

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146588.5 - phase: 0 /pseudo
(637 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
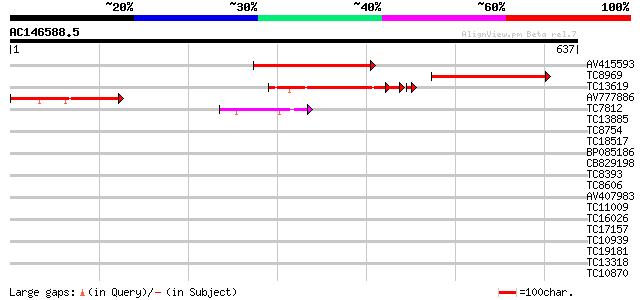
Score E
Sequences producing significant alignments: (bits) Value
AV415593 220 6e-58
TC8969 209 1e-54
TC13619 similar to UP|Q9LFQ3 (Q9LFQ3) Transcriptional regulatory... 148 3e-43
AV777886 125 2e-29
TC7812 UP|Q9LKJ6 (Q9LKJ6) Water-selective transport intrinsic me... 42 4e-04
TC13885 weakly similar to PIR|T14734|T14734 NS5 protein - sorghu... 40 0.001
TC8754 similar to UP|Q9LFQ3 (Q9LFQ3) Transcriptional regulatory-... 40 0.001
TC18517 weakly similar to UP|AAQ82688 (AAQ82688) Epa4p, partial ... 37 0.013
BP085186 35 0.028
CB829198 35 0.037
TC8393 weakly similar to GB|AAM61124.1|21536792|AY084557 prenyla... 35 0.037
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 34 0.063
AV407983 34 0.063
TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Ar... 33 0.11
TC16026 similar to UP|Q9SGZ3 (Q9SGZ3) F28K19.29, partial (32%) 32 0.31
TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly pr... 31 0.53
TC10939 similar to UP|Q941N0 (Q941N0) Drought-induced protein, p... 31 0.53
TC19181 similar to UP|Q40090 (Q40090) SPF1 protein, partial (19%) 31 0.70
TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (... 30 0.91
TC10870 similar to GB|CAA77438.1|2924285|CHNTXX Ycf2 protein {Ni... 30 0.91
>AV415593
Length = 416
Score = 220 bits (560), Expect = 6e-58
Identities = 109/137 (79%), Positives = 118/137 (85%)
Frame = +3
Query: 275 SPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVE 334
SPHRSSDDSENASEN DVSG+ESADGEECSREEHE D +HDNK ESEGEAEGM DA+DVE
Sbjct: 6 SPHRSSDDSENASENVDVSGSESADGEECSREEHE-DREHDNKAESEGEAEGMVDAHDVE 182
Query: 335 GDGASLPYSERFLLTAKPLVKYVSPVFHGKEENVQIFYGNDSFYVLFRLHQTLYERIRSA 394
GDG SL SERFL T KPL KYV H K+ N Q+FYG+DSFYVLFRLHQTLYERI+SA
Sbjct: 183 GDGMSLTISERFLQTVKPLAKYVPAALHEKDRNSQVFYGSDSFYVLFRLHQTLYERIQSA 362
Query: 395 KINSSSAEKKWRASNDT 411
K NSSSAEKKW+AS+DT
Sbjct: 363 KFNSSSAEKKWKASHDT 413
>TC8969
Length = 892
Score = 209 bits (531), Expect = 1e-54
Identities = 104/136 (76%), Positives = 115/136 (84%), Gaps = 2/136 (1%)
Frame = +1
Query: 474 DEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRIECSP--TRLSIQLMDYG 531
+EMDNKLLQLYAYE+SRK G F D+VYHENA V+LH+ENIYR E SP +LSIQLMD G
Sbjct: 1 EEMDNKLLQLYAYEKSRKLGKFIDIVYHENACVILHEENIYRXEYSPGPKKLSIQLMDCG 180
Query: 532 HDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKSGIFLKRNKLKCARSDELSSQVMDGIQ 591
HDKPEVTAVS+DP FSTYLHNDFLSVVPD KEKS +FLKRNK + A S E SSQ M+G+Q
Sbjct: 181 HDKPEVTAVSVDPKFSTYLHNDFLSVVPDKKEKSRLFLKRNKHRYAGSSEFSSQAMEGLQ 360
Query: 592 VINGLECKIACGSSKV 607
V NGLECKIAC SSKV
Sbjct: 361 VFNGLECKIACSSSKV 408
