

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
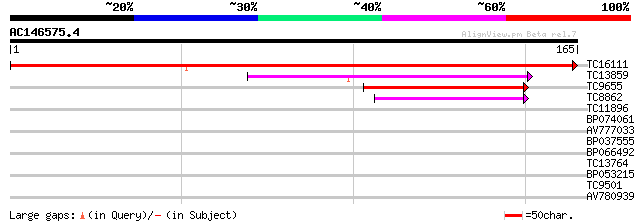
Query= AC146575.4 - phase: 0
(165 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC16111 weakly similar to UP|Q925F6 (Q925F6) Inner mitochondrial... 259 2e-70
TC13859 similar to UP|Q9ZUE6 (Q9ZUE6) F5O8.3 protein, partial (23%) 68 9e-13
TC9655 similar to GB|CAA71502.1|2769566|ATCTPP chloroplast thyla... 45 5e-06
TC8862 similar to UP|Q9LV44 (Q9LV44) Similarity to signal peptid... 44 1e-05
TC11896 29 0.36
BP074061 28 0.61
AV777033 28 1.0
BP037555 27 1.8
BP066492 26 4.0
TC13764 similar to UP|Q9C9Y6 (Q9C9Y6) Thioredoxin-like protein, ... 26 4.0
BP053215 25 5.2
TC9501 weakly similar to UP|Q9MBB6 (Q9MBB6) Pectin methylesteras... 25 6.8
AV780939 25 6.8
>TC16111 weakly similar to UP|Q925F6 (Q925F6) Inner mitochondrial membrane
peptidase 2, partial (25%)
Length = 1171
Score = 259 bits (662), Expect = 2e-70
Identities = 123/169 (72%), Positives = 135/169 (79%), Gaps = 4/169 (2%)
Frame = +3
Query: 1 MGTRNLVWNVTKKLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNSF----TDDYVF 56
MG +WN TKK T GLI+ TVSDRY TVVPVRG SMSPTFNPKT+S +DDYVF
Sbjct: 147 MGPSGFLWNFTKKFVTFGLISVTVSDRYMTVVPVRGGSMSPTFNPKTHSLMGGVSDDYVF 326
Query: 57 VEKLCLDKFKFSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNRHNQDVLKVPEGHCWVE 116
VEK CL K+KFSHGD+V+F SP N KETHIKRIIALPGEW HN DVLK+PEGHCWVE
Sbjct: 327 VEKFCLQKYKFSHGDVVVFRSPLNHKETHIKRIIALPGEWIGAHHNYDVLKIPEGHCWVE 506
Query: 117 GDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIGAVKNTTPERLPSS 165
GDNAASS DSKS+GP+PL L+RGRVTHVVWPP RIGAVK+T P L S
Sbjct: 507 GDNAASSLDSKSFGPIPLALIRGRVTHVVWPPPRIGAVKSTPPAGLSPS 653
Score = 26.6 bits (57), Expect = 2.3
Identities = 10/16 (62%), Positives = 11/16 (68%)
Frame = +1
Query: 145 VWPPQRIGAVKNTTPE 160
VWPP RIG + TT E
Sbjct: 676 VWPPPRIGGERRTTQE 723
>TC13859 similar to UP|Q9ZUE6 (Q9ZUE6) F5O8.3 protein, partial (23%)
Length = 499
Score = 67.8 bits (164), Expect = 9e-13
Identities = 34/89 (38%), Positives = 52/89 (58%), Gaps = 6/89 (6%)
Frame = +2
Query: 70 GDIVIFSSPSNFKETHIKRIIALPGEWF------VNRHNQDVLKVPEGHCWVEGDNAASS 123
GDIV SP+N KR++ + G+ ++ + + VP+GH WVEGDN +S
Sbjct: 5 GDIVCLCSPANPTTFITKRVVGMEGDTVTYLPSPMDSDQLETIVVPKGHVWVEGDNIFNS 184
Query: 124 TDSKSYGPVPLGLVRGRVTHVVWPPQRIG 152
DS+++GPVP GL++GR+ V P + G
Sbjct: 185 KDSRNFGPVPYGLIQGRIFWRVLPLKDFG 271
>TC9655 similar to GB|CAA71502.1|2769566|ATCTPP chloroplast thylakoidal
processing peptidase {Arabidopsis thaliana;} , partial
(19%)
Length = 694
Score = 45.4 bits (106), Expect = 5e-06
Identities = 20/48 (41%), Positives = 30/48 (61%)
Frame = +3
Query: 104 DVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQRI 151
D + VP+GH +V GD+ S DS ++GP+P+ + GR WPP R+
Sbjct: 42 DRMVVPKGHVYVLGDHRNRSFDSHNWGPLPVENLLGRSLFRYWPPSRV 185
>TC8862 similar to UP|Q9LV44 (Q9LV44) Similarity to signal peptidase,
partial (32%)
Length = 605
Score = 43.9 bits (102), Expect = 1e-05
Identities = 20/45 (44%), Positives = 27/45 (59%)
Frame = +2
Query: 107 KVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQRI 151
+VPE + +V GDN +S DS +GP+P + GR WPP RI
Sbjct: 131 RVPENYVFVMGDNRNNSYDSHVWGPLPAKNIIGRSVFRYWPPNRI 265
>TC11896
