

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146664.7 - phase: 0
(2305 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
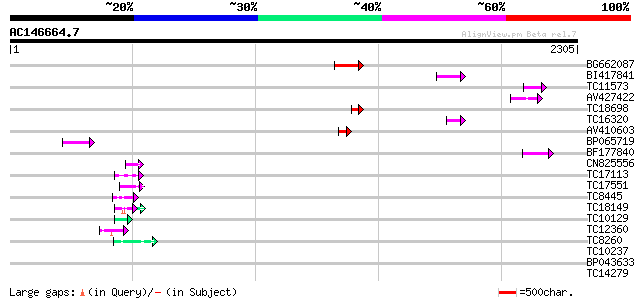
Score E
Sequences producing significant alignments: (bits) Value
BG662087 135 7e-32
BI417841 69 8e-12
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 68 2e-11
AV427422 61 2e-09
TC18698 57 2e-08
TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial ... 53 5e-07
AV410603 50 3e-06
BP065719 50 3e-06
BF177840 49 1e-05
CN825556 45 1e-04
TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protei... 45 1e-04
TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, part... 44 3e-04
TC8445 similar to UP|O80910 (O80910) At2g38410 protein, partial ... 44 4e-04
TC18149 weakly similar to UP|Q91BL3 (Q91BL3) Essential structura... 44 4e-04
TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%) 43 5e-04
TC12360 similar to UP|O22196 (O22196) At2g40820 protein, partial... 43 6e-04
TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment),... 42 8e-04
TC10237 similar to UP|NO75_SOYBN (P08297) Early nodulin 75 precu... 42 0.001
BP043633 41 0.002
TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycop... 41 0.002
>BG662087
Length = 373
Score = 135 bits (340), Expect = 7e-32
Identities = 64/119 (53%), Positives = 84/119 (69%)
Frame = +1
Query: 1321 GKIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFMDGFSGYNQIRMAPEDREK 1380
GK RM VDY DLNKA PKD++PLP ID LVD + ++ S MD +SGY+QI+M P D +K
Sbjct: 16 GKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDEDK 195
Query: 1381 TSFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEIEVYVDDMIVKSGTEEEH 1439
T+F+T +CY +PFGL NAGATYQ M ++F D + + +EVY+D+MIVKS H
Sbjct: 196 TAFMTARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKSALRANH 372
>BI417841
Length = 617
Score = 68.9 bits (167), Expect = 8e-12
Identities = 42/123 (34%), Positives = 65/123 (52%), Gaps = 2/123 (1%)
Frame = +1
Query: 1733 LIFDGAV--NVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVEYEACIMGIEEAIDLRI 1790
L FDG+ N +G GAVL G+ + + + TNN EY I+G++ A +
Sbjct: 142 LEFDGSSKGNPGSAGAGAVLRAEDGSKVYLREGVG-NQTNNQAEYRGLILGLKHAHEQGY 318
Query: 1791 KKIVIYGDSALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKVELHHVPRDENQMADAL 1850
+ I + GDS LV Q++G W+ R+P + + A+ L + F +++HVPR N AD
Sbjct: 319 QHINVKGDSQLVCKQVEGSWKARNPNIASLCNEAKELKSKFQSFDINHVPRQYNSEADVQ 498
Query: 1851 ATL 1853
A L
Sbjct: 499 ANL 507
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 67.8 bits (164), Expect = 2e-11
