
BP047377 70 1e-13
TC19017 similar to UP|Q8S2X6 (Q8S2X6) Glycine-rich RNA binding p... 69 4e-13
BP043331 67 9e-13
TC14208 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, pa... 62 5e-11
TC8014 similar to GB|CAA41023.1|21309|SORBP28 28kD RNA binding p... 61 7e-11
TC16529 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, p... 61 7e-11
TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 61 7e-11
TC10758 61 9e-11
AV409413 60 1e-10
>TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding protein
PsGRBP, partial (88%)
Length = 683
Score = 209 bits (531), Expect = 2e-55
Identities = 109/147 (74%), Positives = 121/147 (82%), Gaps = 1/147 (0%)
Frame = +1
Query: 1 MAFCNKIGNLLRQGATQSTQAPVSSMLNYLRHMSSSKLFIGGLSYNVDDQSLRDAFTTYG 60
MAF NK+GN+LRQGA Q +QAPV+SMLNY+R MSSSKLFIGGLSY VDD+SL DAF+++G
Sbjct: 79 MAFLNKVGNVLRQGAAQRSQAPVASMLNYIRCMSSSKLFIGGLSYGVDDKSLEDAFSSFG 258
Query: 61 DVVEARVITDRETGRSRGFGFVNFTSEESATSAL-SMDGQDLNGRNIRVSYANDRQAGGP 119
V EARVI DR+TGRSRGFGFV+F SEESA+SAL SMDGQDLNGRNIRVSYANDR A GP
Sbjct: 259 TVAEARVIVDRDTGRSRGFGFVSFDSEESASSALSSMDGQDLNGRNIRVSYANDRPA-GP 435
Query: 120 RPGGGYGGGGGYSGGGGGYSGGGGGGW 146
R GGGYG GGGY GGGW
Sbjct: 436 RGGGGYGTGGGYG------DRNSGGGW 498
>TC14758 similar to UP|O24601 (O24601) Glycine-rich RNA binding protein 2,
partial (93%)
Length = 725
Score = 133 bits (334), Expect = 1e-32
Identities = 66/128 (51%), Positives = 87/128 (67%), Gaps = 18/128 (14%)
Frame = +1
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-S 95
+ F+GGL++ D +L AF++YG++++++V+ DRETGRSRGFGFV FTSEE+ SA+
Sbjct: 85 RCFVGGLAWTTDSHTLEQAFSSYGEIIDSKVVNDRETGRSRGFGFVTFTSEEAMRSAIEG 264
Query: 96 MDGQDLNGRNIRVSYANDR---QAGGPRPGGGYGGGG--------------GYSGGGGGY 138
M+G +L+GRNI V+ A R GG R GGGYGGGG GY GGGGGY
Sbjct: 265 MNGNELDGRNITVNEAQARGGGGGGGGRGGGGYGGGGGRREGGYGGGGGSRGYGGGGGGY 444
Query: 139 SGGGGGGW 146
GGGGGG+
Sbjct: 445 GGGGGGGY 468
>BI419822
Length = 380
Score = 119 bits (298), Expect = 2e-28
Identities = 58/108 (53%), Positives = 78/108 (71%), Gaps = 1/108 (0%)
Frame = +2
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-S 95
+ F+GGL++ D+++L AF+ YG++VE++VI DRETGRSRGFGFV F SE++ A+ +
Sbjct: 80 RCFVGGLAWATDNEALEKAFSPYGEIVESKVINDRETGRSRGFGFVTFASEQAMKDAIEA 259
Query: 96 MDGQDLNGRNIRVSYANDRQAGGPRPGGGYGGGGGYSGGGGGYSGGGG 143
M+GQ+L+GRNI V+ A R G GGGG Y GGGGGY GGGG
Sbjct: 260 MNGQNLDGRNITVNEAQSR--------GSSGGGGRYGGGGGGYGGGGG 379
>TC15523 similar to UP|GRPA_MAIZE (P10979) Glycine-rich RNA-binding,
abscisic acid-inducible protein, complete
Length = 972
Score = 103 bits (257), Expect = 1e-23
Identities = 53/86 (61%), Positives = 63/86 (72%), Gaps = 5/86 (5%)
Frame = +2
