

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
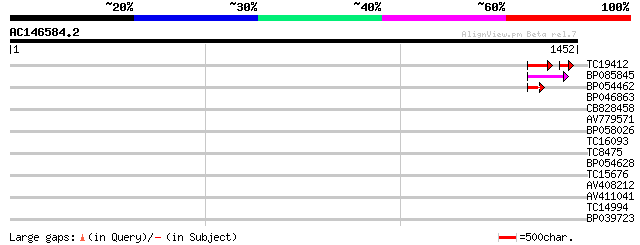
Query= AC146584.2 - phase: 0 /pseudo
(1452 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%) 86 1e-23
BP085845 59 5e-09
BP054462 49 7e-06
BP046863 40 0.003
CB828458 31 1.2
AV779571 31 1.6
BP058026 30 2.1
TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transc... 30 2.1
TC8475 similar to UP|GAL1_ARATH (Q9SEE5) Galactokinase (Galacto... 30 2.7
BP054628 30 3.6
TC15676 similar to GB|AAG24281.1|10880503|AF195894 arabinogalact... 30 3.6
AV408212 29 4.7
AV411041 28 7.9
TC14994 similar to UP|O82469 (O82469) Protein phosphatase-2C , p... 28 7.9
BP039723 28 7.9
>TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%)
Length = 519
Score = 86.3 bits (212), Expect(2) = 1e-23
Identities = 38/64 (59%), Positives = 54/64 (84%)
Frame = +3
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P+FIGP+++LERVG+V+YR+ LPP LS +H VFHVS LRKY+ DPSHVI+ +DVQ+
Sbjct: 48 KLSPRFIGPFEVLERVGSVSYRLALPPDLSAVHPVFHVSMLRKYLYDPSHVIRHEDVQLD 227
Query: 1387 DNLT 1390
+L+
Sbjct: 228 VHLS 239
Score = 42.0 bits (97), Expect(2) = 1e-23
Identities = 17/34 (50%), Positives = 22/34 (64%)
Frame = +2
Query: 1409 KEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
K++ V+V+W G +GE TWE E M E YP LF
Sbjct: 296 KDVGSVKVLWRGPSGEEATWEAEDIMREKYPHLF 397
>BP085845
Length = 464
Score = 58.9 bits (141), Expect = 5e-09
Identities = 34/104 (32%), Positives = 58/104 (55%)
Frame = -2
Query: 1326 KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQV 1385
++ + K IGP++ILE V +AY + +L H V HV+ L + + D S I + V +
Sbjct: 415 RKTESKIIGPFKILEMVCLIAY*LTHSLYLLAAHIVLHVTLLWRNLYDQSQNICHEGVXL 236
Query: 1386 RDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWE 1429
++ + P+ + D KV+ +R K I V+V+W G +G +WE
Sbjct: 235 GEHWSHMEHPIVMVDMKVRCMRPKNIDDVKVIWRGLSG*EKSWE 104
>BP054462
Length = 422
Score = 48.5 bits (114), Expect = 7e-06
Identities = 21/43 (48%), Positives = 32/43 (73%)
Frame = +1
Query: 1326 KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLR 1368
+++ P++ GPY I+ ++G VAYR+ LP H S +H VFHVS L+
Sbjct: 295 EKLSPRYYGPYPIVAKIGAVAYRLELPAH-SRVHPVFHVSLLK 420
>BP046863
Length = 580
Score = 40.0 bits (92), Expect = 0.003
Identities = 20/44 (45%), Positives = 30/44 (67%)
Frame = +1
Query: 1326 KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRK 1369
+++ P+F GP+++LERV VAY + L S +H VFH+S L K
Sbjct: 451 QKLSPRFYGPFKVLERVVQVAY*LDLXSE-SRVHPVFHLSLLEK 579
>CB828458
Length = 516
Score = 31.2 bits (69), Expect = 1.2
Identities = 28/66 (42%), Positives = 38/66 (57%)
Frame = +2
Query: 209 GRARDSRVVLSRTVPLLIKGNRRWSMLGGLRRRMLQRLCVSIVVRKATKATSALRKSRSV 268
GR RDS VV+ + L+ +GNRR GLR R Q C+S R+AT+A S+S
Sbjct: 305 GRGRDSLVVIGCWI-LVPRGNRRLLTGAGLRGRFGQ--CLS--RREATRAA-----SQST 454
Query: 269 SGVARK 274
+G+ RK
Sbjct: 455 AGM*RK 472
>AV779571
Length = 538
Score = 30.8 bits (68), Expect = 1.6
Identities = 20/54 (37%), Positives = 37/54 (68%)
Frame = -2
Query: 331 LAILIILL*LLLLILVLRIVLLLSIVFLLWVLICLI*MERWLSKLQLRVR*LLL 384
+A L+I+ LLL +L+L ++LLL ++ +L + I + M L K++LR++ LL+
Sbjct: 333 IANLLIIYHLLLSLLLLLLLLLLIVIMILSLTIE*VLML*VLLKMRLRLQQLLI 172
>BP058026
Length = 389
Score = 30.4 bits (67), Expect = 2.1
Identities = 13/30 (43%), Positives = 26/30 (86%)
Frame = -2
Query: 330 VLAILIILL*LLLLILVLRIVLLLSIVFLL 359
+ ++L++LL LLLL+L+L ++LLL ++F++
Sbjct: 91 IKSLLLLLLLLLLLLLLLLLLLLLLLLFMI 2
Score = 28.5 bits (62), Expect = 7.9
Identities = 15/37 (40%), Positives = 26/37 (69%)
Frame = -2
