

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146631.5 - phase: 0 /pseudo
(359 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
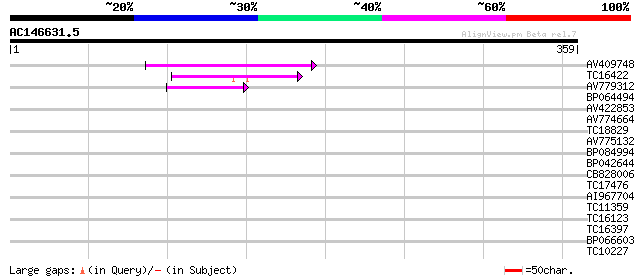
Score E
Sequences producing significant alignments: (bits) Value
AV409748 47 4e-06
TC16422 similar to PIR|T05653|T05653 amino acid transport protei... 43 7e-05
AV779312 42 1e-04
BP064494 31 0.28
AV422853 30 0.48
AV774664 30 0.63
TC18829 homologue to UP|Q8L4X4 (Q8L4X4) Amino acid permease-like... 29 1.4
AV775132 28 1.8
BP084994 28 1.8
BP042644 28 2.4
CB828006 28 2.4
TC17476 similar to UP|Q9SAG9 (Q9SAG9) F23A5.27, partial (44%) 28 2.4
AI967704 28 3.1
TC11359 27 6.9
TC16123 similar to UP|PGC2_HUMAN (O15173) Membrane associated pr... 27 6.9
TC16397 UP|ACCD_LOTJA (Q9BBS1) Acetyl-coenzyme A carboxylase car... 27 6.9
BP066603 26 9.0
TC10227 UP|Q7Q2V3 (Q7Q2V3) EbiP1409 (Fragment), partial (10%) 26 9.0
>AV409748
Length = 417
Score = 47.4 bits (111), Expect = 4e-06
Identities = 29/108 (26%), Positives = 50/108 (45%)
Frame = +2
Query: 87 GYGCSYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFES 146
G S+T +VFN + G G+++ P +K G + +IF + T+ + F
Sbjct: 86 GAAASFTGSVFNLSTTIIGAGIMALPAAIKVLGVAAGIAAIIFLAFLTETSLDILMRFSR 265
Query: 147 REGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTG 194
SY D+ AFG GR+ + I + + V ++ + GD L+G
Sbjct: 266 VGNARSYGDVMGYAFGWVGRLVLQISVLINNFGILVVYVIIIGDVLSG 409
>TC16422 similar to PIR|T05653|T05653 amino acid transport protein homolog
F22I13.20 - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (38%)
Length = 685
Score = 43.1 bits (100), Expect = 7e-05
Identities = 25/90 (27%), Positives = 43/90 (47%), Gaps = 7/90 (7%)
Frame = +2
Query: 103 MAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLM----RHCFESREG---LTSYPD 155
+ G G+L PY K+ GW+ ++ML + Y ++ R +S G L S+ D
Sbjct: 221 IVGAGVLGLPYCFKRTGWLTGLIMLFTVAFLTYHCMMLLVSTRRKLDSLTGYSKLGSFGD 400
Query: 156 IGEAAFGRYGRIFVSIILYTELYSYCVEFI 185
+G A G GR+ V ++ +CV ++
Sbjct: 401 LGFAICGPLGRLAVDAMIVLSQAGFCVSYL 490
>AV779312
Length = 345
Score = 42.4 bits (98), Expect = 1e-04
Identities = 20/53 (37%), Positives = 32/53 (59%), Gaps = 1/53 (1%)
Frame = +3
Query: 100 INVMAGVGLLSTPYTVKQAGW-MGLVLMLIFASVCCYTATLMRHCFESREGLT 151
I + G G+LS + + Q GW +G+ +LIFA V YT+ L+ C+ S + +T
Sbjct: 132 ITAVIGAGVLSLAWAMAQLGWILGVSCILIFAGVTLYTSNLLADCYRSPDPVT 290
>BP064494
Length = 354
Score = 31.2 bits (69), Expect = 0.28
Identities = 19/70 (27%), Positives = 31/70 (44%), Gaps = 6/70 (8%)
Frame = +3
Query: 276 AIGVYGFCFAGHSVFPNIYQSM------ADKKQYTKALITCFVLCILIYGSVAVMGFLSF 329
A+G F +AGH+V I +M KK K ++ ++ Y VA +G+ F
Sbjct: 141 AMGEVAFSYAGHNVVLEIQATMPSTPEKPSKKPMWKGVVVAYIGVAFCYLPVAFIGYYVF 320
Query: 330 GDDTLSQITL 339
G+ I +
Sbjct: 321 GNSVDDNILI 350
>AV422853
Length = 456