>TC13619 similar to UP|Q9LFQ3 (Q9LFQ3) Transcriptional regulatory-like
protein, partial (6%)
Length = 537
Score = 148 bits (374), Expect(3) = 3e-43
Identities = 86/142 (60%), Positives = 100/142 (69%), Gaps = 5/142 (3%)
Frame = +2
Query: 291 DVSGTESADGEECSREEHEEDG----DHDNKVESEGEAEGMADANDVEGDGASLPYSERF 346
DVSG+ESA G+EC +EEHEED D D K ESEGEAEGM DA GD +SLP SERF
Sbjct: 29 DVSGSESA-GDECFQEEHEEDDIEHDDVDGKAESEGEAEGMCDAQG-GGDSSSLP*SERF 202
Query: 347 LLTAKPLVKYVSPVFHGKE-ENVQIFYGNDSFYVLFRLHQTLYERIRSAKINSSSAEKKW 405
L + KPL K+VS V +E ++ ++ GND FY LFRLHQ LYERI SAKINS SAE KW
Sbjct: 203 LSSVKPLTKHVSAVSFAEEMKDSRVLNGNDDFYALFRLHQILYERILSAKINSMSAEMKW 382
Query: 406 RASNDTSSTDRYARFMNSLYSL 427
+A D SS D Y+RFMN + L
Sbjct: 383 KA-KDASSPDPYSRFMNGIVQL 445
Score = 38.9 bits (89), Expect(3) = 3e-43
Identities = 16/21 (76%), Positives = 20/21 (95%)
Frame = +3
Query: 423 SLYSLLDGSSDNSKFEDDCRA 443
+LY+LLDGS +NSKFED+CRA
Sbjct: 432 ALYNLLDGSVENSKFEDECRA 494
Score = 26.2 bits (56), Expect(3) = 3e-43
Identities = 11/12 (91%), Positives = 11/12 (91%)
Frame = +1
Query: 446 GTQSYVLFTLDK 457
G QSYVLFTLDK
Sbjct: 502 GNQSYVLFTLDK 537
>AV777886
Length = 540
Score = 125 bits (314), Expect = 2e-29
Identities = 72/147 (48%), Positives = 90/147 (60%), Gaps = 20/147 (13%)
Frame = +3
Query: 1 MKIWTTFLEPMLGVPSRLRVPEDTEDAVKAKK---------DSAKTGTASIAKGDSSPGV 51
MK+WTTFLEPML VPSR + EDTED VKAK +S K+G S+A+ +PGV
Sbjct: 102 MKVWTTFLEPMLCVPSRAQGAEDTEDVVKAKNTEDVVNTENNSVKSGIVSVAESGGNPGV 281
Query: 52 GATVMSPKNT-----------FEQSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGSS 100
GA M+PK+ +QS+S K WQ NG GV+ED CL SD +V KTET G++
Sbjct: 282 GAITMNPKHMSTSRNGSECMPLDQSDS-KAWQLNGDSGVREDRCLDSDHTVYKTETAGNN 458
Query: 101 TLQGNVHINASIPDEVSRVNKQDHSIE 127
T G +I PDE+S VNKQ+ S E
Sbjct: 459 TQHGKTNIIGFTPDELSGVNKQNQSSE 539
>TC7812 UP|Q9LKJ6 (Q9LKJ6) Water-selective transport intrinsic membrane
protein 1, complete
Length = 1149
Score = 41.6 bits (96), Expect = 4e-04
Identities = 30/115 (26%), Positives = 54/115 (46%), Gaps = 10/115 (8%)
Frame = -2
Query: 236 GLEAVHKGKDCNTSQHYQ------NRREEQICGVAGGENDDES-DGSPHRSSDDSENASE 288
G+ VH+ K + QH + + + E +G E +++ D + SDD ++ +E
Sbjct: 554 GVNCVHQSKGHHDLQHQRVPNSNSS*KTESRYSESGDEGEEQGGDDGSEKLSDDVDDTAE 375
Query: 289 NGDVSGTESADGE---ECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASL 340
GDV+ E A+G+ + + + DGD D EGE G ++ G G ++
Sbjct: 374 EGDVTADEEAEGDGGVDVAAGDVGADGDGD----EEGECVGDGGGDEAGGGGGAV 222
>TC13885 weakly similar to PIR|T14734|T14734 NS5 protein - sorghum
(fragment) {Sorghum bicolor;} , partial (15%)
Length = 410
Score = 40.0 bits (92), Expect = 0.001
Identities = 17/25 (68%), Positives = 21/25 (84%)
Frame = +3
Query: 583 SSQVMDGIQVINGLECKIACGSSKV 607
SS M+G+Q+ NGLEC+IAC SSKV
