

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146583.2 + phase: 0 /pseudo
(577 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
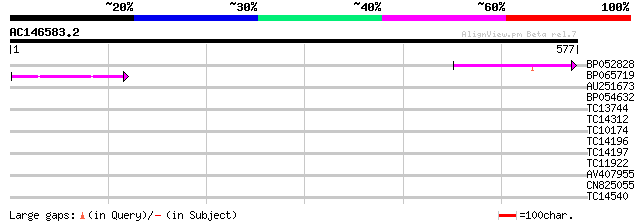
Score E
Sequences producing significant alignments: (bits) Value
BP052828 91 4e-19
BP065719 48 5e-06
AU251673 36 0.015
BP054632 31 0.63
TC13744 weakly similar to UP|Q39126 (Q39126) HYP1 protein (T4P13... 29 2.4
TC14312 similar to UP|Q8J174 (Q8J174) RfeF, partial (7%) 28 4.1
TC10174 28 4.1
TC14196 homologue to UP|Q9FNV7 (Q9FNV7) Auxin-repressed protein,... 28 5.3
TC14197 homologue to UP|Q9FNV7 (Q9FNV7) Auxin-repressed protein,... 28 5.3
TC11922 similar to PIR|T52285|T52285 serine/threonine-specific p... 27 6.9
AV407955 27 9.1
CN825055 27 9.1
TC14540 similar to UP|O49855 (O49855) Acid phosphatase , partial... 27 9.1
>BP052828
Length = 523
Score = 91.3 bits (225), Expect = 4e-19
Identities = 44/128 (34%), Positives = 76/128 (59%), Gaps = 3/128 (2%)
Frame = +2
Query: 452 PCVIGKETVDKALCDLGASVSLLPLSLFKRMGIGELKPTEMILKLADRSTIPLVGYIEDI 511
PC IG + + DLGA ++++P S++ + G L+ T +I++LA+RS G +ED+
Sbjct: 71 PCTIGDNKYENCMLDLGAGINVMPTSIYNILHPGPLQHTGLIVQLANRSNARPKGVVEDV 250
Query: 512 PVKIEGIYIPTDFVVVDIE---EDHDVPIILGRPFLATAGAIVDVQSGRIVFQASDAMIG 568
V++ + P DF ++D+E + PIILGRPF+ TA +DV +G + + D +
Sbjct: 251 LVQVNELIFPADFYILDMEGETKSSRAPIILGRPFMKTAKTKIDVDNGTMSMEFGDIIAK 430
Query: 569 FELENVMK 576
F +++ MK
Sbjct: 431 FNIDDAMK 454
>BP065719
Length = 567
Score = 47.8 bits (112), Expect = 5e-06
Identities = 33/119 (27%), Positives = 54/119 (44%)
Frame = +3
Query: 3 QFSGSPTEDPNLHISSFLRLSGTIKENQEAVRLHLFPFSLRDRASAWFHSLEVGSITSWD 62
+FSG E HI+ + +G + N E +++ FP SL A WF +L S+ +W
Sbjct: 195 KFSGDSGESTVEHIARYQIEAGDLAIN-ENLKMKYFPSSLTKNAFTWFTTLAPRSVHTWA 371
Query: 63 DMRRAFLARFFPPSKTAKLRDQITRFNQKDGESLYEAWEHFKEMLRLCPHHGLEKWLIV 121
+ R F +FF +D + +K ES+ + F+ + C H E L+V
Sbjct: 372 QLERIFHEQFFRGECKVSXKD-LASVKRKPAESIDDYLNRFRMLKSRCFTHVSEHELVV 545
>AU251673
Length = 413
Score = 36.2 bits (82), Expect = 0.015
Identities = 33/115 (28%), Positives = 46/115 (39%), Gaps = 5/115 (4%)
Frame = +2
Query: 35 LHLFPFSLRDRASAWFHSLE----VGSIT-SWDDMRRAFLARFFPPSKTAKLRDQITRFN 89
+ L F L A W++ L VGS +W D F+ RF P S R
Sbjct: 23 VELASFQLEGVARDWYNVLTRAKPVGSPPWTWADFSAEFMNRFLPQSVRDGFVRDFERLE 202
Query: 90 QKDGESLYEAWEHFKEMLRLCPHHGLEKWLIVHTFYNGLSDTTKMSVDAAAGGAL 144
Q +G ++ E HF + R P+ LE+ V F GL + SV + L
Sbjct: 203 QAEGMTVSEYSAHFTHLSRYVPYPLLEEER-VKRFVRGLKEYLFKSVVGSKSSTL 364
>BP054632
Length = 514
Score = 30.8 bits (68), Expect = 0.63
Identities = 23/87 (26%), Positives = 42/87 (47%), Gaps = 4/87 (4%)
Frame = +3
Query: 212 NQNFYKNPQGSYGQTAPPGYTNNQRVAQKSSLEILLENCMMNQNKQLQELKNQTGSLNDS 271
NQ+ YK P +Y Q + Y++ LE+ + +++Q L+++ LN+
Sbjct: 261 NQDAYKKP--NYVQISVESYSHLTG----------LEDQVKTYEEKVQTLEDEINELNEK 404
Query: 272 LS----KLNTKVDSIATHTKMLETQIS 294
LS ++NTK + H K+ E +S
Sbjct: 405 LSAANSEINTKEGLVKQHAKVAEEAVS 485
>TC13744 weakly similar to UP|Q39126 (Q39126) HYP1 protein (T4P13.21
protein), partial (26%)
Length = 744
Score = 28.9 bits (63), Expect = 2.4
Identities = 16/39 (41%), Positives = 21/39 (53%)
Frame = +3
Query: 37 LFPFSLRDRASAWFHSLEVGSITSWDDMRRAFLARFFPP 75
LF L + ASAW SL GS+ S+ + + L FPP
Sbjct: 627 LFVVKLSE*ASAWAFSLGNGSVNSFWKLLQFLLIDVFPP 743
>TC14312 similar to UP|Q8J174 (Q8J174) RfeF, partial (7%)