Score = 30.4 bits (67), Expect = 0.48
Identities = 21/62 (33%), Positives = 29/62 (45%), Gaps = 5/62 (8%)
Frame = -2
Query: 254 SVGTTVGFHHTGRVVNWSGIPFAIGVYGF-----CFAGHSVFPNIYQSMADKKQYTKALI 308
S+ + HHTGR+ W GIP AI A H V+ + MA+ K +L+
Sbjct: 287 SLALRISLHHTGRIEGW-GIPRAISPMALRN*TNPTASHMVW---FNRMANMKPPHFSLV 120
Query: 309 TC 310
TC
Sbjct: 119 TC 114
>AV774664
Length = 405
Score = 30.0 bits (66), Expect = 0.63
Identities = 13/21 (61%), Positives = 15/21 (70%)
Frame = +1
Query: 104 AGVGLLSTPYTVKQAGWMGLV 124
AG G+L TP+ VK GWM LV
Sbjct: 340 AGSGILCTPHGVKGTGWMKLV 402
>TC18829 homologue to UP|Q8L4X4 (Q8L4X4) Amino acid permease-like protein,
partial (27%)
Length = 713
Score = 28.9 bits (63), Expect = 1.4
Identities = 19/86 (22%), Positives = 35/86 (40%), Gaps = 5/86 (5%)
Frame = +1
Query: 97 FNGINVMAGVGLLSTPYTVKQAGW-MGLVLMLIFASVCCYTATLM----RHCFESREGLT 151
F+ + G +L+ PY + GW +G V + + +V Y LM HC +S
Sbjct: 196 FHLTTAIVGPTILTLPYAFRGLGWVLGFVCLTVMGAVTFYAYFLMSKVLEHCEKSGRRHI 375
Query: 152 SYPDIGEAAFGRYGRIFVSIILYTEL 177
+ ++ G + I + T +
Sbjct: 376 RFRELAADVLGSGWMYYFVIFIQTAI 453
>AV775132
Length = 465
Score = 28.5 bits (62), Expect = 1.8
Identities = 11/21 (52%), Positives = 12/21 (56%)
Frame = +1
Query: 62 GLDGTTQSTWWEKRSTQNLVG 82
GLDG T WWE+ S Q G
Sbjct: 16 GLDGPTLWRWWERASVQQFCG 78
>BP084994
Length = 487
Score = 28.5 bits (62), Expect = 1.8
Identities = 15/75 (20%), Positives = 33/75 (44%), Gaps = 2/75 (2%)
Frame = +2
Query: 242 IAATILIIISVFSVGTTVGFHHTGRVVNWS--GIPFAIGVYGFCFAGHSVFPNIYQSMAD 299
++ + +++ V T F + G+++NW +P + F H + Y +
Sbjct: 98 LSPLLFLLLQTLDVVTGDQFLNNGKMLNWKLDSVPGIVD-----FKNHRCYQFQYHITLE 262
Query: 300 KKQYTKALITCFVLC 314
K + +ITC ++C
Sbjct: 263 KTIFCHNIITCSIIC 307
>BP042644
Length = 440
Score = 28.1 bits (61), Expect = 2.4
Identities = 22/65 (33%), Positives = 31/65 (46%)
Frame = +1
Query: 19 DSYTIATAPNFVSILRGPSSIYSSFSNRSKSDLDIDGKTPCLSGLDGTTQSTWWEKRSTQ 78
+S+TIA R +S SS + R +D ID CL +TQ +WE+ T+
Sbjct: 64 ESFTIAAC-----CARQRASPSSSHAGRYFADAGIDDGEGCLQRA-FSTQDMFWEEERTR 225
Query: 79 NLVGE 83
N GE
Sbjct: 226 N*PGE 240
>CB828006
Length = 567
Score = 28.1 bits (61), Expect = 2.4
Identities = 37/124 (29%), Positives = 49/124 (38%), Gaps = 11/124 (8%)
Frame = +2
Query: 181 CVEFITLEGDNLTGLFPGT-SLDIGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLS- 238
CV FI G L LF D L + F V T + IL L +L ++ +S
Sbjct: 119 CVMFIITGGRTLKLLFNTLCDNDCDAHSLTGVEWFLVFTCVAIL-IAQLPNLHSLTAVSL 295
Query: 239 VGGIAA----TILIIISV-----FSVGTTVGFHHTGRVVNWSGIPFAIGVYGFCFAGHSV 289
VG + A T+ +SV V T H V S + AIG+ F GH+V
Sbjct: 296 VGAVTAITYCTLFWFLSVKKGRPHGVSYTSSLSHGSAVDKISDVLNAIGIIVLAFRGHNV 475
Query: 290 FPNI 293
I
Sbjct: 476 LLEI 487
>TC17476 similar to UP|Q9SAG9 (Q9SAG9) F23A5.27, partial (44%)
Length = 804
Score = 28.1 bits (61), Expect = 2.4