Sbjct: 87 SS*AMEGLQIFNGLECEIACSSSKV 161
>TC8754 similar to UP|Q9LFQ3 (Q9LFQ3) Transcriptional regulatory-like
protein, partial (3%)
Length = 566
Score = 40.0 bits (92), Expect = 0.001
Identities = 18/30 (60%), Positives = 25/30 (83%), Gaps = 2/30 (6%)
Frame = +2
Query: 580 DELSS--QVMDGIQVINGLECKIACGSSKV 607
D+LS+ M+G++VINGLECKIAC SS++
Sbjct: 8 DDLSAICAAMEGVKVINGLECKIACNSSQI 97
>TC18517 weakly similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (4%)
Length = 493
Score = 36.6 bits (83), Expect = 0.013
Identities = 27/97 (27%), Positives = 39/97 (39%), Gaps = 26/97 (26%)
Frame = -2
Query: 266 GENDDESDGSPHRSSDDS---------------ENASENGDVSGTESADGEECSREEHEE 310
GEN+ + +P R D++ E E G+ G E DG+ E EE
Sbjct: 474 GENERNNGNNPERGEDETVIDGAGEDNDRAVAEEEEEEPGEEDGEEDDDGDGVPDEGDEE 295
Query: 311 DGDHDNKVESE-----------GEAEGMADANDVEGD 336
D + D+ V +E G AEG+ VEG+
Sbjct: 294 DEEEDHGVVAEEVAEVVLHAEGGFAEGVRAGEGVEGE 184
>BP085186
Length = 388
Score = 35.4 bits (80), Expect = 0.028
Identities = 17/46 (36%), Positives = 26/46 (55%)
Frame = -1
Query: 289 NGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVE 334
NG + ++ADGE EE EE+ + + + E E EAE + N+ E
Sbjct: 304 NGKLISPDNADGEGNEEEEEEEEEEEEEEEEEEEEAEPEEEKNEEE 167
Score = 32.7 bits (73), Expect = 0.18
Identities = 19/61 (31%), Positives = 28/61 (45%)
Frame = -1
Query: 288 ENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERFL 347
+N D G E + EE EE EE+ + + + E E E + + EG+ L FL
Sbjct: 283 DNADGEGNEEEEEEEEEEEEEEEEEEEEAEPEEEKNEEENVNTQEGEGENP*LQVLLLFL 104
Query: 348 L 348
L
Sbjct: 103 L 101
>CB829198
Length = 562
Score = 35.0 bits (79), Expect = 0.037
Identities = 14/43 (32%), Positives = 25/43 (57%)
Frame = +3
Query: 282 DSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEA 324
D + +N DV ++ +D +E + EE +EDGD++N +A
Sbjct: 246 DDQEEPQNDDVEESQHSDSKEETEEEEDEDGDNENNKPKRAKA 374
>TC8393 weakly similar to GB|AAM61124.1|21536792|AY084557 prenylated Rab
receptor 2 {Arabidopsis thaliana;}, partial (78%)
Length = 1012
Score = 35.0 bits (79), Expect = 0.037
Identities = 30/112 (26%), Positives = 49/112 (42%), Gaps = 4/112 (3%)
Frame = -3
Query: 230 AVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENASEN 289
A+ AN G +A H+ +D + + + E++ CG E + +G+ + A E
Sbjct: 590 AMEANKG-DAEHESRDEDGTDTGEENDEDRQCG-ENRERLEIGEGAAEENERAIRRAEEV 417
Query: 290 GDVSGTESA----DGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDG 337
+ G+E A + E E H ED +D V E +ADA G+G
Sbjct: 416 EECPGSEKAGQGDERERIGEERHGEDERNDGVVVDAEVGEILADAEGGIGEG 261
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 34.3 bits (77), Expect = 0.063
Identities = 19/74 (25%), Positives = 32/74 (42%), Gaps = 1/74 (1%)
Frame = -2
Query: 273 DGSPHRSSDDSENASENGDVSGTESADGEECSREEHEE-DGDHDNKVESEGEAEGMADAN 331