Query: 320 RQRTRIV*SEVLAILIILL*LLLLILVLRIVLLLSIV 356
R +I +L +L++LL LLLL+L+L ++LLL ++
Sbjct: 112 RDLNKIHIKSLLLLLLLLLLLLLLLLLLLLLLLLFMI 2
>TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transcription
factor PosF21 (AtbZIP59), partial (17%)
Length = 570
Score = 30.4 bits (67), Expect = 2.1
Identities = 17/39 (43%), Positives = 27/39 (68%)
Frame = -1
Query: 339 *LLLLILVLRIVLLLSIVFLLWVLICLI*MERWLSKLQL 377
*LL+L L+L + LLL +++ W+L+ L+ + WL KL L
Sbjct: 243 *LLMLFLLLLLHLLLWLLY--WLLLLLLLLLNWLLKLML 133
>TC8475 similar to UP|GAL1_ARATH (Q9SEE5) Galactokinase (Galactose kinase)
, partial (86%)
Length = 1662
Score = 30.0 bits (66), Expect = 2.7
Identities = 16/50 (32%), Positives = 31/50 (62%), Gaps = 6/50 (12%)
Frame = +2
Query: 333 ILIILL*LLLL------ILVLRIVLLLSIVFLLWVLICLI*MERWLSKLQ 376
+L+ LL* L+L +L L +++++ ++ +W+L+CL+* W K Q
Sbjct: 779 LLVGLL**LILWRSLRRLLPLPLIIIIGLLNAIWLLLCLL*S*EWNQKKQ 928
>BP054628
Length = 508
Score = 29.6 bits (65), Expect = 3.6
Identities = 21/72 (29%), Positives = 33/72 (45%)
Frame = -3
Query: 498 CLQREKLSFLLILFLERSRSRWHLIVCLLPSYLS*RNSWRTCLRRSLLDQVFHLGERRFC 557
C Q L + ++L SR+ I+C +PS N W L ++ +F +G F
Sbjct: 506 CCQWWPLQLYVSVWLCSSRAHQEFIMCSIPSA---GNHWSMGLSNAVYQMLFAVG-LHFS 339
Query: 558 **RRKMVVCGYV 569
* K + CGY+
Sbjct: 338 S*SVKSISCGYI 303
>TC15676 similar to GB|AAG24281.1|10880503|AF195894 arabinogalactan protein
{Arabidopsis thaliana;} , partial (49%)
Length = 877
Score = 29.6 bits (65), Expect = 3.6
Identities = 15/32 (46%), Positives = 21/32 (64%)
Frame = +1
Query: 333 ILIILL*LLLLILVLRIVLLLSIVFLLWVLIC 364
+L+ L LLLL+L+L ++ S LLW LIC
Sbjct: 193 LLLFLFQLLLLLLLLLLLAFSSENLLLWSLIC 288
>AV408212
Length = 362
Score = 29.3 bits (64), Expect = 4.7
Identities = 23/56 (41%), Positives = 33/56 (58%)
Frame = -1
Query: 329 EVLAILIILL*LLLLILVLRIVLLLSIVFLLWVLICLI*MERWLSKLQLRVR*LLL 384
E+ +I +LL*L LL+L L ++LLL ++ LL +L+ M L L V LLL
Sbjct: 197 EMCSIPTLLL*LWLLLL*LLLLLLLLLLLLLLLLLLESCMPLQQDLLHLHVLLLLL 30
>AV411041
Length = 428
Score = 28.5 bits (62), Expect = 7.9
Identities = 17/35 (48%), Positives = 25/35 (70%)
Frame = -3
Query: 320 RQRTRIV*SEVLAILIILL*LLLLILVLRIVLLLS 354
R+ T ++ VL + I+LL LLLL+L+LRI+L S
Sbjct: 309 RRTTPVLHLLVLQLNILLLLLLLLMLLLRIILTRS 205
>TC14994 similar to UP|O82469 (O82469) Protein phosphatase-2C , partial
(89%)
Length = 1672
Score = 28.5 bits (62), Expect = 7.9
Identities = 15/34 (44%), Positives = 27/34 (79%)
Frame = -3
Query: 330 VLAILIILL*LLLLILVLRIVLLLSIVFLLWVLI 363
+L +L++LL LLLL+L+L+++LLL + L+ + I
Sbjct: 233 LLLLLLLLLLLLLLLLLLQLLLLLLLQ*LIEIEI 132
>BP039723
Length = 587
Score = 28.5 bits (62), Expect = 7.9
Identities = 11/39 (28%), Positives = 25/39 (63%)
Frame = +1
Query: 721 VMVLQWIRLKLKRYHNGRLLSQLLRLEVSWV*LVTIEGL 759
VMVL W+++ + + R+ +++ + V WV*++ ++ L
Sbjct: 349 VMVLNWVQMMVLDWVEMRVQNRIQEMVVDWV*VIQLQDL 465
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.365 0.164 0.591
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,358,932
Number of Sequences: 28460
Number of extensions: 399048
Number of successful extensions: 5815
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 3079
Number of HSP's successfully gapped in prelim test: 210
Number of HSP's that attempted gapping in prelim test: 2418
Number of HSP's gapped (non-prelim): 3635
length of query: 1452
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1350
effective length of database: 1,994,680
effective search space: 2692818000
effective search space used: 2692818000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146584.2


