

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146568.12 + phase: 0 /pseudo
(1737 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
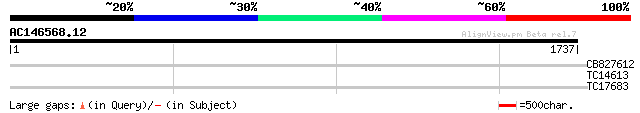
Sequences producing significant alignments: (bits) Value
CB827612 41 0.001
TC14613 homologue to UP|PAL2_PEA (Q04593) Phenylalanine ammonia-... 30 3.3
TC17683 weakly similar to UP|P93816 (P93816) F19P19.10, partial ... 28 9.5
>CB827612
Length = 277
Score = 41.2 bits (95), Expect = 0.001
Identities = 33/91 (36%), Positives = 49/91 (53%)
Frame = +2
Query: 1593 VIFQ*S*STKLIGPSKLLILIKL*LVRKDS*SSMS*KK*DLVRMRTPSFTKRELRSIMTR 1652
V F * *+T LIG K++I V K++ + S K *D M+ F KR+L M +
Sbjct: 5 VTFL*R*NTVLIGQ*KVVIXRCNKQVLKENSNYSSLKN*D*GLMKALGFIKRKLSIXMIK 184
Query: 1653 VW*GESFMLDNLFCCLIPG*NYFPGN*NPSG 1683
+ G + + N F CLIP + + GN +P+G
Sbjct: 185 *FLGRNSLWANKFYCLIPALSLWRGNFDPNG 277
>TC14613 homologue to UP|PAL2_PEA (Q04593) Phenylalanine ammonia-lyase 2 ,
partial (47%)
Length = 1048
Score = 30.0 bits (66), Expect = 3.3
Identities = 11/26 (42%), Positives = 18/26 (68%)
Frame = +1
Query: 1015 LRYLWTISLYLVSRLTNACFISMLSL 1040
L +LWT +L+ +L N+C ++ LSL
Sbjct: 676 LEFLWTTHAWLLQQLANSCLLNSLSL 753
>TC17683 weakly similar to UP|P93816 (P93816) F19P19.10, partial (5%)
Length = 561
Score = 28.5 bits (62), Expect = 9.5
Identities = 13/24 (54%), Positives = 17/24 (70%), Gaps = 2/24 (8%)
Frame = -3
Query: 289 IPCFLIHNN--NNNQESLQVLRKQ 310
IPCFLIHN +N+ S+ + RKQ
Sbjct: 553 IPCFLIHNRGASNSVSSMYIRRKQ 482
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.369 0.165 0.601
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,462,759
Number of Sequences: 28460
Number of extensions: 405158
Number of successful extensions: 4949
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 2684
Number of HSP's successfully gapped in prelim test: 143
Number of HSP's that attempted gapping in prelim test: 2166
Number of HSP's gapped (non-prelim): 3020
length of query: 1737
length of database: 4,897,600
effective HSP length: 103
effective length of query: 1634
effective length of database: 1,966,220
effective search space: 3212803480
effective search space used: 3212803480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146568.12