Length = 648
Score = 29.3 bits (64), Expect = 0.36
Identities = 23/71 (32%), Positives = 32/71 (44%)
Frame = +2
Query: 41 PTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNR 100
P N K SF FVEKLCL +K ++ +P+N K T+ ++L V
Sbjct: 248 PVTNSKLISFRICGAFVEKLCLVAYKIK*FCHILTPNPTNTKLTNDNV*VSLEEMSCVFT 427
Query: 101 HNQDVLKVPEG 111
N DV + G
Sbjct: 428 CNIDVNNIKTG 460
>BP074061
Length = 379
Score = 28.5 bits (62), Expect = 0.61
Identities = 12/27 (44%), Positives = 19/27 (69%)
Frame = -1
Query: 54 YVFVEKLCLDKFKFSHGDIVIFSSPSN 80
Y+++ CLD+F FS I+IFS+ S+
Sbjct: 217 YMYILTTCLDQFLFSCKSILIFSTESS 137
>AV777033
Length = 592
Score = 27.7 bits (60), Expect = 1.0
Identities = 16/56 (28%), Positives = 24/56 (42%)
Frame = +2
Query: 75 FSSPSNFKETHIKRIIALPGEWFVNRHNQDVLKVPEGHCWVEGDNAASSTDSKSYG 130
F+S SNF + +A+PG V++ D+L P W D T+ G
Sbjct: 182 FASASNFAARSRRFSVAIPGPCIVHKRGADILHDP----WFNKDTGFPLTERDRLG 337
>BP037555
Length = 549
Score = 26.9 bits (58), Expect = 1.8
Identities = 12/27 (44%), Positives = 18/27 (66%)
Frame = -2
Query: 114 WVEGDNAASSTDSKSYGPVPLGLVRGR 140
+V GDN +S DS +GP+P+ + GR
Sbjct: 545 YVLGDNRNNSYDSHVWGPLPVKNIIGR 465
>BP066492
Length = 469
Score = 25.8 bits (55), Expect = 4.0
Identities = 14/29 (48%), Positives = 18/29 (61%), Gaps = 4/29 (13%)
Frame = +2
Query: 130 GPVP---LGL-VRGRVTHVVWPPQRIGAV 154
G VP LGL +R R+TH++W P I V
Sbjct: 341 GTVPGLRLGLRLRFRLTHIIWAPSSISMV 427
>TC13764 similar to UP|Q9C9Y6 (Q9C9Y6) Thioredoxin-like protein, partial
(73%)
Length = 585
Score = 25.8 bits (55), Expect = 4.0
Identities = 12/46 (26%), Positives = 24/46 (52%)
Frame = -2
Query: 67 FSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNRHNQDVLKVPEGH 112
F+ D + ++P +E H++ +I E + RH++ +K GH
Sbjct: 509 FAKHDGLTTATPL*IEEIHVRCLICTSKEKIMQRHSRQNMKNNHGH 372
>BP053215
Length = 275
Score = 25.4 bits (54), Expect = 5.2
Identities = 10/20 (50%), Positives = 13/20 (65%)
Frame = +3
Query: 1 MGTRNLVWNVTKKLATIGLI 20
+G RN+ W V+ LATI I
Sbjct: 207 LGLRNMAWRVSNHLATINSI 266
>TC9501 weakly similar to UP|Q9MBB6 (Q9MBB6) Pectin methylesterase ,
partial (29%)
Length = 934
Score = 25.0 bits (53), Expect = 6.8
Identities = 14/34 (41%), Positives = 21/34 (61%), Gaps = 4/34 (11%)
Frame = -2
Query: 117 GDNAASSTDSKSYGPVPLGL----VRGRVTHVVW 146
G+NAAS T S+S+ +P+G+ V VT + W
Sbjct: 450 GNNAASGTISQSWITIPVGMFFVGVPSAVTVLNW 349
>AV780939
Length = 548
Score = 25.0 bits (53), Expect = 6.8
Identities = 10/23 (43%), Positives = 12/23 (51%)
Frame = +1
Query: 92 LPGEWFVNRHNQDVLKVPEGHCW 114
LP W ++RH Q V V CW
Sbjct: 82 LPRRWLIHRHRQQVAPV----CW 138
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.136 0.422
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,051,042
Number of Sequences: 28460
Number of extensions: 42026
Number of successful extensions: 182
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 180
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 180
length of query: 165
length of database: 4,897,600
effective HSP length: 83
effective length of query: 82
effective length of database: 2,535,420
effective search space: 207904440
effective search space used: 207904440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146575.4