Length = 622
Score = 28.1 bits (61), Expect = 4.1
Identities = 13/19 (68%), Positives = 14/19 (73%)
Frame = +1
Query: 216 YKNPQGSYGQTAPPGYTNN 234
Y PQGSYGQ APP Y N+
Sbjct: 199 YGAPQGSYGQ-APPSYGNS 252
>TC10174
Length = 1063
Score = 28.1 bits (61), Expect = 4.1
Identities = 15/43 (34%), Positives = 24/43 (54%), Gaps = 4/43 (9%)
Frame = +1
Query: 445 DPGSFVIPCVIGKETVDKALCDLGASVSLLPLS----LFKRMG 483
D SF++ CV+ ++ ++K L G +L PL LFK +G
Sbjct: 397 DLASFIVDCVLSEDKINKVLPIGGPGKALTPLEQGEMLFKLVG 525
>TC14196 homologue to UP|Q9FNV7 (Q9FNV7) Auxin-repressed protein, partial
(95%)
Length = 1040
Score = 27.7 bits (60), Expect = 5.3
Identities = 15/32 (46%), Positives = 19/32 (58%)
Frame = -2
Query: 445 DPGSFVIPCVIGKETVDKALCDLGASVSLLPL 476
DPG +P V+G +DK LC+L S S L L
Sbjct: 205 DPGVVGVPGVVGVGGMDKFLCNLLPSPSSLML 110
>TC14197 homologue to UP|Q9FNV7 (Q9FNV7) Auxin-repressed protein, partial
(95%)
Length = 976
Score = 27.7 bits (60), Expect = 5.3
Identities = 15/32 (46%), Positives = 19/32 (58%)
Frame = -2
Query: 445 DPGSFVIPCVIGKETVDKALCDLGASVSLLPL 476
DPG +P V+G +DK LC+L S S L L
Sbjct: 348 DPGVVGVPGVVGVGGMDKFLCNLLPSPSSLML 253
>TC11922 similar to PIR|T52285|T52285 serine/threonine-specific protein
kinase APK2a [imported] - Arabidopsis
thaliana {Arabidopsis thaliana;} , partial (22%)
Length = 550
Score = 27.3 bits (59), Expect = 6.9
Identities = 16/60 (26%), Positives = 32/60 (52%), Gaps = 2/60 (3%)
Frame = +2
Query: 310 FPGQTETNPKAHVNAISLGGNKLEDTITK--AKSVKGESVKLLGEKDAIKTLLDKNKALN 367
F G + P AHV++ S GNK + K + S++ ++ + E + K++ +K K+ +
Sbjct: 101 FMGNSCAKPVAHVSSSSFAGNKRPQSRPKQHSNSLEQKAAMQISEANVEKSISNKLKSFS 280
>AV407955
Length = 402
Score = 26.9 bits (58), Expect = 9.1
Identities = 20/68 (29%), Positives = 31/68 (45%)
Frame = -2
Query: 6 GSPTEDPNLHISSFLRLSGTIKENQEAVRLHLFPFSLRDRASAWFHSLEVGSITSWDDMR 65
GS E P + + G ++ ++ LFP S+RDR+ + VG S ++R
Sbjct: 221 GSSKEPPPAPLEGLRPVFGNLRMSR*-----LFP*SVRDRSGDFSWDA*VGETFSEGELR 57
Query: 66 RAFLARFF 73
FLA F
Sbjct: 56 VEFLAELF 33
>CN825055
Length = 593
Score = 26.9 bits (58), Expect = 9.1
Identities = 17/48 (35%), Positives = 20/48 (41%)
Frame = +2
Query: 187 VSEYNHLAAKVEQLNYAQYKQGMRPNQNFYKNPQGSYGQTAPPGYTNN 234
V +YNH AK Q +G+ N Y N S GQT NN
Sbjct: 446 VEDYNHFLAKAIAGGTGQIVKGIFMCSNAYTNQVQSGGQTILTDNKNN 589
>TC14540 similar to UP|O49855 (O49855) Acid phosphatase , partial (88%)
Length = 1144
Score = 26.9 bits (58), Expect = 9.1
Identities = 16/54 (29%), Positives = 28/54 (51%)
Frame = +1
Query: 400 LSRMPLYTKFLKEIFSKKKAMDHKETIALTRESSAIVKKQPPKLRDPGSFVIPC 453
L + L+T +E +KK + ET+ E+S ++ +PP+ S +IPC
Sbjct: 661 LVKQQLHTNQRRERNWRKKDTESLETL----ETSGVIS*EPPQATGLSSCLIPC 810
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.134 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,765,988
Number of Sequences: 28460
Number of extensions: 111041
Number of successful extensions: 507
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 505
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 506
length of query: 577
length of database: 4,897,600
effective HSP length: 95
effective length of query: 482
effective length of database: 2,193,900
effective search space: 1057459800
effective search space used: 1057459800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146583.2