+G P + EN + D G + D ++ ++ EE DG D + E E + +
Sbjct: 473 EGGPGAGAGPEENGEDEEDEDGEDQEDDDDDDEDDDEEDDGGEDEEEEGVEEEDNEDEEE 294
Query: 332 DVEGDGASLPYSER 345
D E + A P +R
Sbjct: 293 DEEDEEALQPPKKR 252
Score = 28.9 bits (63), Expect = 2.6
Identities = 20/71 (28%), Positives = 30/71 (42%), Gaps = 2/71 (2%)
Frame = -2
Query: 268 NDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGM 327
N S +P ENG+ E + +E ++ E+D + D+ E E E EG+
Sbjct: 500 NKSNSKKAPEGGPGAGAGPEENGEDEEDEDGEDQEDDDDDDEDDDEEDDGGEDE-EEEGV 324
Query: 328 --ADANDVEGD 336
D D E D
Sbjct: 323 EEEDNEDEEED 291
>AV407983
Length = 417
Score = 34.3 bits (77), Expect = 0.063
Identities = 24/75 (32%), Positives = 30/75 (40%), Gaps = 2/75 (2%)
Frame = -2
Query: 264 AGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEE--CSREEHEEDGDHDNKVESE 321
+ E DE+D + E E G G EE C EE E G+ + E E
Sbjct: 341 SASEGFDEADEDEVEEGAEDEREEEVGAERGGRRGGEEEGECDGEEEAEIGEVF*EAEEE 162
Query: 322 GEAEGMADANDVEGD 336
EG + DVEGD
Sbjct: 161 FWGEGCGEGMDVEGD 117
>TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;}, partial (44%)
Length = 700
Score = 33.5 bits (75), Expect = 0.11
Identities = 24/92 (26%), Positives = 37/92 (40%), Gaps = 11/92 (11%)
Frame = +2
Query: 257 EEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTE--------SADGEECSREEH 308
EE+ GV GG DD + E + GTE +A+ EE S +
Sbjct: 125 EEEEGGVEGGRGGGGDPDDDDDDDDDDDEEEEEEEDLGTEYLVRRTVAAAEDEEASSDFE 304
Query: 309 EEDGDHDNKVESEGEAEGMAD---ANDVEGDG 337
E+ + D+ +GE G+ +D +G G
Sbjct: 305 PEEDEGDDNDNDDGEKAGVPSKRKRSDKDGSG 400
Score = 28.9 bits (63), Expect = 2.6
Identities = 18/62 (29%), Positives = 28/62 (45%)
Frame = +2
Query: 265 GGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEA 324
GG++DD D E+ E G V G G+ ++ ++D D D++ E E E
Sbjct: 95 GGDDDD-----------DEEDEEEEGGVEGGRGGGGDP---DDDDDDDDDDDEEEEEEED 232
Query: 325 EG 326
G
Sbjct: 233 LG 238
>TC16026 similar to UP|Q9SGZ3 (Q9SGZ3) F28K19.29, partial (32%)
Length = 1183
Score = 32.0 bits (71), Expect = 0.31
Identities = 19/48 (39%), Positives = 25/48 (51%), Gaps = 3/48 (6%)
Frame = -2
Query: 282 DSENASENGDVSGTES---ADGEECSREEHEEDGDHDNKVESEGEAEG 326
D E SE V+ E GEE R+E +EDG+ + + E E E EG
Sbjct: 780 DVEQPSEKKGVTEGEE*KELGGEEQGRDEFDEDGEGEEEEEEEEEEEG 637
>TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly protein
1-like protein 3, partial (81%)
Length = 1298
Score = 31.2 bits (69), Expect = 0.53
Identities = 18/61 (29%), Positives = 27/61 (43%)
Frame = +2
Query: 281 DDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASL 340
+D E E+GD E D +E EED D D++ + EGE + + + G
Sbjct: 764 EDIEEDDEDGDEDEDEDDDDDE-----EEEDDDDDDEDDEEGEGKSKSKSGSKARPGKDQ 928
Query: 341 P 341
P
Sbjct: 929 P 931
Score = 29.6 bits (65), Expect = 1.5
