

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146574.11 + phase: 0 /pseudo
(738 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
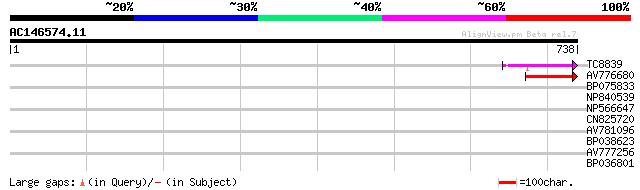
Score E
Sequences producing significant alignments: (bits) Value
TC8839 weakly similar to PIR|AD1564|AD1564 B. subtilis YktB prot... 110 1e-24
AV776680 80 9e-16
BP075833 37 0.009
NP840539 hypothetical protein [Lotus corniculatus var. japonicus] 32 0.36
NP566647 unnamed protein product [Lotus japonicus] 32 0.36
CN825720 31 0.81
AV781096 31 0.81
BP038623 28 5.3
AV777256 28 5.3
BP036801 28 6.9
>TC8839 weakly similar to PIR|AD1564|AD1564 B. subtilis YktB protein
homolog lin1053 [imported] - Listeria
innocua (strain Clip11262) {Listeria innocua;}, partial
(9%)
Length = 751
Score = 110 bits (274), Expect = 1e-24
Identities = 62/131 (47%), Positives = 73/131 (55%), Gaps = 34/131 (25%)
Frame = +2
Query: 642 MKIHRWLAGEEDDFELKLESVIEGCNGTWLR----------------------------- 672
MKI R ++G ED LK++S+IE CN TW+R
Sbjct: 2 MKIQR-ISGGEDVLALKIDSIIEVCNDTWVRNQGSMCPLIEDQCFPPYAKRKRLTDAVLM 178
Query: 673 -----KLDGICHENNWILPTYSVSLSDGEFHATVRVKGVDFEYSCEGNTCPFPREARDSA 727
+LDG+CH NNWILPTYS+S S G F A + VKG DFE SC GN C PREAR+SA
Sbjct: 179 QSPCQELDGVCHANNWILPTYSLSQSAGGFQANLTVKGPDFESSCGGNICCHPREARESA 358
Query: 728 AAQMLTKFRSM 738
AAQML RSM
Sbjct: 359 AAQMLVNLRSM 391
>AV776680
Length = 501
Score = 80.5 bits (197), Expect = 9e-16
Identities = 38/67 (56%), Positives = 49/67 (72%)
Frame = -3
Query: 672 RKLDGICHENNWILPTYSVSLSDGEFHATVRVKGVDFEYSCEGNTCPFPREARDSAAAQM 731
++LDG+C ENNW+LPTY + S G F A V VKGV+F+ S G+ C +EAR+SAAA+M
Sbjct: 457 QELDGVCRENNWLLPTYHLFQSVGGFQANVTVKGVEFQCSYAGDLCSDSQEARESAAAKM 278
Query: 732 LTKFRSM 738
LT RSM
Sbjct: 277 LTNLRSM 257
>BP075833
Length = 408
Score = 37.4 bits (85), Expect = 0.009
Identities = 18/30 (60%), Positives = 21/30 (70%)
Frame = -2
Query: 709 EYSCEGNTCPFPREARDSAAAQMLTKFRSM 738
E SC G+ C PR AR+SAAAQ+L RSM
Sbjct: 407 ESSCGGDICCHPRAARESAAAQILVYLRSM 318
>NP840539 hypothetical protein [Lotus corniculatus var. japonicus]
Length = 3654
Score = 32.0 bits (71), Expect = 0.36
Identities = 38/180 (21%), Positives = 62/180 (34%), Gaps = 40/180 (22%)
Frame = +1
Query: 453 SHVSSKGLY---IDLDNKPGSDTKVALNQKEKNGCGITKRCDS----------------- 492
S VS G Y I+LD+ DT + +++ + C + DS
Sbjct: 2560 SEVSYDGEYMENIELDDSDSQDTDMNWEEEQFSACDYKQEIDSSVIMQKTGSKFEASSES 2739
Query: 493 ---VKEDW----------DMDVDKSL--VLPSKNKECQKHTANTLHISEDQKIENPSVQH 537
+ E W D+ V+ +L V KN + T T + ED + H
Sbjct: 2740 LCGISEMWLDDILSNHYADILVEVALQAVKEEKNTHFEAQTHGTKSVLEDIEFNTQETDH 2919
Query: 538 HSNECTRPSKAEKAVSKRKHIT---EGGIKDQSAFDK--ICAGTTFENDSVEKCILNANS 592
SN + +H T + + + D CA + D++E C + NS
Sbjct: 2920 LSNAASHEHDQSSTEEVFEHFTNTRDNNRESEKHMDNEVSCASKVLDEDAIENCEGHTNS 3099
>NP566647 unnamed protein product [Lotus japonicus]
Length = 2205
Score = 32.0 bits (71), Expect = 0.36