Query: 66 RVITDRETGRSRGFGFVNFTSEESATSALS-MDGQDLNGRNIRVSYANDRQAGGPRPGGG 124
+V+ DRETGRSRGFGFV F SE+S A+ M+GQD++GRN+ V+ A R GG GGG
Sbjct: 446 QVVNDRETGRSRGFGFVTFASEQSMKDAIEGMNGQDIDGRNVTVNEAQARGRGGGGGGGG 625
Query: 125 YGGG---GGYSGGGGG-YSGGGGGGW 146
YGGG GGY GGGG YSGGGGGG+
Sbjct: 626 YGGGRREGGYGGGGGSRYSGGGGGGY 703
Score = 37.4 bits (85), Expect = 0.001
Identities = 13/31 (41%), Positives = 24/31 (76%)
Frame = +2
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARV 67
+ F+GGL++ D + L AF+TYG++V+++V
Sbjct: 77 RCFVGGLAWATDSEGLEKAFSTYGEIVDSKV 169
>TC14147 similar to UP|ROC1_NICSY (Q08935) 29 kDa ribonucleoprotein A,
chloroplast precursor (CP29A), partial (63%)
Length = 1602
Score = 82.8 bits (203), Expect = 2e-17
Identities = 41/82 (50%), Positives = 59/82 (71%), Gaps = 1/82 (1%)
Frame = +3
Query: 34 SSSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSA 93
S +++ +G L++ VD+ +L F G VV+A+VI DRE+GRSRGFGFV F+S + SA
Sbjct: 732 SENRVHVGNLAWGVDNAALESLFREQGRVVDAKVIYDRESGRSRGFGFVTFSSPDEVNSA 911
Query: 94 L-SMDGQDLNGRNIRVSYANDR 114
+ S+DG DLNGR I+VS A+ +
Sbjct: 912 IRSLDGADLNGRAIKVSQADSK 977
Score = 70.1 bits (170), Expect = 1e-13
Identities = 42/120 (35%), Positives = 59/120 (49%), Gaps = 10/120 (8%)
Frame = +3
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTS-EESATSALS 95
K+F+G L ++VD L + F G+V VI D+ TG SRGF FV +S E+ +A
Sbjct: 348 KVFVGNLPFSVDSAQLAELFQDAGNVEVVEVIYDKMTGNSRGFAFVTMSSAAEAEAAAQQ 527
Query: 96 MDGQDLNGRNIRVSYA----NDRQAGGPRP-----GGGYGGGGGYSGGGGGYSGGGGGGW 146
+ +L GR +RV+ N+ + P GG Y GG GY GGG G+
Sbjct: 528 FNNYELEGRALRVNSGPPPKNENRGFNENPRFRNNSFNRGGSDSYRGGSDGYRGGGSDGY 707
>TC8015 similar to UP|ROC4_NICSY (P19683) 31 kDa ribonucleoprotein,
chloroplast precursor, partial (28%)
Length = 717
Score = 81.3 bits (199), Expect = 6e-17
Identities = 40/85 (47%), Positives = 55/85 (64%), Gaps = 1/85 (1%)
Frame = +3
Query: 34 SSSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSA 93
S +++G L ++VD L + F +G+V AR++ DRETGRSRGFGFV +SE A
Sbjct: 72 SGLSVYVGNLPWSVDTARLEEIFREHGNVENARIVMDRETGRSRGFGFVTMSSEADINGA 251
Query: 94 L-SMDGQDLNGRNIRVSYANDRQAG 117
+ ++DGQ L+GR IRVS A R G
Sbjct: 252 IAALDGQSLDGRTIRVSVAEGRSGG 326
>TC14420 similar to UP|ROC3_NICSY (P19682) 28 kDa ribonucleoprotein,
chloroplast precursor (28RNP), partial (34%)
Length = 637
Score = 80.1 bits (196), Expect = 1e-16
Identities = 38/79 (48%), Positives = 53/79 (66%), Gaps = 1/79 (1%)
Frame = +1
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-S 95
++++G L ++ D+ L F+ +G VV ARV+ DRETGRSRGFGFV + E A+ +
Sbjct: 43 RVYVGNLPWDFDNSQLEQVFSEHGKVVSARVVYDRETGRSRGFGFVTMSDEAELNDAIAA 222
Query: 96 MDGQDLNGRNIRVSYANDR 114
+DGQ L GR IRV+ A DR
