

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146664.9 - phase: 0
(59 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
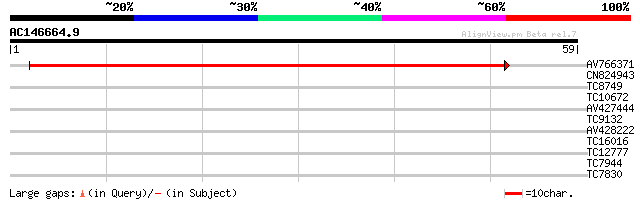
Score E
Sequences producing significant alignments: (bits) Value
AV766371 67 5e-13
CN824943 31 0.056
TC8749 26 1.8
TC10672 weakly similar to PIR|T00710|T00710 thioredoxin homolog ... 25 3.0
AV427444 24 5.2
TC9132 similar to UP|Q8GSM7 (Q8GSM7) Hydroxycinnamoyl transferas... 24 6.8
AV428222 23 8.9
TC16016 23 8.9
TC12777 similar to UP|Q9SYP6 (Q9SYP6) F9H16.10 protein, partial ... 23 8.9
TC7944 homologue to UP|Q9ASR1 (Q9ASR1) At1g56070/T6H22_13 (Elong... 23 8.9
TC7830 UP|Q8H946 (Q8H946) Phosphoenolpyruvate carboxylase , com... 23 8.9
>AV766371
Length = 483
Score = 67.4 bits (163), Expect = 5e-13
Identities = 30/50 (60%), Positives = 36/50 (72%)
Frame = -2
Query: 3 HLSTLACPKVHEEARYLPNMISANFLQKSTVWPESFKNSGTNNFSIGIYF 52
HLSTLACPKV EE R P ++SAN L +S VWP+ F SG N+ SI +YF
Sbjct: 389 HLSTLACPKVLEETRLFPEVLSANLLPRSAVWPKGFSESGPNDESIALYF 240
>CN824943
Length = 719
Score = 30.8 bits (68), Expect = 0.056
Identities = 17/54 (31%), Positives = 31/54 (56%), Gaps = 2/54 (3%)
Frame = +3
Query: 3 HLSTLACPKVHEEA-RYLPNMISANFLQKSTVWPESF-KNSGTNNFSIGIYFLS 54
HLS+ A PKV + ++LP +S + + + + WP F + G+ +I +YF +
Sbjct: 159 HLSSCASPKVLDVVNKFLPE-VSLHEVSRLSTWPSQFHQGGGSKEDNIALYFFA 317
>TC8749
Length = 643
Score = 25.8 bits (55), Expect = 1.8
Identities = 11/25 (44%), Positives = 17/25 (68%), Gaps = 1/25 (4%)
Frame = -2
Query: 3 HLSTLACPKVHEE-ARYLPNMISAN 26
HLS+L C +HE+ A Y+ M+ +N
Sbjct: 228 HLSSLCCNPIHEQVASYISEMMVSN 154
>TC10672 weakly similar to PIR|T00710|T00710 thioredoxin homolog F22O13.5 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(12%)
Length = 511
Score = 25.0 bits (53), Expect = 3.0
Identities = 14/38 (36%), Positives = 19/38 (49%), Gaps = 4/38 (10%)
Frame = +2
Query: 23 ISANFLQK----STVWPESFKNSGTNNFSIGIYFLSPH 56
+S NFL+K S V+ ESF G + G + PH
Sbjct: 98 LSFNFLEKF*RKSCVFLESFNG*GFYQYQFGFFLAWPH 211
>AV427444
Length = 428
Score = 24.3 bits (51), Expect = 5.2
Identities = 12/33 (36%), Positives = 21/33 (63%), Gaps = 2/33 (6%)
Frame = -3
Query: 20 PNMISANFLQKSTVWPESFKNSGT--NNFSIGI 50
P++ +F + S ++ SFKN+GT NF+ G+
Sbjct: 345 PSLFINDFSRLSFIFVISFKNNGTPDTNFTTGL 247
>TC9132 similar to UP|Q8GSM7 (Q8GSM7) Hydroxycinnamoyl transferase,
partial (53%)
Length = 878
Score = 23.9 bits (50), Expect = 6.8
Identities = 8/15 (53%), Positives = 12/15 (79%)
Frame = +3
Query: 18 YLPNMISANFLQKST 32
++PN +S+NFL ST
Sbjct: 75 FIPNFLSSNFLSSST 119
>AV428222
Length = 416
Score = 23.5 bits (49), Expect = 8.9
Identities = 9/29 (31%), Positives = 16/29 (55%)
Frame = -1
Query: 24 SANFLQKSTVWPESFKNSGTNNFSIGIYF 52
+ N + ++ +WP + + NNF GI F
Sbjct: 212 TTNNIIEALIWPNKIR*TEINNFHHGIRF 126
>TC16016
Length = 602
Score = 23.5 bits (49), Expect = 8.9
Identities = 10/28 (35%), Positives = 17/28 (60%)
Frame = -1
Query: 2 GHLSTLACPKVHEEARYLPNMISANFLQ 29
G L T +C KV++E ++ ISA ++
Sbjct: 299 GSLHTESCQKVNKEVKFESKQISAEKME 216
>TC12777 similar to UP|Q9SYP6 (Q9SYP6) F9H16.10 protein, partial (10%)
Length = 500
Score = 23.5 bits (49), Expect = 8.9
Identities = 11/26 (42%), Positives = 17/26 (65%)
Frame = +3
Query: 14 EEARYLPNMISANFLQKSTVWPESFK 39
EEA+Y P+++ A L + TV P+ K
Sbjct: 69 EEAKYAPDLVKALELSEQTV-PDDLK 143
>TC7944 homologue to UP|Q9ASR1 (Q9ASR1) At1g56070/T6H22_13 (Elongation factor
EF-2), complete
Length = 2971
Score = 23.5 bits (49), Expect = 8.9
Identities = 13/43 (30%), Positives = 21/43 (48%), Gaps = 5/43 (11%)
Frame = -2
Query: 21 NMISANFLQKSTVWPESFKNSGTNNFSIGIYFLS-----PHNP 58
N ++ + S ++ +S N FSI ++FLS HNP
Sbjct: 1563 NCVTTRNILNSLLFLSHPNDSSLNTFSIQVFFLSRNIIWSHNP 1435
>TC7830 UP|Q8H946 (Q8H946) Phosphoenolpyruvate carboxylase , complete
Length = 3376
Score = 23.5 bits (49), Expect = 8.9
Identities = 10/38 (26%), Positives = 20/38 (52%)
Frame = +3
Query: 2 GHLSTLACPKVHEEARYLPNMISANFLQKSTVWPESFK 39
GH S L PK+ + + + +S ++T+ P S++
Sbjct: 1578 GHCSDLTSPKLRKSEMFWTHFMSLPNFHRTTLEPISYQ 1691
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.133 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,186,559
Number of Sequences: 28460
Number of extensions: 10893
Number of successful extensions: 75
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 75
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 75
length of query: 59
length of database: 4,897,600
effective HSP length: 35
effective length of query: 24
effective length of database: 3,901,500
effective search space: 93636000
effective search space used: 93636000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 49 (23.5 bits)
Medicago: description of AC146664.9


