

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146632.13 - phase: 0
(126 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
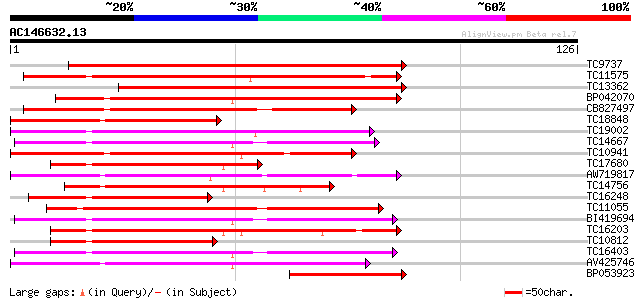
Score E
Sequences producing significant alignments: (bits) Value
TC9737 similar to UP|Q9LRT1 (Q9LRT1) Receptor protein kinase, pa... 145 3e-36
TC11575 similar to UP|Q9LN92 (Q9LN92) T12C24.1, partial (52%) 81 5e-17
TC13362 weakly similar to UP|CAE54285 (CAE54285) Receptor-like k... 76 2e-15
BP042070 60 8e-11
CB827497 55 3e-09
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 55 4e-09
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 55 5e-09
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 54 6e-09
TC10941 homologue to UP|Q7Y229 (Q7Y229) At1g56720, partial (27%) 54 6e-09
TC17680 similar to UP|O65440 (O65440) CLV1 receptor kinase like ... 54 1e-08
AW719817 53 1e-08
TC14756 weakly similar to UP|Q41944 (Q41944) Receptor-like prote... 53 2e-08
TC16248 similar to UP|BAC87845 (BAC87845) Leucine-rich repeat re... 53 2e-08
TC11055 similar to UP|Q9LR04 (Q9LR04) F10A5.16, partial (12%) 52 4e-08
BI419694 52 4e-08
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 51 5e-08
TC10812 similar to UP|O65440 (O65440) CLV1 receptor kinase like ... 51 7e-08
TC16403 homologue to UP|Q9LKY4 (Q9LKY4) Pti1 kinase-like protein... 50 9e-08
AV425746 50 9e-08
BP053923 48 4e-07
>TC9737 similar to UP|Q9LRT1 (Q9LRT1) Receptor protein kinase, partial
(11%)
Length = 563
Score = 145 bits (365), Expect = 3e-36
Identities = 67/75 (89%), Positives = 71/75 (94%)
Frame = +2
Query: 14 PELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVRVLLEHGNALECV 73
PELACQ LRVNEKCDVYGFGVMILE+VTG+RPVEYGEDNVLILNDHVRVLLEHGN LECV
Sbjct: 2 PELACQSLRVNEKCDVYGFGVMILELVTGRRPVEYGEDNVLILNDHVRVLLEHGNVLECV 181
Query: 74 DPSLMSEYPEDEVLP 88
DPS+ EYP+DEVLP
Sbjct: 182DPSMSHEYPDDEVLP 226
>TC11575 similar to UP|Q9LN92 (Q9LN92) T12C24.1, partial (52%)
Length = 647
Score = 81.3 bits (199), Expect = 5e-17
Identities = 43/85 (50%), Positives = 64/85 (74%), Gaps = 1/85 (1%)
Frame = +1
Query: 4 RFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDN-VLILNDHVRV 62
+F + +GYVAPELA Q +R +EKCDVY FGV++LE+VTG++PVE N V++L ++VR
Sbjct: 19 KFHNVVGYVAPELA-QSMRQSEKCDVYSFGVILLELVTGRKPVESPTSNEVVVLCEYVRG 195
Query: 63 LLEHGNALECVDPSLMSEYPEDEVL 87
LLE G+A C D +L+ + E+E++
Sbjct: 196 LLETGSASNCFDRNLVG-FAENELI 267
>TC13362 weakly similar to UP|CAE54285 (CAE54285) Receptor-like kinase
with LRR repeats (Fragment), partial (60%)
Length = 519
Score = 75.9 bits (185), Expect = 2e-15
Identities = 35/64 (54%), Positives = 45/64 (69%)
Frame = +3
Query: 25 EKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVRVLLEHGNALECVDPSLMSEYPED 84
