

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
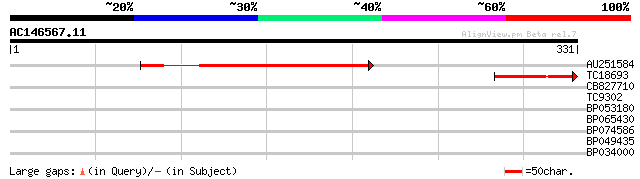
Query= AC146567.11 - phase: 0
(331 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AU251584 208 9e-55
TC18693 weakly similar to UP|Q84TE3 (Q84TE3) At2g03510, partial ... 76 9e-15
CB827710 36 0.008
TC9302 similar to UP|BGLT_TRIRP (P26205) Cyanogenic beta-glucosi... 28 2.1
BP053180 27 3.6
BP065430 27 3.6
BP074586 26 8.1
BP049435 26 8.1
BP034000 26 8.1
>AU251584
Length = 350
Score = 208 bits (530), Expect = 9e-55
Identities = 109/136 (80%), Positives = 110/136 (80%)
Frame = +1
Query: 77 GTKGGVMIVFGKIEVVNRLHKESVYETLLNYGVQYDKTWIYDKIHHEINQFCSSHSLQQV 136
GTKGGVMI F KIE YDKTWIYDKIHHEINQFCSSHSLQQV
Sbjct: 1 GTKGGVMINFEKIE--------------------YDKTWIYDKIHHEINQFCSSHSLQQV 120
Query: 137 YIDVFDQIDEKMKDALQVDCTRYAPGIEIIGVRVTKPNIPESIRHNFEQMEEERTKVLIA 196
YIDVFDQIDEKMKDALQ DCTRYAPGIEIIGVRVTKP IP SIR NFEQMEEERTKVLIA
Sbjct: 121 YIDVFDQIDEKMKDALQGDCTRYAPGIEIIGVRVTKPTIPSSIRRNFEQMEEERTKVLIA 300
Query: 197 IEKQKVSEKEAETMKK 212
IEKQ+VSEKEAETMKK
Sbjct: 301 IEKQRVSEKEAETMKK 348
>TC18693 weakly similar to UP|Q84TE3 (Q84TE3) At2g03510, partial (10%)
Length = 427
Score = 75.9 bits (185), Expect = 9e-15
Identities = 38/48 (79%), Positives = 42/48 (87%)
Frame = +3
Query: 284 KFIESIANNTKIFFGDKIPNMILDQRLLGNFLVEEVPRGAATKTKADI 331
KFIES+ NNTKIFFG+KIP MILDQRLLGN L++E RGAATK KADI
Sbjct: 3 KFIESVTNNTKIFFGEKIPGMILDQRLLGN-LLQEASRGAATKAKADI 143
>CB827710
Length = 560
Score = 36.2 bits (82), Expect = 0.008
Identities = 23/79 (29%), Positives = 45/79 (56%), Gaps = 3/79 (3%)
Frame = +1
Query: 179 IRHNFEQMEEERTKVLIAIEK-QKVSEKEAET--MKKMAISEAEKNANVSKILMEQKLSE 235
I + EEE TK+L +EK QKV+ +++ T +K++ + +++A++ +IL E + +
Sbjct: 100 ISSEIQNREEEHTKLLADLEKQQKVASRKSYTHRIKEITKNSRKQDADIERILKETREVQ 279
Query: 236 KDSARRQEEIENAMYLARE 254
+S QE + +A E
Sbjct: 280 LESNSIQERLHRTYAVADE 336
>TC9302 similar to UP|BGLT_TRIRP (P26205) Cyanogenic beta-glucosidase
precursor (Linamarase) (Fragment) , partial (58%)
Length = 867
Score = 28.1 bits (61), Expect = 2.1
Identities = 15/62 (24%), Positives = 34/62 (54%)
Frame = +2
Query: 172 KPNIPESIRHNFEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQ 231
KP+I ++ HN+ + +R+ +A+++ +++ MK M + + + S+IL +
Sbjct: 212 KPSIWDTYAHNYSERIVDRSNGDVAVDEYHRYKEDVGIMKSMNMDAYRFSISWSRILPKG 391
Query: 232 KL 233
KL
Sbjct: 392 KL 397
>BP053180
Length = 497
Score = 27.3 bits (59), Expect = 3.6
Identities = 8/27 (29%), Positives = 16/27 (58%)
Frame = -3
Query: 2 PYYLQVLVPSASPSFKNTMAIVHQVPE 28
P+ ++ + P FK T+ ++H +PE
Sbjct: 150 PFEAELFTSTVPPCFKTTLKVIHNLPE 70
>BP065430
Length = 164
Score = 27.3 bits (59), Expect = 3.6
Identities = 11/19 (57%), Positives = 13/19 (67%)
Frame = +1
Query: 122 HEINQFCSSHSLQQVYIDV 140
H INQFC SL +VY D+
Sbjct: 16 HSINQFCYKVSLYRVYSDI 72
>BP074586
Length = 482
Score = 26.2 bits (56), Expect = 8.1
Identities = 8/16 (50%), Positives = 13/16 (81%)
Frame = -1
Query: 171 TKPNIPESIRHNFEQM 186
T+PN+PE + H+FE +
Sbjct: 194 TEPNLPEDLHHSFEDI 147
>BP049435
Length = 485
Score = 26.2 bits (56), Expect = 8.1
Identities = 14/37 (37%), Positives = 21/37 (55%)
Frame = +3
Query: 187 EEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNAN 223
EEE V +E+++ E E K++A EA+ NAN
Sbjct: 309 EEEPESVYEVLERKRQREIEEWRAKQIASGEAKDNAN 419
>BP034000
Length = 369
Score = 26.2 bits (56), Expect = 8.1
Identities = 10/22 (45%), Positives = 12/22 (54%)
Frame = -3
Query: 112 DKTWIYDKIHHEINQFCSSHSL 133
D W+ HH + FC SHSL
Sbjct: 271 DSYWV*VTHHHRVQFFCVSHSL 206
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.378
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,467,838
Number of Sequences: 28460
Number of extensions: 50286
Number of successful extensions: 257
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 255
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 255
length of query: 331
length of database: 4,897,600
effective HSP length: 91
effective length of query: 240
effective length of database: 2,307,740
effective search space: 553857600
effective search space used: 553857600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146567.11


