

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
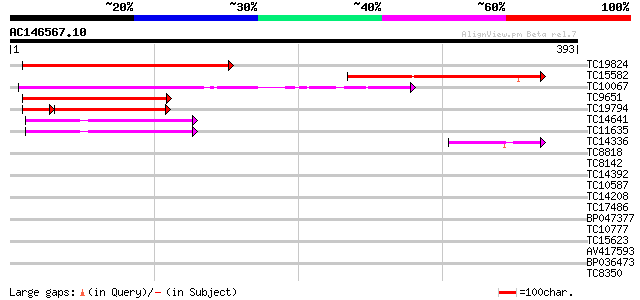
Query= AC146567.10 + phase: 0
(393 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC19824 weakly similar to UP|Q8LEI7 (Q8LEI7) Ras-GTPase-activati... 249 7e-67
TC15582 weakly similar to GB|AAO23575.1|27764908|BT003010 At5g60... 197 2e-51
TC10067 weakly similar to UP|Q8VYJ4 (Q8VYJ4) AT3g25150/MJL12_9, ... 175 1e-44
TC9651 weakly similar to GB|AAO23575.1|27764908|BT003010 At5g609... 131 2e-31
TC19794 similar to GB|AAO23575.1|27764908|BT003010 At5g60980/MSL... 93 3e-24
TC14641 similar to UP|NTF2_ARATH (Q9C7F5) Nuclear transport fact... 57 5e-09
TC11635 similar to PIR|H86398|H86398 protein F17L21.10 [imported... 52 2e-07
TC14336 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, p... 41 3e-04
TC8818 weakly similar to PIR|T48456|T48456 rna binding protein-l... 39 0.001
TC8142 UP|BAD11133 (BAD11133) Glutamate-rich protein, complete 38 0.003
TC14392 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, pa... 37 0.007
TC10587 weakly similar to GB|AAC50895.1|1518802|HSU63289 CUG-BP/... 36 0.013
TC14208 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, pa... 35 0.017
TC17486 similar to UP|RU17_ARATH (Q42404) U1 small nuclear ribon... 33 0.063
BP047377 32 0.14
TC10777 similar to UP|Q9SCL3 (Q9SCL3) Pre-mRNA splicing factor S... 32 0.14
TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding p... 32 0.14
AV417593 32 0.14
BP036473 32 0.24
TC8350 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, par... 32 0.24
>TC19824 weakly similar to UP|Q8LEI7 (Q8LEI7) Ras-GTPase-activating protein
SH3-domain binding protein-like, partial (25%)
Length = 552
Score = 249 bits (635), Expect = 7e-67
Identities = 117/146 (80%), Positives = 133/146 (90%)
Frame = +3
Query: 10 PTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSL 69
PTPQMVGNAFVEQYYSILH DP+QVHRFYH++S++SRPEEDGTMTTVTTT EI+KKI S
Sbjct: 114 PTPQMVGNAFVEQYYSILHHDPEQVHRFYHETSILSRPEEDGTMTTVTTTVEINKKILSQ 293
Query: 70 EYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVF 129
+Y SFRVE+LSAD+QPSY GV+V VTGCLTGTDN+KRKFAQSFFLAPQDKG++VLNDVF
Sbjct: 294 DYKSFRVEILSADSQPSYKEGVIVAVTGCLTGTDNMKRKFAQSFFLAPQDKGYFVLNDVF 473
Query: 130 RYVDAYKSIDIESVPANDADESAPSE 155
RYVD YKS+DIESVP+ DADE S+
Sbjct: 474 RYVDEYKSVDIESVPSRDADEITLSD 551
>TC15582 weakly similar to GB|AAO23575.1|27764908|BT003010
At5g60980/MSL3_100 {Arabidopsis thaliana;}, partial
(16%)
Length = 1053
Score = 197 bits (502), Expect = 2e-51
Identities = 105/139 (75%), Positives = 115/139 (82%), Gaps = 2/139 (1%)
Frame = +3
Query: 235 EQPTSIEKVASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAKPAVHPPRVHSVPAPEA 294
E +S EK SNTQED PKKSFASIVNAL +N+APFH+R SP KP V PRV S PA EA
Sbjct: 9 EHHSSSEKADSNTQEDAPKKSFASIVNALNENAAPFHVRTSPMKP-VEQPRVSSKPASEA 185
Query: 295 PTPNMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNK--G 352
P D+ LEKNNENAG+AHAIFVANLPM+ATVEQL+RAF +FGPIK +GIQVRSNK G
Sbjct: 186 QAPIADVLLEKNNENAGKAHAIFVANLPMNATVEQLERAFVEFGPIKTNGIQVRSNKQQG 365
