

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
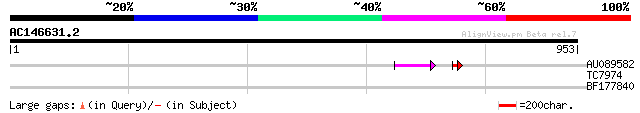
Query= AC146631.2 + phase: 0 /pseudo
(953 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AU089582 56 2e-08
TC7974 similar to GB|AAA77675.1|607870|DSU11801 period protein {... 31 1.1
BF177840 28 6.8
>AU089582
Length = 383
Score = 55.8 bits (133), Expect(2) = 2e-08
Identities = 27/69 (39%), Positives = 39/69 (56%)
Frame = +1
Query: 647 DLYS*YCEVAWSSVEYCVGKRPEVHFKILEEFARCIGFKVEVEFGLSSANRWSVGENDSI 706
DL * C +AW + Y R +H LE F+ C G ++ E+ SS+N WSV E+ S
Sbjct: 28 DLLG*DCFLAWCTCVYNFRSRSSIHITFLEVFSNCFGNSIKNEYRFSSSN*WSVREDYSD 207
Query: 707 VRGFAKRMC 715
+RG+A +C
Sbjct: 208 LRGYASCLC 234
Score = 20.4 bits (41), Expect(2) = 2e-08
Identities = 8/18 (44%), Positives = 13/18 (71%)
Frame = +2
Query: 744 GTVQSFVWSEVEDSFMLV 761
GT+ S VW E++ ++ LV
Sbjct: 320 GTI*SLVW*EMQVTYRLV 373
>TC7974 similar to GB|AAA77675.1|607870|DSU11801 period protein {Drosophila
simulans;} , partial (46%)
Length = 689
Score = 30.8 bits (68), Expect = 1.1
Identities = 15/36 (41%), Positives = 22/36 (60%)
Frame = -3
Query: 138 MNIR*CLSVILMRLLCLWSI*TEFSMLIWINLWLCL 173
+N+ CL + L + LCL+ * F +W+ LWLCL
Sbjct: 99 LNLYQCLCLCLFQCLCLFLC*FLF---LWVCLWLCL 1
>BF177840
Length = 410
Score = 28.1 bits (61), Expect = 6.8
Identities = 15/39 (38%), Positives = 20/39 (50%)
Frame = +1
Query: 661 EYCVGKRPEVHFKILEEFARCIGFKVEVEFGLSSANRWS 699
E+ VG ++H LE G KV V + LSS N W+
Sbjct: 13 EHSVGSGHQIHKPFLENIMG*GGNKVVVLYNLSSTNGWA 129
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.373 0.168 0.680
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,546,860
Number of Sequences: 28460
Number of extensions: 287397
Number of successful extensions: 3529
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 1868
Number of HSP's successfully gapped in prelim test: 96
Number of HSP's that attempted gapping in prelim test: 1599
Number of HSP's gapped (non-prelim): 2070
length of query: 953
length of database: 4,897,600
effective HSP length: 99
effective length of query: 854
effective length of database: 2,080,060
effective search space: 1776371240
effective search space used: 1776371240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146631.2