EKCDVYGFG++ILE VTGKRPVEY ED+ ++L + VR LE G CVD L+ + +
Sbjct: 3 EKCDVYGFGILILEXVTGKRPVEYLEDDXVVLCELVRGALEEGKVEPCVDGRLLGNFAAE 182
Query: 85 EVLP 88
E +P
Sbjct: 183EAIP 194
>BP042070
Length = 460
Score = 60.5 bits (145), Expect = 8e-11
Identities = 34/78 (43%), Positives = 48/78 (60%), Gaps = 1/78 (1%)
Frame = +3
Query: 11 YVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY-GEDNVLILNDHVRVLLEHGNA 69
Y+APE A + + +K DVY FG++ LEIV+GK Y ++ + L D VL E GN
Sbjct: 3 YMAPEYAMRGY-LTDKADVYSFGIVALEIVSGKSNTNYRPKEEFIYLLDWAYVLQEQGNL 179
Query: 70 LECVDPSLMSEYPEDEVL 87
LE VDP L S+Y ++E +
Sbjct: 180LELVDPGLGSKYSQEEAM 233
>CB827497
Length = 550
Score = 55.5 bits (132), Expect = 3e-09
Identities = 26/74 (35%), Positives = 49/74 (66%)
Frame = +2
Query: 4 RFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVRVL 63
+F+ GY+APE + V+EK DV+ FGV++LE+V+G+R +++ + ++++ + L
Sbjct: 209 KFEGTFGYLAPEYLLHGI-VDEKTDVFAFGVLLLELVSGRRALDFSQQSLVL---WAKPL 376
Query: 64 LEHGNALECVDPSL 77
L+ +E +DPSL
Sbjct: 377 LKKNEIMELIDPSL 418
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 55.1 bits (131), Expect = 4e-09
Identities = 24/47 (51%), Positives = 36/47 (76%)
Frame = +2
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVE 47
M+ R + LGY+APE A +V+E CDVY FG+++LE+VTG++P+E
Sbjct: 776 MTTRVKGTLGYLAPEYAMWG-KVSESCDVYSFGILLLELVTGRKPIE 913
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 54.7 bits (130), Expect = 5e-09
Identities = 31/82 (37%), Positives = 46/82 (55%), Gaps = 1/82 (1%)
Frame = +1
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNV-LILNDH 59
++ R + LGY+APE A + + NE CDV+ FG+++LE+ +GK+P+E V +ND
Sbjct: 538 VTTRVKGTLGYLAPEYA-MLGKANECCDVFSFGILLLELASGKKPLEKLSSTVKRSINDW 714
Query: 60 VRVLLEHGNALECVDPSLMSEY 81
L E DP L EY
Sbjct: 715 ALPLACAKKFTEFADPRLNGEY 780
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 54.3 bits (129), Expect = 6e-09
Identities = 31/85 (36%), Positives = 47/85 (54%), Gaps = 4/85 (4%)
Frame = +2
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY----GEDNVLILN 57
S R GY APE A ++ +K DVY FGV++LE++TG++PV++ G+ +++
Sbjct: 857 STRVLGTFGYHAPEYA-MTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLV--- 1024
Query: 58 DHVRVLLEHGNALECVDPSLMSEYP 82
L +CVDP L EYP
Sbjct: 1025TWATPRLSEDKVKQCVDPKLKGEYP 1099
>TC10941 homologue to UP|Q7Y229 (Q7Y229) At1g56720, partial (27%)
Length = 831
Score = 54.3 bits (129), Expect = 6e-09
Identities = 33/78 (42%), Positives = 49/78 (62%), Gaps = 1/78 (1%)
Frame = +2
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGE-DNVLILNDH 59
++ R GYVAPE A L +NEK DVY FGV++LE +TG+ PV+YG N + L D
Sbjct: 23 VTTRVMGTFGYVAPEYANTGL-LNEKSDVYSFGVLLLEGITGRDPVDYGRPTNEVNLVDW 199
Query: 60 VRVLLEHGNALECVDPSL 77
++ ++ + E VDP++