Sbjct: 223 LDGQSLGGRTIRVNVAEDR 279
>AV773507
Length = 496
Score = 76.3 bits (186), Expect = 2e-15
Identities = 40/102 (39%), Positives = 58/102 (56%), Gaps = 1/102 (0%)
Frame = +2
Query: 41 GGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-SMDGQ 99
GGLS++V ++ L AF G ++E +++ +R+TGR RGFGF+ F A+ M G+
Sbjct: 26 GGLSWDVTERQLEHAFDRDGKILECQIMMERDTGRPRGFGFITFADRRGMEDAIKEMHGR 205
Query: 100 DLNGRNIRVSYANDRQAGGPRPGGGYGGGGGYSGGGGGYSGG 141
++ R I V+ A + GG GY GGG SGG G Y G
Sbjct: 206 EIGDRIISVNKAQPKM-GGDDADQGYRGGGYSSGGRGSYGAG 328
>TC16457 similar to UP|Q41124 (Q41124) Chloroplast RNA binding protein
precursor, partial (30%)
Length = 655
Score = 74.3 bits (181), Expect = 8e-15
Identities = 40/83 (48%), Positives = 55/83 (66%), Gaps = 1/83 (1%)
Frame = +2
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTS-EESATSALS 95
KLF+G LS+ V +SL F YG VV ARV+ D ETGRSRG+GFV ++ E T+ +S
Sbjct: 17 KLFVGNLSWTVTSESLIQVFPEYGTVVGARVLYDGETGRSRGYGFVCYSKRSELETALIS 196
Query: 96 MDGQDLNGRNIRVSYANDRQAGG 118
++ +L GR IRVS A +++ G
Sbjct: 197 LNNVELEGRAIRVSLAEGKRS*G 265
>TC12531 similar to UP|Q9ASQ8 (Q9ASQ8) At1g22910/F19G10_13, partial (39%)
Length = 688
Score = 74.3 bits (181), Expect = 8e-15
Identities = 41/116 (35%), Positives = 64/116 (54%), Gaps = 9/116 (7%)
Frame = -1
Query: 36 SKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSALS 95
+K+F+GGL++ ++++ F +GD++EA VITD+ TGRS+G+GFV F E+A A
Sbjct: 361 TKVFVGGLAWETQKETMKKYFEQFGDILEAVVITDKATGRSKGYGFVTFREPEAAMRACV 182
Query: 96 MDGQDLNGRNIRVSYAN-DRQAGGPRPGGGYGGGGG--------YSGGGGGYSGGG 142
++GR + A+ Q P +GG G +GGGGG+ G G
Sbjct: 181 DAAPVIDGRRANCNLASLGVQRSKPSTPKQHGGAGRNFRVMSSFQTGGGGGFGGVG 14
>TC17853 similar to UP|Q42412 (Q42412) RNA-binding protein RZ-1, partial
(58%)
Length = 881
Score = 70.9 bits (172), Expect = 8e-14
Identities = 41/118 (34%), Positives = 62/118 (51%), Gaps = 21/118 (17%)
Frame = +2
Query: 38 LFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-SM 96
LF +++ D+ L+DAF +G + EA+V+ D+ +GRSRGFGFV F +++ A+ +M
Sbjct: 62 LFYWWPAWSTSDRKLKDAFEKFGKLTEAKVVVDKFSGRSRGFGFVTFDDKKAMDEAIDAM 241
Query: 97 DGQDLNGRNIRVSYANDRQAGGPRPGG--------------------GYGGGGGYSGG 134
+G DL+GR I V A + + R G GYGG G +GG
Sbjct: 242 NGMDLDGRTITVDKAQPQGSSRDRDDGDRYRERGRDRDDRGDRDRSRGYGGSRGSNGG 415
Score = 25.0 bits (53), Expect = 5.3
Identities = 17/42 (40%), Positives = 17/42 (40%), Gaps = 15/42 (35%)
Frame = +3
Query: 118 GPRPGGGYGG------GGGY---------SGGGGGYSGGGGG 144
G R GG YGG GGGY SGG GG G
Sbjct: 471 GKRGGGRYGGRESRXSGGGYGPDRNADRSSGGRSSRDGGSHG 596
>BP047377
Length = 564
Score = 70.5 bits (171), Expect = 1e-13
Identities = 37/115 (32%), Positives = 64/115 (55%), Gaps = 3/115 (2%)