Identities = 37/96 (38%), Positives = 52/96 (53%), Gaps = 1/96 (1%)
Frame = +2
Query: 2087 DNGTNLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEAANKNIKKIIQKMVVTYKD-WHE 2145
+NG + ++ CD IQ SS PQ NG EAANK I K I++ + + W +
Sbjct: 8 ENGIQFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWID 187
Query: 2146 MLPYALYGYRTSVRTSTGATPFSLVYGMEAVLPVEV 2181
LP L+ Y T ++S TPF L YG + +LPVE+
Sbjct: 188 ELPIVLWSYNTMPQSSIKETPF*LTYGADTMLPVEI 295
>AV427422
Length = 417
Score = 61.2 bits (147), Expect = 2e-09
Identities = 39/136 (28%), Positives = 65/136 (47%), Gaps = 5/136 (3%)
Frame = +1
Query: 2036 SNGHRFILVAIDYFTKWVEAASYAN-VTKQVVVRFIKNNLISRYGVPNRIITDNGTNLNN 2094
S G+ +LV +D +K+ + T +V+ ++ +GVP I++D +
Sbjct: 19 SKGYEAVLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDPLFMS 198
Query: 2095 NMMKELCDDFKIQHHN---SSPYRPQMNGAVEAANKNIKKIIQKMVVTY-KDWHEMLPYA 2150
N KEL FK+Q S+ Y P+ +G E N+ ++ ++ + K W +P+A
Sbjct: 199 NFWKEL---FKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPWA 369
Query: 2151 LYGYRTSVRTSTGATP 2166
Y Y TS STG TP
Sbjct: 370 EYWYNTSYHVSTGQTP 417
>TC18698
Length = 808
Score = 57.4 bits (137), Expect = 2e-08
Identities = 30/50 (60%), Positives = 33/50 (66%)
Frame = -2
Query: 1390 FCYVVMPFGLINAGATYQRGMTKIFHDMIHKEIEVYVDDMIVKSGTEEEH 1439
+ Y VMP GL N TYQR M KIFH I K +EVYV+DMIVKS E H
Sbjct: 771 YYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKSSQE*FH 622
>TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (11%)
Length = 632
Score = 53.1 bits (126), Expect = 5e-07
Identities = 26/77 (33%), Positives = 42/77 (53%)
Frame = +2
Query: 1775 YEACIMGIEEAIDLRIKKIVIYGDSALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKV 1834
Y I+G++ AI K I + GDS LV NQ++G W+ ++ + A+ L F
Sbjct: 2 YRGLILGLKHAIKEGYKHIQVKGDSMLVCNQVQGLWKIKNQNIASLCSEAKELKNKFLSF 181
Query: 1835 ELHHVPRDENQMADALA 1851
+++H+PR+ N AD A
Sbjct: 182 KINHIPREYNSEADVQA 232
>AV410603
Length = 162
Score = 50.4 bits (119), Expect = 3e-06
Identities = 25/51 (49%), Positives = 33/51 (64%)
Frame = +1
Query: 1338 KDNFPLPHIDVLVDNTAKCKVFSFMDGFSGYNQIRMAPEDREKTSFITPWG 1388
KD+FP+P +D L+D + FS +D SGY+QI + PEDR KT F T G
Sbjct: 10 KDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTHHG 162
>BP065719
Length = 567
Score = 50.4 bits (119), Expect = 3e-06
Identities = 40/132 (30%), Positives = 60/132 (45%), Gaps = 4/132 (3%)
Frame = +3
Query: 215 VPDVNIPPKFKMPVFEKYQGDT--CPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAH 272
V + +P +K+P F K+ GD+ H+ Y N L + F SLT A