Sbjct: 200LK-MVGSRRSEEVVDPNI 250
>TC17680 similar to UP|O65440 (O65440) CLV1 receptor kinase like protein,
partial (16%)
Length = 685
Score = 53.5 bits (127), Expect = 1e-08
Identities = 26/48 (54%), Positives = 39/48 (81%), Gaps = 1/48 (2%)
Frame = +1
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPV-EYGEDNVLIL 56
GY+APE A L+V+EK DVY FGV++LE++TG+RPV ++GE+ + I+
Sbjct: 160 GYIAPEYA-YTLKVDEKSDVYSFGVVLLELLTGRRPVGDFGEEGLNIV 300
>AW719817
Length = 499
Score = 53.1 bits (126), Expect = 1e-08
Identities = 36/89 (40%), Positives = 50/89 (55%), Gaps = 2/89 (2%)
Frame = +1
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGK--RPVEYGEDNVLILND 58
+S R GY+APE A ++ K DVY FGV+ILE+V+GK +G N +L +
Sbjct: 124 ISTRIAGTTGYLAPEYAMGG-QLTMKADVYSFGVLILEVVSGKGSARTNWGGSNKYLL-E 297
Query: 59 HVRVLLEHGNALECVDPSLMSEYPEDEVL 87
L E G LE VDP M ++PE+ V+
Sbjct: 298 WAWQLHEEGKLLELVDPD-MDDFPEE*VI 381
>TC14756 weakly similar to UP|Q41944 (Q41944) Receptor-like protein kinase
like (Fragment), partial (60%)
Length = 566
Score = 52.8 bits (125), Expect = 2e-08
Identities = 33/70 (47%), Positives = 45/70 (64%), Gaps = 10/70 (14%)
Frame = +3
Query: 13 APELACQILRVNEKCDVYGFGVMILEIVTGKRPV--EYGEDNVLI------LNDHVRVL- 63
APELA I + EK DVY FGV++LE+V+G++P+ EYGE ++ LNDH +L
Sbjct: 3 APELAYTI-DITEKSDVYSFGVVLLELVSGRKPIEEEYGEAKDIVYWVLTHLNDHESILN 179
Query: 64 -LEHGNALEC 72
L+ ALEC
Sbjct: 180ILDDRVALEC 209
>TC16248 similar to UP|BAC87845 (BAC87845) Leucine-rich repeat receptor-like
protein kinase 1, partial (17%)
Length = 866
Score = 52.8 bits (125), Expect = 2e-08
Identities = 25/41 (60%), Positives = 28/41 (67%)
Frame = +2
Query: 5 FQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRP 45
F GY APELA Q ++VNEKCDVY FGV LEI+ G P
Sbjct: 353 FAGTFGYAAPELA-QTMQVNEKCDVYSFGVFALEIIMGMHP 472
>TC11055 similar to UP|Q9LR04 (Q9LR04) F10A5.16, partial (12%)
Length = 590
Score = 51.6 bits (122), Expect = 4e-08
Identities = 27/75 (36%), Positives = 46/75 (61%)
Frame = +1
Query: 9 LGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVRVLLEHGN 68
+GY++PE A +++ DVY FG+++LE++TGKRPV + +D ++ V+ L G
Sbjct: 91 VGYISPE-AILTGEASKESDVYSFGIVLLELLTGKRPVMFTQDEDIV--KWVKKQLPKGQ 261
Query: 69 ALECVDPSLMSEYPE 83
E ++P L+ PE
Sbjct: 262 ITELLEPGLLELDPE 306
>BI419694
Length = 623
Score = 51.6 bits (122), Expect = 4e-08
Identities = 32/89 (35%), Positives = 47/89 (51%), Gaps = 4/89 (4%)
Frame = +3
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY----GEDNVLILN 57
S R GY APE A ++N K DVY FGV++LE++TG++PV++ G+ +++
Sbjct: 123 STRVLGTFGYHAPEYA-MTGQLNAKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLV--- 290
Query: 58 DHVRVLLEHGNALECVDPSLMSEYPEDEV 86
L +CVD L EYP V
Sbjct: 291 TWATPKLSEDKVRQCVDARLGGEYPPKAV 377
