

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146693.3 - phase: 0 /pseudo
(634 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
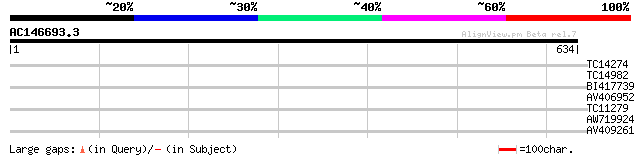
Score E
Sequences producing significant alignments: (bits) Value
TC14274 weakly similar to GB|AAF79559.1|8778551|AC022464 F22G5.3... 30 0.90
TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%) 28 3.4
BI417739 28 5.9
AV406952 28 5.9
TC11279 28 5.9
AW719924 28 5.9
AV409261 27 7.6
>TC14274 weakly similar to GB|AAF79559.1|8778551|AC022464 F22G5.35
{Arabidopsis thaliana;} , partial (16%)
Length = 1208
Score = 30.4 bits (67), Expect = 0.90
Identities = 18/53 (33%), Positives = 24/53 (44%), Gaps = 1/53 (1%)
Frame = +1
Query: 327 PNSLPRLILS-PHTPKCLRPKSPKWPNKWPPLLKH*ESFPVNPKQTPKLMSTP 378
P LP L P TP+ RP P+ P + P L+H + P P + P P
Sbjct: 277 PRDLPLQTLRYPQTPRRHRPPPPRRPPRPP*FLQHPQIHPPPPLRPPPRQDPP 435
>TC14982 similar to UP|Q9FPI2 (Q9FPI2) At1g17620, partial (32%)
Length = 1036
Score = 28.5 bits (62), Expect = 3.4
Identities = 26/87 (29%), Positives = 36/87 (40%), Gaps = 3/87 (3%)
Frame = +1
Query: 337 PHTPKCLRPKSPKWPNKWPPLLKH*ESFPVNPKQTPKLMSTPFLMGVVSWKGQLLKPKEL 396
PH +C R P P P LL+H P P L+ P L + + KP++
Sbjct: 235 PHHRRCWRRLLPPLPPPPPHLLRH---LPQTLLPQPHLLLNPLLQ--IRPHRRRHKPQQE 399
Query: 397 RERVL---GFLVRKMRNRLIRTRPLTH 420
+ R+L F +R R R R TH
Sbjct: 400 KHRILLPTHFHHHTLRRRRRRRRNHTH 480
>BI417739
Length = 591
Score = 27.7 bits (60), Expect = 5.9
Identities = 11/23 (47%), Positives = 15/23 (64%)
Frame = -3
Query: 58 LVPFPRSRFHHFMGRYEASIPCP 80
+ PFP S HH + R++AS P P
Sbjct: 358 IFPFPDSCCHHLLFRFQASSPPP 290
>AV406952
Length = 441
Score = 27.7 bits (60), Expect = 5.9
Identities = 17/55 (30%), Positives = 26/55 (46%), Gaps = 3/55 (5%)
Frame = +1
Query: 327 PNSLPRLILSPHTPKCLRPK---SPKWPNKWPPLLKH*ESFPVNPKQTPKLMSTP 378
P ++ R HTP+ + PK SP W + P + H P+ P TP ++ P
Sbjct: 7 PPNVKRSTDEGHTPEPVSPKSVASPPWEIEHIPPVTH----PIFPITTPSILPAP 159
>TC11279
Length = 455
Score = 27.7 bits (60), Expect = 5.9
Identities = 14/37 (37%), Positives = 21/37 (55%)
Frame = -1
Query: 313 PRTEKPNRISERLSPNSLPRLILSPHTPKCLRPKSPK 349
PRTE+P +I++ +P SL L+ H C P + K
Sbjct: 161 PRTERPKKITQLFAPVSLNSLV-GHHYHPCNPPNNEK 54
>AW719924
Length = 505
Score = 27.7 bits (60), Expect = 5.9
Identities = 13/24 (54%), Positives = 15/24 (62%)
Frame = -1
Query: 324 RLSPNSLPRLILSPHTPKCLRPKS 347
R P SLP + LSPHT + PKS
Sbjct: 493 RREPPSLPYVRLSPHTARTKTPKS 422
>AV409261
Length = 355
Score = 27.3 bits (59), Expect = 7.6
Identities = 15/45 (33%), Positives = 19/45 (41%)
Frame = +2
Query: 312 SPRTEKPNRISERLSPNSLPRLILSPHTPKCLRPKSPKWPNKWPP 356
SP T + ++ + S P L P TP SP WP W P
Sbjct: 230 SPPTPSTSPLASPATSPSPPHSSLPPWTPS----PSPPWPPPWQP 352
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.364 0.164 0.631
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,978,952
Number of Sequences: 28460
Number of extensions: 201579
Number of successful extensions: 2949
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 1688
Number of HSP's successfully gapped in prelim test: 114
Number of HSP's that attempted gapping in prelim test: 1221
Number of HSP's gapped (non-prelim): 1884
length of query: 634
length of database: 4,897,600
effective HSP length: 96
effective length of query: 538
effective length of database: 2,165,440
effective search space: 1165006720
effective search space used: 1165006720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146693.3