Sbjct: 147 VLESELPRGWKVPKFTKFSGDSGESTVEHIARYQIEAGDLAINENLKMKYFPSSLTKNAF 326
Query: 273 TWFMGL--KGVTTFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKA 330
TWF L + V T+ QL F +Q+ + S K+L S+ +K ES +Y RF
Sbjct: 327 TWFTTLAPRSVHTWAQLERIFHEQF-FRGECKVSXKDLASVKRKPAESIDDYLNRFRMLK 503
Query: 331 AQIRPPLDEREL 342
++ + E EL
Sbjct: 504 SRCFTHVSEHEL 539
>BF177840
Length = 410
Score = 48.5 bits (114), Expect = 1e-05
Identities = 29/129 (22%), Positives = 65/129 (49%), Gaps = 3/129 (2%)
Frame = +2
Query: 2084 IITDNGTNLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEAANKNIKKIIQKMVV-TYKD 2142
I++D T ++ + L + S+ PQ +G E NK + +++ ++ K
Sbjct: 17 IVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNLKM 196
Query: 2143 WHEMLPYALYGYRTSVRTSTGATPFSLVYGMEAVLPVE-VEIPSLRVLMEAE-LSEAEWC 2200
W LP+ + Y V ++T +PF +VYG + P++ + +P+ +L + ++AE+
Sbjct: 197 WETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDLLPMPNTYLLKHKDGKAKAEFV 376
Query: 2201 QSRYDQLNL 2209
+ ++++ L
Sbjct: 377 KRLHEKIKL 403
>CN825556
Length = 655
Score = 45.1 bits (105), Expect = 1e-04
Identities = 27/71 (38%), Positives = 36/71 (50%)
Frame = +1
Query: 472 PQQLSQPPYYPQQPYQQRPQQQPRPQVPHNQQYQRQQFDPLPMTYGALLPSLLAQNLVQT 531
PQ + PP PQ Q +PQ QP+PQ P Q + Q P P+ A LP Q L+
Sbjct: 433 PQPVPPPPSLPQPQPQPQPQPQPQPQ-PQPQPQPQPQPQPQPVPPPASLP----QPLLAP 597
Query: 532 IPPPRIPDSLP 542
+P P +P+ P
Sbjct: 598 LPSPGLPEQKP 630
>TC17113 weakly similar to UP|Q9LFU8 (Q9LFU8) Proline-rich protein, partial
(25%)
Length = 1077
Score = 45.1 bits (105), Expect = 1e-04
Identities = 37/118 (31%), Positives = 50/118 (42%)
Frame = +1
Query: 425 PQPVYPNHPFVANITPQMTAPQNPNYQPPRPQGPSLYYPPLYQQPYNPQQLSQPPYYPQQ 484
P P+ PN PF + P + P +QPP P PPL+ P+ P S PP P
Sbjct: 490 PPPLIPN-PFQPSPPPLLPDP----FQPPSP-------PPLFPNPFQPP--SPPPLIPN- 624
Query: 485 PYQQRPQQQPRPQVPHNQQYQRQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDSLP 542
P+Q P P P +P+ Q + P P+ P ++ L PPP P P
Sbjct: 625 PFQPPP--SPPPFIPNPFQPPPSKQPPSPL---FPFPPIVIPGLTPPPPPPPPPPFFP 783
Score = 33.5 bits (75), Expect = 0.38
Identities = 31/101 (30%), Positives = 41/101 (39%), Gaps = 6/101 (5%)
Frame = +1
Query: 453 PRPQGPSLYYPPLYQQPYNPQQL-----SQPPYYPQQPYQQRPQQQPRPQVPHNQQYQRQ 507
P P P+ + PPL P+ P L PP P P+ Q P P +P +Q
Sbjct: 403 PPPLIPNPFQPPLVPNPFQPPPLIPNPFQPPPLIP-NPF----QPSPPPLLP--DPFQPP 561
Query: 508 QFDPL-PMTYGALLPSLLAQNLVQTIPPPRIPDSLPRWYRP 547
PL P + P L N Q PPP P +P ++P
Sbjct: 562 SPPPLFPNPFQPPSPPPLIPNPFQ--PPPSPPPFIPNPFQP 678
Score = 29.3 bits (64), Expect = 7.3
Identities = 22/79 (27%), Positives = 30/79 (37%)