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 51.2 bits (121), Expect = 5e-08
Identities = 36/84 (42%), Positives = 52/84 (61%), Gaps = 6/84 (7%)
Frame = +2
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPV-EYGE--DNVLILNDHVRVLLEH 66
GY+APE A L+V+EK DVY FGV++LE++ G++PV E+G+ D V +N + L +
Sbjct: 2693 GYIAPEYA-YTLKVDEKSDVYSFGVVLLELIIGRKPVGEFGDGVDIVGWVNKTMSELSQP 2869
Query: 67 GN---ALECVDPSLMSEYPEDEVL 87
+ L VDP L S YP V+
Sbjct: 2870 SDTALVLAVVDPRL-SGYPLTSVI 2938
>TC10812 similar to UP|O65440 (O65440) CLV1 receptor kinase like protein,
partial (14%)
Length = 671
Score = 50.8 bits (120), Expect = 7e-08
Identities = 23/37 (62%), Positives = 31/37 (83%)
Frame = +3
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPV 46
GY+APE A L+V+EK DVY FGV++LE++TG+RPV
Sbjct: 105 GYIAPEYA-YTLKVDEKSDVYSFGVVLLELITGRRPV 212
>TC16403 homologue to UP|Q9LKY4 (Q9LKY4) Pti1 kinase-like protein, partial
(55%)
Length = 876
Score = 50.4 bits (119), Expect = 9e-08
Identities = 31/89 (34%), Positives = 46/89 (50%), Gaps = 4/89 (4%)
Frame = +3
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY----GEDNVLILN 57
S R GY APE A ++ K DVY FGV++LE++TG++PV++ G+ +++
Sbjct: 231 STRVLGTFGYHAPEYA-MTGQLTSKSDVYSFGVVLLELLTGRKPVDHTLPRGQQSLV--- 398
Query: 58 DHVRVLLEHGNALECVDPSLMSEYPEDEV 86
L +CVD L EYP V
Sbjct: 399 TWATPKLSEDKVKQCVDARLKGEYPVKSV 485
>AV425746
Length = 415
Score = 50.4 bits (119), Expect = 9e-08
Identities = 28/83 (33%), Positives = 47/83 (55%), Gaps = 3/83 (3%)
Frame = +1
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY---GEDNVLILN 57
++ R GY+APE A ++ EK DVY FGV++LEI+ G++ ++ G ++
Sbjct: 43 LTTRVAGTHGYLAPEYALYG-QLTEKSDVYSFGVVVLEILCGRKALDLSSSGSPRAFLIT 219
Query: 58 DHVRVLLEHGNALECVDPSLMSE 80
D L++ GN E +D SL+ +
Sbjct: 220 DWAWSLVKSGNIDEALDSSLLKD 288
>BP053923
Length = 471
Score = 48.1 bits (113), Expect = 4e-07
Identities = 20/26 (76%), Positives = 22/26 (83%)
Frame = -2
Query: 63 LLEHGNALECVDPSLMSEYPEDEVLP 88
LLEHGN LECVDPS+ EYP+DEV P
Sbjct: 470 LLEHGNLLECVDPSMSHEYPDDEVXP 393
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.141 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,401,280
Number of Sequences: 28460
Number of extensions: 14167
Number of successful extensions: 204
Number of sequences better than 10.0: 151
Number of HSP's better than 10.0 without gapping: 189
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 190
length of query: 126
length of database: 4,897,600
effective HSP length: 80
effective length of query: 46
effective length of database: 2,620,800
effective search space: 120556800
effective search space used: 120556800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 50 (23.9 bits)
Medicago: description of AC146632.13


