

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146567.9 + phase: 0
(80 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
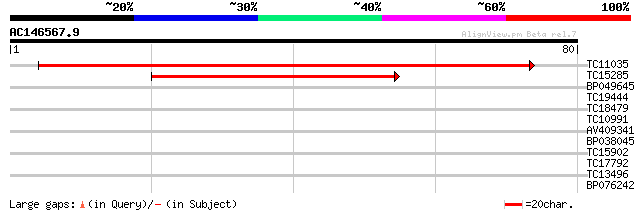
Score E
Sequences producing significant alignments: (bits) Value
TC11035 homologue to UP|Q9C6K5 (Q9C6K5) Sm-like protein, partial... 135 2e-33
TC15285 homologue to PIR|T10586|T10586 small nuclear ribonucleop... 38 3e-04
BP049645 35 0.002
TC19444 weakly similar to UP|LSM7_HUMAN (Q9UK45) U6 snRNA-associ... 34 0.006
TC18479 similar to UP|O80960 (O80960) Nodulin-like protein (At2g... 29 0.18
TC10991 similar to UP|Q8VYA0 (Q8VYA0) Aspartate--tRNA ligase-lik... 26 1.2
AV409341 26 1.2
BP038045 26 1.5
TC15902 similar to UP|Q948P0 (Q948P0) Aspartic proteinase alpha,... 24 5.8
TC17792 24 5.8
TC13496 23 7.5
BP076242 23 7.5
>TC11035 homologue to UP|Q9C6K5 (Q9C6K5) Sm-like protein, partial (96%)
Length = 576
Score = 135 bits (339), Expect = 2e-33
Identities = 68/70 (97%), Positives = 69/70 (98%)
Frame = +2
Query: 5 EEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDD 64
EEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDD
Sbjct: 23 EEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDD 202
Query: 65 ETYEEIVRVS 74
ETYEEIVR +
Sbjct: 203ETYEEIVRTT 232
>TC15285 homologue to PIR|T10586|T10586 small nuclear
ribonucleoprotein-associated protein homolog
F9F13.90 - Arabidopsis thaliana {Arabidopsis
thaliana;}, partial (61%)
Length = 587
Score = 38.1 bits (87), Expect = 3e-04
Identities = 15/35 (42%), Positives = 26/35 (73%)
Frame = +3
Query: 21 LDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEE 55
++ R+ V ++ R+L GK A+D+H+N++LGD EE
Sbjct: 114 INYRMRVTIQDGRQLVGKFMAFDRHMNLVLGDCEE 218
>BP049645
Length = 415
Score = 35.0 bits (79), Expect = 0.002
Identities = 13/37 (35%), Positives = 27/37 (72%)
Frame = -3
Query: 24 RIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTV 60
++ + R++++L G++ A+D+H NM+L +V E+ T V
Sbjct: 380 QVLINCRNNKKLLGRVRAFDRHCNMVLENVREMWTEV 270
>TC19444 weakly similar to UP|LSM7_HUMAN (Q9UK45) U6 snRNA-associated
Sm-like protein LSm7, partial (88%)
Length = 565
Score = 33.9 bits (76), Expect = 0.006
Identities = 17/44 (38%), Positives = 27/44 (60%)
Frame = +3
Query: 14 LDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIV 57
LDL + +D+ + VKL R++ G L YDQ LN++L + E +
Sbjct: 135 LDLAKF-VDKGVQVKLTGGRQVTGTLKGYDQLLNLVLDEAVEFL 263
>TC18479 similar to UP|O80960 (O80960) Nodulin-like protein
(At2g39210/T16B24.15), partial (27%)
Length = 524
Score = 28.9 bits (63), Expect = 0.18
Identities = 19/54 (35%), Positives = 27/54 (49%), Gaps = 1/54 (1%)
Frame = -1
Query: 28 KLRSDRELRGKLHAYDQH-LNMILGDVEEIVTTVEIDDETYEEIVRVSHNPNHS 80
KLR D R KL YDQH + L E V + + ++I R H+P++S
Sbjct: 392 KLRKDHWAR*KLSEYDQHRIRAELESALEAVENCTLSEHVLKQIQR-KHDPHNS 234
>TC10991 similar to UP|Q8VYA0 (Q8VYA0) Aspartate--tRNA ligase-like protein,
partial (42%)
Length = 680
Score = 26.2 bits (56), Expect = 1.2
Identities = 15/36 (41%), Positives = 19/36 (52%)
Frame = +3
Query: 14 LDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMI 49
LDL+R+ L L D +G LHA+ HLN I
Sbjct: 456 LDLVRVEL-------LFLDWTTKGNLHAWPNHLNFI 542
>AV409341
Length = 321
Score = 26.2 bits (56), Expect = 1.2
Identities = 12/22 (54%), Positives = 14/22 (63%)
Frame = +2
Query: 22 DERIYVKLRSDRELRGKLHAYD 43
+ R YVK RSD +LR K YD
Sbjct: 224 NHRRYVKSRSDTQLRSKAAEYD 289
>BP038045
Length = 556
Score = 25.8 bits (55), Expect = 1.5
Identities = 12/38 (31%), Positives = 22/38 (57%)
Frame = +1
Query: 35 LRGKLHAYDQHLNMILGDVEEIVTTVEIDDETYEEIVR 72
+RG LH+ DQ+LN+ ++ T +D + Y +V+
Sbjct: 103 IRGTLHSVDQYLNI------KLENTRVVDQDKYPHMVK 198
>TC15902 similar to UP|Q948P0 (Q948P0) Aspartic proteinase alpha, partial
(25%)
Length = 517
Score = 23.9 bits (50), Expect = 5.8
Identities = 19/57 (33%), Positives = 27/57 (47%)
Frame = +3
Query: 11 KEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETY 67
K PLDL ++ +++ +LRS R + G L LN+ D E IV D Y
Sbjct: 237 KRPLDLQSINAAKKVSGELRSVRPMLGAL-----DLNIGKPDGEAIVPLKNYLDAQY 392
>TC17792
Length = 665
Score = 23.9 bits (50), Expect = 5.8
Identities = 13/37 (35%), Positives = 23/37 (62%), Gaps = 1/37 (2%)
Frame = +1
Query: 31 SDRELRGKLHAYDQHLNMILGD-VEEIVTTVEIDDET 66
S+ + G + A +HLN+ILG + I T+++ D+T
Sbjct: 7 SEVKKMGGVRAEKEHLNLILGSHLTTIHDTLQVLDQT 117
>TC13496
Length = 576
Score = 23.5 bits (49), Expect = 7.5
Identities = 10/23 (43%), Positives = 13/23 (56%)
Frame = +2
Query: 1 MAGTEEESAVKEPLDLIRLSLDE 23
M G S V P DL+R+ +DE
Sbjct: 122 MTGQGHASNVMMPFDLLRIKIDE 190
>BP076242
Length = 470
Score = 23.5 bits (49), Expect = 7.5
Identities = 8/20 (40%), Positives = 14/20 (70%)
Frame = +2
Query: 31 SDRELRGKLHAYDQHLNMIL 50
S++ + KLH ++ HLN +L
Sbjct: 179 SNQRIHSKLHLWNHHLNPLL 238
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.134 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,048,184
Number of Sequences: 28460
Number of extensions: 9254
Number of successful extensions: 33
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 33
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 33
length of query: 80
length of database: 4,897,600
effective HSP length: 56
effective length of query: 24
effective length of database: 3,303,840
effective search space: 79292160
effective search space used: 79292160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 48 (23.1 bits)
Medicago: description of AC146567.9