Identities = 17/50 (34%), Positives = 27/50 (54%)
Frame = +2
Query: 290 GDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGAS 339
G+ ++ D EE E+ +ED D D+ + E E + D +D EG+G S
Sbjct: 740 GEAEQSDFEDIEE-DDEDGDEDEDEDDDDDEEEEDDDDDDEDDEEGEGKS 886
>TC10939 similar to UP|Q941N0 (Q941N0) Drought-induced protein, partial
(59%)
Length = 564
Score = 31.2 bits (69), Expect = 0.53
Identities = 19/92 (20%), Positives = 37/92 (39%), Gaps = 6/92 (6%)
Frame = +3
Query: 238 EAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESDGSPHRSS-----DDSENASENGDV 292
EA+H G + + + E G D+ G H+ D + E G+
Sbjct: 45 EALHVGGGHKKEEEHHKKEEHHGDAHKGEHKSDQHHGGEHKEGIVDKIKDKIHGDEKGEK 224
Query: 293 SGTESA-DGEECSREEHEEDGDHDNKVESEGE 323
++ D ++ +++HE DHD+ S+ +
Sbjct: 225 HDEKNKKDKKDKKKKKHEHGHDHDSSSSSDSD 320
>TC19181 similar to UP|Q40090 (Q40090) SPF1 protein, partial (19%)
Length = 402
Score = 30.8 bits (68), Expect = 0.70
Identities = 23/65 (35%), Positives = 29/65 (44%), Gaps = 5/65 (7%)
Frame = +3
Query: 271 ESDGSPHRSSD-DSENASENGDVS----GTESADGEECSREEHEEDGDHDNKVESEGEAE 325
E G+ H S DS EN +S E + + EE +ED H K EGE E
Sbjct: 87 EIQGATHVSGQMDSVATPENSSLSMGGDDFEQSSMSKSGGEELDEDEPHGKKWRMEGENE 266
Query: 326 GMADA 330
GM+ A
Sbjct: 267 GMSAA 281
>TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (SPBC3D6.14C
protein) (Fragment), partial (15%)
Length = 543
Score = 30.4 bits (67), Expect = 0.91
Identities = 17/67 (25%), Positives = 34/67 (50%), Gaps = 1/67 (1%)
Frame = +3
Query: 273 DGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADAND 332
DG P D+E ++ + E D ++ ++ ++DG+ + + + E E ++ +
Sbjct: 219 DGPPKPVQTDNEEEEDDDEEEDDEDEDDDD--DDDDDDDGEEEEDDDDDDEEEEGSEEVE 392
Query: 333 VEG-DGA 338
VEG DGA
Sbjct: 393 VEGKDGA 413
>TC10870 similar to GB|CAA77438.1|2924285|CHNTXX Ycf2 protein {Nicotiana
tabacum;}, complete
Length = 6897
Score = 30.4 bits (67), Expect = 0.91
Identities = 13/42 (30%), Positives = 23/42 (53%)
Frame = +1
Query: 475 EMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRI 516
E+ N LY++ Q +GSF + +H+ + + L D I+ I
Sbjct: 55 EIKNSRYFLYSWTQFNSTGSFIHIFFHQESFIKLLDSRIWSI 180
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.129 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,688,810
Number of Sequences: 28460
Number of extensions: 122961
Number of successful extensions: 674
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 638
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 660
length of query: 637
length of database: 4,897,600
effective HSP length: 96
effective length of query: 541
effective length of database: 2,165,440
effective search space: 1171503040
effective search space used: 1171503040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146588.5