Frame = +1
Query: 461 YYPPLYQQPYNPQQLSQPPYYPQQPYQQRPQQQPRPQVPHNQQYQRQQFDPLPMTYGALL 520
++PP+ P+ P L P+ P P VP+ F P P+
Sbjct: 379 FFPPI---PFFPPPLIPNPFQP-------------PLVPN-------PFQPPPLIPNPFQ 489
Query: 521 PSLLAQNLVQTIPPPRIPD 539
P L N Q PPP +PD
Sbjct: 490 PPPLIPNPFQPSPPPLLPD 546
>TC17551 weakly similar to UP|AAR24662 (AAR24662) At5g26960, partial (22%)
Length = 505
Score = 43.9 bits (102), Expect = 3e-04
Identities = 36/100 (36%), Positives = 48/100 (48%)
Frame = +1
Query: 448 PNYQPPRPQGPSLYYPPLYQQPYNPQQLSQPPYYPQQPYQQRPQQQPRPQVPHNQQYQRQ 507
P PP P P L PPL P P+ L PP P P RP+ P + R+
Sbjct: 175 PTLFPPFPL-PRL--PPLVPPPPLPRLLRPPPPPPPPP-PLRPRPHRHRLRPLLRHPPRR 342
Query: 508 QFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDSLPRWYRP 547
Q +PLP LP LL + L + +P P +P + PR++RP
Sbjct: 343 QMEPLP------LPPLLQRRLPRDLPLPPLPRA-PRFHRP 441
>TC8445 similar to UP|O80910 (O80910) At2g38410 protein, partial (7%)
Length = 841
Score = 43.5 bits (101), Expect = 4e-04
Identities = 36/105 (34%), Positives = 44/105 (41%), Gaps = 2/105 (1%)
Frame = +1
Query: 419 MVAHGGPQPV-YPNHPFVANITPQMTAPQNPNYQPPRPQGPSLYYPPLYQQPYNPQQLSQ 477
M H PQP+ Y P + + PQ+ + P PQ P L QP PQ + Q
Sbjct: 7 MQLHNEPQPLQYQPQPHLQH------QPQHHPHFQPHPQ------PQLQSQP-QPQHIQQ 147
Query: 478 PPYYPQQPYQQRPQQQPRPQVPHNQQYQRQ-QFDPLPMTYGALLP 521
PQ Q Q QP+PQ+ H QQ Q Q QF Y P
Sbjct: 148 SQPQPQSQPQHIQQPQPQPQLQHIQQSQPQPQFQNQHFQYPGTYP 282
Score = 30.0 bits (66), Expect = 4.3
Identities = 20/62 (32%), Positives = 28/62 (44%), Gaps = 1/62 (1%)
Frame = +1
Query: 477 QPPYYPQQPY-QQRPQQQPRPQVPHNQQYQRQQFDPLPMTYGALLPSLLAQNLVQTIPPP 535
QP Y QP+ Q +PQ P Q PH Q + Q P + P Q++ Q P P
Sbjct: 28 QPLQYQPQPHLQHQPQHHPHFQ-PHPQPQLQSQPQPQHIQQSQPQPQSQPQHIQQPQPQP 204
Query: 536 RI 537
++
Sbjct: 205 QL 210
>TC18149 weakly similar to UP|Q91BL3 (Q91BL3) Essential structural protein
pp78/81, partial (5%)
Length = 710
Score = 43.5 bits (101), Expect = 4e-04
Identities = 33/103 (32%), Positives = 44/103 (42%), Gaps = 9/103 (8%)
Frame = +3
Query: 425 PQPVYPNHP-FVANITPQMTAPQNPNYQPPRPQGPSLYYPPLYQQPYNPQQLSQPPYYPQ 483
PQP P H + PQ + P P +QPP P PP Q + +PPY P
Sbjct: 306 PQPGAPPHQQYQQTPHPQYSQPAPPQHQPPIPS----INPPQLQSSLG-HHVEEPPYVPS 470
Query: 484 QPY----QQRPQQQPR-PQVPHNQQY---QRQQFDPLPMTYGA 518
Q Y +Q P Q P P P +QQ+ ++P P G+
Sbjct: 471 QTYPPNLRQPPSQPPSGPPAPPSQQFYGTAPHAYEPPPSRSGS 599
Score = 43.1 bits (100), Expect = 5e-04
Identities = 41/142 (28%), Positives = 55/142 (37%), Gaps = 19/142 (13%)
Frame = +3
Query: 427 PVYPNHPFVANITPQMTAPQNPNYQPPR----------PQGPSLYYPPL---------YQ 467