Frame = +3
Query: 18 STQAPVSSMLNYLRHMSSSK--LFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGR 75
S +P SS ++++ S +F+G + Y+ ++ L + G VV R++ DRETG+
Sbjct: 69 SRASPSSSAVDFMASSQSQHRCVFVGNIPYDATEEQLIEICQEVGPVVSFRLVIDRETGK 248
Query: 76 SRGFGFVNFTSEESATSA-LSMDGQDLNGRNIRVSYANDRQAGGPRPGGGYGGGG 129
+G+GF + EE+A SA ++ G ++NGR +RV +A + + G GG G
Sbjct: 249 PKGYGFCEYKDEETALSARRNLQGYEINGRQLRVDFAENDKGNDRNREQGRGGPG 413
>TC19017 similar to UP|Q8S2X6 (Q8S2X6) Glycine-rich RNA binding protein
(Fragment), partial (26%)
Length = 320
Score = 68.6 bits (166), Expect = 4e-13
Identities = 35/71 (49%), Positives = 53/71 (74%), Gaps = 5/71 (7%)
Frame = +2
Query: 37 KLFIGGLSYNVDDQSLRDAFT----TYGDVVEARVITDRETGRSRGFGFVNFTSEESATS 92
+LF+G LS+N D+S+R AF G V++++VI DRETGRSRGFGFV F+S + A++
Sbjct: 107 RLFVGNLSWNTTDESMRMAFEDGAGVPGSVLDSKVILDRETGRSRGFGFVTFSSPDLASA 286
Query: 93 ALS-MDGQDLN 102
A++ M+G D++
Sbjct: 287 AIAKMNGMDVD 319
>BP043331
Length = 420
Score = 67.4 bits (163), Expect = 9e-13
Identities = 40/116 (34%), Positives = 64/116 (54%), Gaps = 8/116 (6%)
Frame = +3
Query: 6 KIGNLLRQGATQS-------TQAPVSSMLNYLRHMSSSKLFIGGLSYNVDDQSLRDAFTT 58
++GNLL+ + ++ + +N +S LF+GG+S D+ L DAF+
Sbjct: 12 RLGNLLKHAVNRQIISSEIRSRFLATRSINKAASAPTSMLFVGGISPTTTDRHLDDAFSI 191
Query: 59 YGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-SMDGQDLNGRNIRVSYAND 113
+GDV A V + G+ R FG V FT+ E AT AL +++GQ L GR +R++Y +
Sbjct: 192 HGDVYLATVKHTKVDGKPRQFGLVAFTTVECATRALQALNGQVLQGRRLRINYTKN 359
>TC14208 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, partial (41%)
Length = 837
Score = 61.6 bits (148), Expect = 5e-11
Identities = 36/111 (32%), Positives = 54/111 (48%), Gaps = 2/111 (1%)
Frame = +2
Query: 34 SSSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSA 93
S++ +F+G L NV+D LR F+ YG++V ++ + GFV F A A
Sbjct: 185 SNTTIFVGNLDPNVNDDHLRQVFSPYGELVHVKIPAGKRC------GFVQFADRSGAEEA 346
Query: 94 LS-MDGQDLNGRNIRVSYANDRQAGGPRPGGG-YGGGGGYSGGGGGYSGGG 142
L ++G L G+N+R+S+ +P Y GGY G G GY G
Sbjct: 347 LRVLNGTLLGGQNVRLSWGRSPSNKQAQPDSNQYAAAGGYYGYGQGYENYG 499
>TC8014 similar to GB|CAA41023.1|21309|SORBP28 28kD RNA binding protein
{Spinacia oleracea;} , partial (24%)
Length = 563
Score = 61.2 bits (147), Expect = 7e-11
Identities = 31/54 (57%), Positives = 38/54 (69%), Gaps = 1/54 (1%)
Frame = +2
Query: 62 VVEARVITDRETGRSRGFGFVNFTSEESATSAL-SMDGQDLNGRNIRVSYANDR 114
V AR++ DRETGRSRGFGFV +SE A+ ++DGQ L+GR IRVS A R
Sbjct: 2 VENARIVMDRETGRSRGFGFVTISSEADINGAIPALDGQSLDGRTIRVSVAEGR 163
>TC16529 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, partial
(22%)
Length = 761
Score = 61.2 bits (147), Expect = 7e-11