Query: 353 SCFGFVEFESAASMQSALE 371
SCFGFVEFESAASMQSALE
Sbjct: 366 SCFGFVEFESAASMQSALE 422
>TC10067 weakly similar to UP|Q8VYJ4 (Q8VYJ4) AT3g25150/MJL12_9, partial
(24%)
Length = 903
Score = 175 bits (443), Expect = 1e-44
Identities = 112/276 (40%), Positives = 152/276 (54%), Gaps = 1/276 (0%)
Frame = +1
Query: 7 VQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKI 66
V+TP +VGNAFV+QYY +LH P+ VHRFY + S + RPE++G M TT +I+KKI
Sbjct: 169 VKTPAADIVGNAFVDQYYHMLHESPELVHRFYQEVSKLGRPEQNGLMGITTTMLDINKKI 348
Query: 67 QSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLN 126
SL Y E++S DAQ SY GV+V+VTG + G D IK+KF Q FFLAPQ+KG++VLN
Sbjct: 349 LSLGYDELSAEIISVDAQESYGGGVLVLVTGFMIGKDAIKKKFTQCFFLAPQEKGYFVLN 528
Query: 127 DVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIP 186
DVFRYVDA IE A+D P + P V E IP
Sbjct: 529 DVFRYVDAN---GIEG-SAHDIGSPVPPDNAADPSVLETQVSEEIP-------------- 654
Query: 187 PTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASN 246
+ A+ + +EV P ENG+ S+ E PV V + + P+ + +AS
Sbjct: 655 ----LAAEDGDV--EEVYNP-ENGQASIEEEEAPVPEVLD-----EIPDDSQVVAGLASP 798
Query: 247 TQEDTPKKSFASIVNALKDNSAPFH-LRASPAKPAV 281
+ PKKS+A IV +K+ + P + A P K A+
Sbjct: 799 V-DGAPKKSYAYIVKVMKEVTTPSSTVTAVPVKSAI 903
>TC9651 weakly similar to GB|AAO23575.1|27764908|BT003010
At5g60980/MSL3_100 {Arabidopsis thaliana;}, partial
(21%)
Length = 521
Score = 131 bits (330), Expect = 2e-31
Identities = 62/103 (60%), Positives = 79/103 (76%)
Frame = +1
Query: 10 PTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSL 69
P+PQ+V NAFV+QYY ILH P+ V+RFY DSSV+SRP+ +G MT+VTT I++ I SL
Sbjct: 211 PSPQLVCNAFVDQYYHILHHTPESVYRFYQDSSVVSRPDSNGVMTSVTTMKGINEMIISL 390
Query: 70 EYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQS 112
+ F+ E+ +ADAQ SY GV V+VTGCLTG DN+ RKFAQS
Sbjct: 391 NFKDFKAEIKTADAQKSYKEGVTVLVTGCLTGKDNLTRKFAQS 519
>TC19794 similar to GB|AAO23575.1|27764908|BT003010 At5g60980/MSL3_100
{Arabidopsis thaliana;}, partial (22%)
Length = 472
Score = 93.2 bits (230), Expect(2) = 3e-24
Identities = 45/80 (56%), Positives = 57/80 (71%)
Frame = +1
Query: 32 DQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSFRVEVLSADAQPSYNNGV 91
D VHRF +SS +SR + G MTTVTTT I++ I SL Y + E+ +ADAQ SY GV
Sbjct: 232 DLVHRFDQESSFLSRSDSKGVMTTVTTTKAINENIISLNYEDYTAEIKTADAQESYEKGV 411
Query: 92 MVVVTGCLTGTDNIKRKFAQ 111
+V+VTGCLTG DN+ RKF+Q
Sbjct: 412 IVLVTGCLTGKDNVGRKFSQ 471
Score = 35.0 bits (79), Expect(2) = 3e-24
Identities = 14/22 (63%), Positives = 18/22 (81%)
Frame = +3
Query: 10 PTPQMVGNAFVEQYYSILHRDP 31
P+ Q+VG+A VEQYY ILH+ P
Sbjct: 165 PSAQVVGHALVEQYYHILHQSP 230
>TC14641 similar to UP|NTF2_ARATH (Q9C7F5) Nuclear transport factor 2
(NTF-2), partial (98%)
Length = 760
Score = 57.0 bits (136), Expect = 5e-09
Identities = 40/122 (32%), Positives = 61/122 (49%), Gaps = 3/122 (2%)
Frame = +1
Query: 12 PQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEY 71
P + AFVE YYS + + Y + S+++ + + + I K+ SL +
Sbjct: 40 PDALAKAFVEHYYSTFDGNRAGLANLYQEGSMLTFEGQ-----KIQGSTNIVAKLTSLPF 204