PV P + PQ P P++ PP PQ + PP+ YQ
Sbjct: 108 PVVPPPNAPPQLPPQQGLP--PSFPPPNQFSQTQIPAVPQRDPYFPPPVQSQETPNQQYQ 281
Query: 468 QPYNPQQLSQPPYYPQQPYQQRPQQQPRPQVPHNQQYQRQQFDPLPMTYGALLPSLLAQN 527
QP PQ QP P Q YQQ P Q ++Q Q P+P L S L +
Sbjct: 282 QPLAPQPHPQPGAPPHQQYQQTPHPQ------YSQPAPPQHQPPIPSINPPQLQSSLGHH 443
Query: 528 LVQTIPPPRIPDSLPRWYRPDL 549
+ + PP +P + Y P+L
Sbjct: 444 VEE---PPYVPS---QTYPPNL 491
>TC10129 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (10%)
Length = 562
Score = 43.1 bits (100), Expect = 5e-04
Identities = 33/92 (35%), Positives = 36/92 (38%), Gaps = 17/92 (18%)
Frame = +1
Query: 425 PQPVY--------PNHPFVANITPQMTAPQ---NPNYQPPR----PQGPSLYYPPLYQQ- 468
P PVY P P N P P +P+Y PP P+ PS P Y
Sbjct: 1 PSPVYTPPSIPTPPKTPSPGNQPPHTPIPPKTPSPSYSPPNVPSPPKAPSPNNHPPYTPT 180
Query: 469 -PYNPQQLSQPPYYPQQPYQQRPQQQPRPQVP 499
P P SQPPY P P P QP P VP
Sbjct: 181 PPKTPSPTSQPPYIPLPPKTPSPISQP-PHVP 273
Score = 41.2 bits (95), Expect = 0.002
Identities = 27/72 (37%), Positives = 34/72 (46%), Gaps = 3/72 (4%)
Frame = +1
Query: 431 NHPFVANITPQMTAPQN-PNYQPPRPQGPS-LYYPP-LYQQPYNPQQLSQPPYYPQQPYQ 487
NHP P+ +P + P Y P P+ PS + PP + P P +SQPPY P P
Sbjct: 157 NHPPYTPTPPKTPSPTSQPPYIPLPPKTPSPISQPPHVPTPPSTPSPISQPPYTPTPPKT 336
Query: 488 QRPQQQPRPQVP 499
P QP P P
Sbjct: 337 PSPTSQP-PHTP 369
Score = 40.0 bits (92), Expect = 0.004
Identities = 28/82 (34%), Positives = 35/82 (42%), Gaps = 9/82 (10%)
Frame = +1
Query: 427 PVYPNHPFVANITPQMTAPQN-------PNYQPPRPQGPS-LYYPPLYQQPYN-PQQLSQ 477
P+ P P + P + P + P Y P P+ PS PP P N P +SQ
Sbjct: 223 PLPPKTPSPISQPPHVPTPPSTPSPISQPPYTPTPPKTPSPTSQPPHTPTPPNTPSPISQ 402
Query: 478 PPYYPQQPYQQRPQQQPRPQVP 499
PPY P P P QP P +P
Sbjct: 403 PPYTPTPPKTPSPTNQP-PHIP 465
Score = 40.0 bits (92), Expect = 0.004
Identities = 26/96 (27%), Positives = 33/96 (34%), Gaps = 6/96 (6%)
Frame = +1
Query: 410 PKRNAQEVGMVAHGGPQPVYPNHPFVANITPQMTAPQNPNYQPPRPQGPSLYYPPLYQQP 469
P + + H P P+ TP +P QPP P P+ Q P
Sbjct: 229 PPKTPSPISQPPHVPTPPSTPSPISQPPYTPTPPKTPSPTSQPPHTPTPPNTPSPISQPP 408
Query: 470 YNP------QQLSQPPYYPQQPYQQRPQQQPRPQVP 499
Y P +QPP+ P P P QP P P
Sbjct: 409 YTPTPPKTPSPTNQPPHIPSPPNSSSPTSQP-PNTP 513
Score = 39.7 bits (91), Expect = 0.005
Identities = 29/95 (30%), Positives = 37/95 (38%), Gaps = 21/95 (22%)
Frame = +1
Query: 427 PVYPNHPFVANITPQMTAPQ---NPNYQPPRPQGPSLYYPPLYQQPY------NPQQLSQ 477
P+ P P + P + +P +PN PP P P Q PY P +SQ
Sbjct: 79 PIPPKTPSPSYSPPNVPSPPKAPSPNNHPPYTPTPPKTPSPTSQPPYIPLPPKTPSPISQ 258
Query: 478 PPYYP----------QQPYQQRPQQQPRP--QVPH 500