Identities = 33/79 (41%), Positives = 46/79 (57%), Gaps = 1/79 (1%)
Frame = +1
Query: 35 SSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL 94
++ L++G L NV+D L D F G VV RV D T RS G+G+VNF++ A AL
Sbjct: 328 TTSLYVGDLDQNVNDSQLYDLFNQVGQVVSVRVCRDLTTRRSLGYGYVNFSNPPDAARAL 507
Query: 95 S-MDGQDLNGRNIRVSYAN 112
++ LN R IRV Y++
Sbjct: 508 DVLNFTPLNNRPIRVMYSH 564
>TC11095 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (89%)
Length = 912
Score = 61.2 bits (147), Expect = 7e-11
Identities = 44/130 (33%), Positives = 65/130 (49%), Gaps = 10/130 (7%)
Frame = +1
Query: 26 MLNYLRHMSS---------SKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRS 76
+L +L H++S S++++G L V ++ L D F +G + V R
Sbjct: 31 LLGFLSHIASGFF*KAETMSRVYVGNLDSRVSERDLEDEFRVFGVIRSVWVAR-----RP 195
Query: 77 RGFGFVNFTSEESATSAL-SMDGQDLNGRNIRVSYANDRQAGGPRPGGGYGGGGGYSGGG 135
G+ F++F A A+ +DG+ NG + +S+ N R GG GGGGG G G
Sbjct: 196 PGYAFIDFDDRRDAQDAIRELDGK--NGWRVELSH-NSR--------GGGGGGGG--GRG 336
Query: 136 GGYSGGGGGG 145
GG SGGGGGG
Sbjct: 337 GGRSGGGGGG 366
>TC10758
Length = 532
Score = 60.8 bits (146), Expect = 9e-11
Identities = 31/74 (41%), Positives = 45/74 (59%)
Frame = +1
Query: 31 RHMSSSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESA 90
R + KLF+ GL+ ++LR F+ YGD+ EA VI D+ TGRS+G+GFV F + A
Sbjct: 292 RDTAVRKLFVRGLACETTTETLRSVFSAYGDLDEAIVIFDKSTGRSKGYGFVVFKHIDGA 471
Query: 91 TSALSMDGQDLNGR 104
AL + ++GR
Sbjct: 472 ILALKDPSKKIDGR 513
>AV409413
Length = 422
Score = 60.5 bits (145), Expect = 1e-10
Identities = 37/117 (31%), Positives = 57/117 (48%), Gaps = 12/117 (10%)
Frame = +2
Query: 40 IGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSALSMDGQ 99
+GG+ +V + L+ F+ YG VVE +I D T RSRGFGF+ F +E+ + L+ DG
Sbjct: 2 VGGIPTSVSEDELKGFFSKYGKVVEHEIIRDHTTKRSRGFGFIVFDNEKVVDTILT-DGN 178
Query: 100 --DLNGRNIRVSYANDRQ----------AGGPRPGGGYGGGGGYSGGGGGYSGGGGG 144
D+ G + + A ++ A RP G G++ G + GG G
Sbjct: 179 MIDMAGTQVEIKKAEPKKSSNSGSLPPFASDSRPRSYNDGFSGFADSYGSFGSGGYG 349
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.314 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,962,323
Number of Sequences: 28460
Number of extensions: 29300
Number of successful extensions: 4077
Number of sequences better than 10.0: 566
Number of HSP's better than 10.0 without gapping: 1046
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2438
length of query: 146
length of database: 4,897,600
effective HSP length: 82
effective length of query: 64
effective length of database: 2,563,880
effective search space: 164088320
effective search space used: 164088320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 51 (24.3 bits)
Medicago: description of AC146705.10