Query: 72 TSFRVEVLSADAQPS-YNNGVMVVVTGCL-TGTDNIKRKFAQSFFLAPQDKG-FYVLNDV 128
+ + D QPS NNG++V V+G L + KF+Q F L P +G +YVLNDV
Sbjct: 205 QQCLHSISTVDCQPSGVNNGMLVFVSGNLQLAGEQHALKFSQMFHLIPTPQGSYYVLNDV 384
Query: 129 FR 130
FR
Sbjct: 385 FR 390
>TC11635 similar to PIR|H86398|H86398 protein F17L21.10 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(98%)
Length = 633
Score = 51.6 bits (122), Expect = 2e-07
Identities = 36/122 (29%), Positives = 59/122 (47%), Gaps = 3/122 (2%)
Frame = +3
Query: 12 PQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEY 71
P + AFVE YYS + + Y + S++S + + ++ I K+ SL +
Sbjct: 51 PDALAKAFVEHYYSTFDSNRAALANLYQEGSMLSFEGQ-----KIQGSSNIVAKLTSLPF 215
Query: 72 TSFRVEVLSADAQPS-YNNGVMVVVTGCL-TGTDNIKRKFAQSFFLAPQDKG-FYVLNDV 128
+ + D QPS + ++V V+G L + F+Q F L P +G +YVLND+
Sbjct: 216 QQCHHSITTVDCQPSGATSAMLVFVSGNLQLAGEQHALNFSQMFHLMPTQQGSYYVLNDI 395
Query: 129 FR 130
FR
Sbjct: 396 FR 401
>TC14336 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, partial
(85%)
Length = 1917
Score = 41.2 bits (95), Expect = 3e-04
Identities = 25/72 (34%), Positives = 37/72 (50%), Gaps = 5/72 (6%)
Frame = +2
Query: 305 KNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIK-----RDGIQVRSNKGSCFGFVE 359
+++ + + + +FV NL S T ++L F +FG I RDG K CFGFV
Sbjct: 338 ESSTDKAKFNNVFVKNLSESTTDDELKNVFGEFGTITSAVVMRDG----DGKSKCFGFVN 505
Query: 360 FESAASMQSALE 371
FE+A A+E
Sbjct: 506 FENADDAARAVE 541
Score = 30.4 bits (67), Expect = 0.53
Identities = 18/57 (31%), Positives = 30/57 (52%), Gaps = 1/57 (1%)
Frame = +2
Query: 316 IFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSN-KGSCFGFVEFESAASMQSALE 371
IF+ NL + + L F FG I + S+ + +GFV+FE+ + Q+A+E
Sbjct: 98 IFIKNLDKAIDHKALHDTFSTFGNILSCKVATDSSGQSKGYGFVQFETEEAAQNAIE 268
>TC8818 weakly similar to PIR|T48456|T48456 rna binding protein-like -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(90%)
Length = 1079
Score = 39.3 bits (90), Expect = 0.001
Identities = 29/93 (31%), Positives = 46/93 (49%), Gaps = 2/93 (2%)
Frame = +3
Query: 273 RASPAKPAVHPPRVHSVPAPEAPTPNMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDR 332
R + AKP+ P + P P +P++ EN G ++V +P ++++
Sbjct: 111 RVTAAKPSDGPDFL-----PLEGGPARKLPVQTPLENKGTV--LYVGRIPHGFYEKEIEG 269
Query: 333 AFKKFGPIKRDGI--QVRSNKGSCFGFVEFESA 363
F +FG IKR I ++ K FGF+EFESA
Sbjct: 270 YFGQFGTIKRLRIARNKKTGKSRHFGFIEFESA 368
>TC8142 UP|BAD11133 (BAD11133) Glutamate-rich protein, complete
Length = 999
Score = 38.1 bits (87), Expect = 0.003
Identities = 41/161 (25%), Positives = 60/161 (36%), Gaps = 19/161 (11%)
Frame = +1
Query: 108 KFAQSFFLAPQDKGFYVLNDVFRYVDAYKSIDIE---SVPANDADESAPSEAI----ITP 160
+F SF L F L+ +F ++ +E P E+ P+E P
Sbjct: 1 EFGTSFSLC-HPCSFPFLSALFESFSPFQMASVEVAQQAPTTVVQENEPTEVTKIQETIP 177
Query: 161 EPEPVHVPEVIPPTQ------------TVIPTAQTVIPPTQTVIADTETIISKEVSLPLE 208
EPE EV P Q T P +T P +T +TE ++ E PLE
Sbjct: 178 EPEQPAATEVAAPEQPAAEVPAPAEPATEEPKEETTEEPKETE-TETEAPVAPETQDPLE 354
Query: 209 NGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASNTQE 249
VTE P E+ ++ E+P + E A+ E