PP+ P Q PY P + P P Q PH
Sbjct: 259 PPHVPTPPSTPSPISQPPYTPTPPKTPSPTSQPPH 363
Score = 33.1 bits (74), Expect = 0.50
Identities = 21/68 (30%), Positives = 26/68 (37%)
Frame = +1
Query: 427 PVYPNHPFVANITPQMTAPQNPNYQPPRPQGPSLYYPPLYQQPYNPQQLSQPPYYPQQPY 486
P PN P +P P P PP+ P+ P + P + SQPP P P
Sbjct: 367 PTPPNTP-----SPISQPPYTPT--PPKTPSPTNQPPHIPSPPNSSSPTSQPPNTPSPPK 525
Query: 487 QQRPQQQP 494
P QP
Sbjct: 526 TPSPTSQP 549
>TC12360 similar to UP|O22196 (O22196) At2g40820 protein, partial (11%)
Length = 1030
Score = 42.7 bits (99), Expect = 6e-04
Identities = 39/134 (29%), Positives = 55/134 (40%), Gaps = 15/134 (11%)
Frame = +3
Query: 363 SQKFTDLVETGMRIEEWARKGAAVSGSSSGGSSGVSSNGNK-KFGNGYP----------- 410
S K + V+ +R+ E K S GG +S +GNK +FG YP
Sbjct: 429 SNKPSGFVDVSIRVFE--DKEEVDSHPGEGGGILLSDHGNKGRFGQAYPHQMAPASFHGP 602
Query: 411 KRNAQEVGMVAHGGPQPVYPNHPFVANITPQMTAPQNPNYQPPRPQGPSLYYPPL---YQ 467
++ AQ +H P P ++P+V P A P+YQPPR P PP Y
Sbjct: 603 EKQAQINTPYSHPVPFPANYSNPYVGG--PSYPAAAGPSYQPPRTHPPPPPPPPSNTGYI 776
Query: 468 QPYNPQQLSQPPYY 481
++P P Y
Sbjct: 777 PSFHPSNDGLGPSY 818
>TC8260 similar to UP|Q94ES8 (Q94ES8) Nodule extensin (Fragment), partial
(79%)
Length = 1017
Score = 42.4 bits (98), Expect = 8e-04
Identities = 47/190 (24%), Positives = 69/190 (35%), Gaps = 10/190 (5%)
Frame = +3
Query: 421 AHGGPQPVYPNHPFVANITPQMTAPQNPNYQPPRPQGPSLYY--PPLYQQPYNPQQLSQP 478
A+ P P P P+ + P P Y+ P P +Y+ PP Y++P+ P S P
Sbjct: 111 AYSSPPP--PPKPYSYHSPPPPVHSPPPPYEKPHP----VYHSPPPPYEKPH-PVYHSPP 269
Query: 479 PYYPQQPYQQRPQQQPRPQVPHNQQYQRQQFDPLP-----MTYGALLPSLLA---QNLVQ 530
P P +PY+ P + PH + P P Y + P + + V
Sbjct: 270 PPPPHKPYKYPSPPPPPHKYPHPHPHPVYHSPPPPPPKKHYKYSSPPPPVHTYPHPHPVY 449
Query: 531 TIPPPRIPDSLPRWYRPDLHCIYHQGAPGHDVERCFALKKEVQKLINSKELTFTDPDAVA 590
PPP P P H +YH P E+ + ++ T P V
Sbjct: 450 HSPPP------PVHTYPHPHPVYHSPPPPPPHEKPYKYASPPPPPVH------TYPHPVY 593
Query: 591 QNNPLPTHGP 600
+ P P H P
Sbjct: 594 HSPPPPVHSP 623
Score = 35.0 bits (79), Expect = 0.13
Identities = 26/75 (34%), Positives = 32/75 (42%), Gaps = 8/75 (10%)
Frame = +3
Query: 420 VAHGGPQPV--YPNHPFVANITPQMTAPQNPNYQPPRPQGPSLY-YP-PLYQQPYNPQQL 475
V H P PV YP HP +P P Y+ P P ++ YP P+Y P P
Sbjct: 444 VYHSPPPPVHTYP-HPHPVYHSPPPPPPHEKPYKYASPPPPPVHTYPHPVYHSPPPPVHS 620
Query: 476 SQPPYY----PQQPY 486
PP+Y P PY
Sbjct: 621 PPPPHYYYKSPPPPY 665
Score = 30.4 bits (67), Expect = 3.3
Identities = 21/67 (31%), Positives = 32/67 (47%), Gaps = 2/67 (2%)
Frame = +2
Query: 44 STPVVVVTAAPEGPPQPSPTTTTTIDLTQPLMTDFSSGNMATMGNSGFRPLGP--QGPFA 101