Identities = 11/16 (68%), Positives = 11/16 (68%)
Frame = +2
Query: 69 STWWEKRSTQNLVGEM 84
S WW K TQNL GEM
Sbjct: 395 SWWWRKPMTQNLYGEM 442
>AI967704
Length = 385
Score = 27.7 bits (60), Expect = 3.1
Identities = 11/46 (23%), Positives = 23/46 (49%)
Frame = +2
Query: 287 HSVFPNIYQSMADKKQYTKALITCFVLCILIYGSVAVMGFLSFGDD 332
H P + + D+ ++ K L F L+Y + G+++FG++
Sbjct: 248 HPPSPELKEQQKDRSKFPKLLAQTFSGITLVYILFGLCGYMAFGEE 385
>TC11359
Length = 544
Score = 26.6 bits (57), Expect = 6.9
Identities = 11/21 (52%), Positives = 14/21 (66%)
Frame = -3
Query: 177 LYSYCVEFITLEGDNLTGLFP 197
+ SYC+ F+ L GDNL FP
Sbjct: 323 ILSYCLLFVDLFGDNLHHSFP 261
>TC16123 similar to UP|PGC2_HUMAN (O15173) Membrane associated progesterone
receptor component 2 (Progesterone membrane binding
protein) (Steroid receptor protein DG6), partial (15%)
Length = 604
Score = 26.6 bits (57), Expect = 6.9
Identities = 20/60 (33%), Positives = 29/60 (48%), Gaps = 3/60 (5%)
Frame = -2
Query: 216 VLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGTTVGFHH---TGRVVNWSG 272
VL+ ++LP + DL V + VGG +L + S G + G H TG V +SG
Sbjct: 363 VLSYELVLPVIENADLFVGEAIEVGGDIFLVLAHLR*GSAGVSSGEHRVSTTGAVEGFSG 184
>TC16397 UP|ACCD_LOTJA (Q9BBS1) Acetyl-coenzyme A carboxylase carboxyl
transferase subunit beta (ACCASE beta chain) , complete
Length = 1514
Score = 26.6 bits (57), Expect = 6.9
Identities = 15/45 (33%), Positives = 24/45 (53%), Gaps = 1/45 (2%)
Frame = +3
Query: 271 SGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQY-TKALITCFVLC 314
+GIP AIG+ F F G S+ + + + +Y T L+ V+C
Sbjct: 981 NGIPVAIGIMDFEFMGGSMGSVVGEKITRLVEYATNQLLPLIVVC 1115
>BP066603
Length = 446
Score = 26.2 bits (56), Expect = 9.0
Identities = 8/13 (61%), Positives = 13/13 (99%)
Frame = -3
Query: 172 ILYTELYSYCVEF 184
I+YT++YS+C+EF
Sbjct: 75 IMYTQIYSWCLEF 37
>TC10227 UP|Q7Q2V3 (Q7Q2V3) EbiP1409 (Fragment), partial (10%)
Length = 624
Score = 26.2 bits (56), Expect = 9.0
Identities = 11/43 (25%), Positives = 20/43 (45%)
Frame = +3
Query: 142 HCFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEF 184
HC R+ + +G + G+Y +S+ +Y +Y Y F
Sbjct: 486 HCISGRKD*LACMALGSGSHGKYDVSSLSLSIYIYIYIYIYHF 614
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.140 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,686,585
Number of Sequences: 28460
Number of extensions: 91261
Number of successful extensions: 588
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 586
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 587
length of query: 359
length of database: 4,897,600
effective HSP length: 91
effective length of query: 268
effective length of database: 2,307,740
effective search space: 618474320
effective search space used: 618474320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146631.5