Identities = 38/180 (21%), Positives = 62/180 (34%), Gaps = 40/180 (22%)
Frame = +1
Query: 453 SHVSSKGLY---IDLDNKPGSDTKVALNQKEKNGCGITKRCDS----------------- 492
S VS G Y I+LD+ DT + +++ + C + DS
Sbjct: 1108 SEVSYDGEYMENIELDDSDSQDTDMNWEEEQFSACDYKQEIDSSVIMQKTGSKFEASSES 1287
Query: 493 ---VKEDW----------DMDVDKSL--VLPSKNKECQKHTANTLHISEDQKIENPSVQH 537
+ E W D+ V+ +L V KN + T T + ED + H
Sbjct: 1288 LCGISEMWLDDILSNHYADILVEVALQAVKEEKNTHFEAQTHGTKSVLEDIEFNTQETDH 1467
Query: 538 HSNECTRPSKAEKAVSKRKHIT---EGGIKDQSAFDK--ICAGTTFENDSVEKCILNANS 592
SN + +H T + + + D CA + D++E C + NS
Sbjct: 1468 LSNAASHEHDQSSTEEVFEHFTNTRDNNRESEKHMDNEVSCASKVLDEDAIENCEGHTNS 1647
>CN825720
Length = 699
Score = 30.8 bits (68), Expect = 0.81
Identities = 19/100 (19%), Positives = 41/100 (41%)
Frame = +1
Query: 215 DGVWSVIEKDVGSSDQISEDMNELKHTYQKRRVIQNPTKNGLNVDDDEFLQVGYSAVKEA 274
DG+ +++ + D +S E ++ R +QN L DD + V+
Sbjct: 358 DGLKTLMAESSEDKDMLSMATEEFGQAKEEERRLQNMLLKSLLPKDDADERDSILEVRAG 537
Query: 275 TGVNSNDIMLLESYTVYSQRKEKTASRFYIMKCSQSTADG 314
+G + ++ + +Y + K +F ++ +QS G
Sbjct: 538 SGGEEASLFAMDIFKMYEKYAHKNGWKFEVVDIAQSDMKG 657
>AV781096
Length = 497
Score = 30.8 bits (68), Expect = 0.81
Identities = 17/41 (41%), Positives = 24/41 (58%)
Frame = +2
Query: 395 NIFHQIISHFHFMFSLEPLSVR*G*LFV*NFLCRETFSNSL 435
++F+QII H +F L S++*G F NF C+ T SL
Sbjct: 83 SLFNQIILFSHPIFPLNTASIQ*GQSFFYNFTCQGTTLTSL 205
>BP038623
Length = 538
Score = 28.1 bits (61), Expect = 5.3
Identities = 13/70 (18%), Positives = 30/70 (42%)
Frame = +2
Query: 515 QKHTANTLHISEDQKIENPSVQHHSNECTRPSKAEKAVSKRKHITEGGIKDQSAFDKICA 574
++ L + + + + SNE T AEK ++ IT+ G + F+ +
Sbjct: 167 EQRVITDLSLGSEDPLHQEATSRDSNEATDHEPAEKLETQETEITQEGNAKEDEFENVII 346
Query: 575 GTTFENDSVE 584
++ + D ++
Sbjct: 347 KSSRDTDMMK 376
>AV777256
Length = 390
Score = 28.1 bits (61), Expect = 5.3
Identities = 13/32 (40%), Positives = 15/32 (46%), Gaps = 2/32 (6%)
Frame = -2
Query: 669 TWLRKLDGICHENNWI--LPTYSVSLSDGEFH 698
TWL D +C E W+ P S S S FH
Sbjct: 101 TWLHMFDSLCEEVFWV*ATPLDSFSFSSSVFH 6
>BP036801
Length = 540
Score = 27.7 bits (60), Expect = 6.9
Identities = 11/34 (32%), Positives = 19/34 (55%)
Frame = +1
Query: 206 CLLRFCSTTDGVWSVIEKDVGSSDQISEDMNELK 239
C+ CS+T G W +++ +G S + MN+ K
Sbjct: 61 CITTTCSSTGGHWVQLQRSIGISAMHMQVMNDNK 162
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,656,492
Number of Sequences: 28460
Number of extensions: 174846
Number of successful extensions: 1064
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1055
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1064
length of query: 738
length of database: 4,897,600
effective HSP length: 97
effective length of query: 641
effective length of database: 2,136,980
effective search space: 1369804180
effective search space used: 1369804180
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146574.11