STP V + P P P +TTTT + Q L++ +S ++ + S PL P
Sbjct: 308 STPTTQVPSPPPSPSLPLSSTTTT*EALQVLISSTTSSHLPSPSPS--LPLSPTTSSHLP 481
Query: 102 TPQYSMP 108
+P S+P
Sbjct: 482 SPTPSLP 502
>TC10237 similar to UP|NO75_SOYBN (P08297) Early nodulin 75 precursor (N-75)
(NGM-75), partial (49%)
Length = 803
Score = 42.0 bits (97), Expect = 0.001
Identities = 30/106 (28%), Positives = 42/106 (39%), Gaps = 13/106 (12%)
Frame = +1
Query: 410 PKRNAQEVGMVAHGGPQPVYPNHPFVANITPQMTAPQN---PNYQPP---------RPQG 457
P+ V + + P P YP P P+ PQ P Y PP P
Sbjct: 154 PQEKPPPVYLPPYWKPPPAYP--PPYEKPPPEYEPPQEKPPPVYSPPYERPPTGHPTPYP 327
Query: 458 PSLYYPPLYQQPY-NPQQLSQPPYYPQQPYQQRPQQQPRPQVPHNQ 502
P + PP+Y PY P L QPP+ Y+ ++ P + PH +
Sbjct: 328 PFYHLPPVYHPPYEKPPPLYQPPHEKPPIYEPPHEKPPIYEPPHEK 465
Score = 41.6 bits (96), Expect = 0.001
Identities = 39/140 (27%), Positives = 53/140 (37%), Gaps = 18/140 (12%)
Frame = +1
Query: 426 QPVYPNHPFVANITPQMTAPQN----PNYQPPRPQGPSLYYPP------------LYQQP 469
QP + PF P + P + P YQPP + PS Y PP +Y P
Sbjct: 19 QPPHEKPPFEK---PPLEKPPHEKPPPEYQPPHKKPPSQYQPPPEYEPPQEKPPPVYLPP 189
Query: 470 Y-NPQQLSQPPYYPQQPYQQRPQQQPRPQVPHNQQYQRQQF-DPLPMTYGALLPSLLAQN 527
Y P PPY P + PQ++P P ++ Y+R P P LP +
Sbjct: 190 YWKPPPAYPPPYEKPPPEYEPPQEKPPP--VYSPPYERPPTGHPTPYPPFYHLPPVYHPP 363
Query: 528 LVQTIPPPRIPDSLPRWYRP 547
+ P + P P Y P
Sbjct: 364 YEKPPPLYQPPHEKPPIYEP 423
Score = 36.6 bits (83), Expect = 0.045
Identities = 17/54 (31%), Positives = 25/54 (45%), Gaps = 5/54 (9%)
Frame = +1
Query: 448 PNYQPPRPQGPSLYYPP-----LYQQPYNPQQLSQPPYYPQQPYQQRPQQQPRP 496
P Y PP + P LY PP +Y+ P+ + +PP+ Y+ P P P
Sbjct: 346 PVYHPPYEKPPPLYQPPHEKPPIYEPPHEKPPIYEPPHEKPPLYEPPPGYNPPP 507
Score = 35.8 bits (81), Expect = 0.078
Identities = 17/57 (29%), Positives = 26/57 (44%)
Frame = +1
Query: 458 PSLYYPPLYQQPYNPQQLSQPPYYPQQPYQQRPQQQPRPQVPHNQQYQRQQFDPLPM 514
P Y PP + P+ L +PP+ P Q P ++P Q +Y+ Q P P+
Sbjct: 7 PPKYQPPHEKPPFEKPPLEKPPHEKPPPEYQPPHKKPPSQYQPPPEYEPPQEKPPPV 177
Score = 31.6 bits (70), Expect = 1.5
Identities = 22/67 (32%), Positives = 27/67 (39%), Gaps = 3/67 (4%)
Frame = +1
Query: 427 PVYPNHPFVANITPQMTAPQN--PNYQPPRPQGPSLYYPPLYQQPY-NPQQLSQPPYYPQ 483
PVY HP P P P Y+PP + PP+Y+ P+ P PP Y
Sbjct: 346 PVY--HPPYEKPPPLYQPPHEKPPIYEPPHEK------PPIYEPPHEKPPLYEPPPGYNP 501
Query: 484 QPYQQRP 490
PY P
Sbjct: 502 PPYGHYP 522
>BP043633
Length = 542
Score = 41.2 bits (95), Expect = 0.002
Identities = 30/90 (33%), Positives = 35/90 (38%), Gaps = 15/90 (16%)
Frame = +1
Query: 425 PQPVYPNHPFVANITPQMTAPQNPNYQP------------PRPQ-GPSLYYPPLYQQPYN 471
P P PNH P PQ P Y P P PQ GP PP PY+
Sbjct: 265 PPPPDPNHHHHFPQIPPPHPPQAPWYSPQFQYHHPSQTPSPPPQWGPPPPPPPPPSNPYS 444
Query: 472 --PQQLSQPPYYPQQPYQQRPQQQPRPQVP 499
P+Q PP P+ P+ PQ P +P
Sbjct: 445 YHPKQFPPPPPPPRPPHPPHPQFPPHSHIP 534
>TC14279 similar to UP|Q09085 (Q09085) Hydroxyproline-rich glycoprotein
(HRGP) (Fragment), partial (57%)
Length = 941
Score = 41.2 bits (95), Expect = 0.002
Identities = 36/129 (27%), Positives = 44/129 (33%), Gaps = 14/129 (10%)
Frame = +1
Query: 421 AHGGPQPVYPNHPFVANITPQMTAPQNPNYQ----PPRPQGPSLYYPPLYQQPYNPQQLS 476
+H P P Y P P +P P Y PP P P YY Y+ P P +
Sbjct: 157 SHSPPPPYYYKSP-----PPPSPSPPPPYYYKSPPPPSPSPPPPYY---YKSPPPPSPIP 312
Query: 477 QPPYY----------PQQPYQQRPQQQPRPQVPHNQQYQRQQFDPLPMTYGALLPSLLAQ 526
PPYY P PY P P P Y P P + PS
Sbjct: 313 HPPYYYKSPPPPTSSPPPPYHYVSPPPPSPSPPPPYHYA----SPPPPS-----PSPAPT 465
Query: 527 NLVQTIPPP 535
+ ++ PPP
Sbjct: 466 YIYKSPPPP 492
Score = 33.9 bits (76), Expect = 0.29
Identities = 34/125 (27%), Positives = 43/125 (34%), Gaps = 4/125 (3%)
Frame = +1
Query: 425 PQPVYPNHPFVANITPQMTAPQNPNYQ----PPRPQGPSLYYPPLYQQPYNPQQLSQPPY 480
P P P + + P +P P Y PP P P PP Y + P S PP
Sbjct: 106 PSPSPPPPYYYQSPPPPSHSPPPPYYYKSPPPPSPSPP----PPYYYKSPPPPSPSPPP- 270
Query: 481 YPQQPYQQRPQQQPRPQVPHNQQYQRQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDS 540
PY + P P +PH Y + P P T P PPP P
Sbjct: 271 ----PYYYKSPPPPSP-IPHPPYYYK---SPPPPTSSPPPPYHYVS------PPPPSPSP 408
Query: 541 LPRWY 545
P ++
Sbjct: 409 PPPYH 423
Score = 30.8 bits (68), Expect = 2.5
Identities = 26/96 (27%), Positives = 34/96 (35%)
Frame = +1
Query: 450 YQPPRPQGPSLYYPPLYQQPYNPQQLSQPPYYPQQPYQQRPQQQPRPQVPHNQQYQRQQF 509
Y+ P P PP +PY + PPYY Y+ P P P P+ Q
Sbjct: 13 YKSPPP-------PPPVPKPYYYKSPPPPPYY----YKSPPPPSPSPPPPYYYQ------ 141
Query: 510 DPLPMTYGALLPSLLAQNLVQTIPPPRIPDSLPRWY 545
P P ++ P PPP P P +Y
Sbjct: 142 SPPPPSHSPPPPYYYKS------PPPPSPSPPPPYY 231
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 40,030,376
Number of Sequences: 28460
Number of extensions: 605203
Number of successful extensions: 4492
Number of sequences better than 10.0: 175
Number of HSP's better than 10.0 without gapping: 3755
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4192
length of query: 2305
length of database: 4,897,600
effective HSP length: 105
effective length of query: 2200
effective length of database: 1,909,300
effective search space: 4200460000
effective search space used: 4200460000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146664.7