Sbjct: 355 VENKEVTEEAKPEEVKPETEKTEEKTEEPKTEEPAATTETE 477
>TC14392 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, partial (77%)
Length = 1576
Score = 36.6 bits (83), Expect = 0.007
Identities = 29/100 (29%), Positives = 43/100 (43%)
Frame = +3
Query: 292 PEAPTPNMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNK 351
P+A N P NEN IFV NL + T + L + F +G D + V+ +
Sbjct: 885 PKASYQNSQGP---QNENDPNNTTIFVGNLDPNVTDDHLRQVFGLYG----DLVHVKIPQ 1043
Query: 352 GSCFGFVEFESAASMQSALEVWSISINSCYQQGCKLNVGW 391
G GFV+F + + AL V +N G + + W
Sbjct: 1044GKRCGFVQFADRSCAEEALRV----LNGTLLGGQNVRLSW 1151
>TC10587 weakly similar to GB|AAC50895.1|1518802|HSU63289 CUG-BP/hNab50
{Homo sapiens;} , partial (19%)
Length = 482
Score = 35.8 bits (81), Expect = 0.013
Identities = 24/73 (32%), Positives = 37/73 (49%), Gaps = 2/73 (2%)
Frame = +1
Query: 316 IFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKG--SCFGFVEFESAASMQSALEVW 373
+F+ ++P + L AF+ FG + + V G CFGFV ++S + QSA
Sbjct: 19 LFIYHIPQEFGDQDLANAFQPFGRVISAKVFVDKATGVSKCFGFVSYDSPEAAQSA---- 186
Query: 374 SISINSCYQQGCK 386
IS+ + YQ G K
Sbjct: 187 -ISMMNGYQLGGK 222
>TC14208 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, partial (41%)
Length = 837
Score = 35.4 bits (80), Expect = 0.017
Identities = 28/111 (25%), Positives = 41/111 (36%), Gaps = 3/111 (2%)
Frame = +2
Query: 284 PRVHSVPAPEAPTPNMDI---PLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPI 340
P PA P P NEN IFV NL + + L + F +G +
Sbjct: 92 PAATKTPATNQGQPRASYQNHPQGSQNENDPSNTTIFVGNLDPNVNDDHLRQVFSPYGEL 271
Query: 341 KRDGIQVRSNKGSCFGFVEFESAASMQSALEVWSISINSCYQQGCKLNVGW 391
+ V+ G GFV+F + + AL V +N G + + W
Sbjct: 272 ----VHVKIPAGKRCGFVQFADRSGAEEALRV----LNGTLLGGQNVRLSW 400
>TC17486 similar to UP|RU17_ARATH (Q42404) U1 small nuclear
ribonucleoprotein 70 kDa (U1 snRNP 70 kDa) (snRNP70)
(U1-70K), partial (39%)
Length = 595
Score = 33.5 bits (75), Expect = 0.063
Identities = 23/78 (29%), Positives = 33/78 (41%), Gaps = 2/78 (2%)
Frame = +3
Query: 267 SAPFHLRASPA--KPAVHPPRVHSVPAPEAPTPNMDIPLEKNNENAGRAHAIFVANLPMS 324
S P + +PA + VH R+ A A P N + +FVA L
Sbjct: 339 SPPIPVAETPAQRRERVHKLRLEKGAAKAAEELEKYDPQNDPNVSGDPYKTLFVAKLSYE 518
Query: 325 ATVEQLDRAFKKFGPIKR 342
T ++ R F+ +GPIKR
Sbjct: 519 TTESRIKREFESYGPIKR 572
>BP047377
Length = 564
Score = 32.3 bits (72), Expect = 0.14
Identities = 22/66 (33%), Positives = 35/66 (52%), Gaps = 5/66 (7%)
Frame = +3
Query: 316 IFVANLPMSATVEQLDRAFKKFGPIK--RDGIQVRSNKGSCFGFVEF---ESAASMQSAL 370
+FV N+P AT EQL ++ GP+ R I + K +GF E+ E+A S + L
Sbjct: 135 VFVGNIPYDATEEQLIEICQEVGPVVSFRLVIDRETGKPKGYGFCEYKDEETALSARRNL 314
Query: 371 EVWSIS 376
+ + I+
Sbjct: 315 QGYEIN 332
>TC10777 similar to UP|Q9SCL3 (Q9SCL3) Pre-mRNA splicing factor SF2-like
protein, partial (46%)
Length = 597
Score = 32.3 bits (72), Expect = 0.14
Identities = 22/84 (26%), Positives = 40/84 (47%)
Frame = +1
Query: 306 NNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGSCFGFVEFESAAS 365
+N + + I+V NLP +++ F K+G I ++V + C+ FVEF++A
Sbjct: 232 SNMSGRFSRTIYVGNLPGDIRESEIEDIFYKYGRILDIELKV-PPRPPCYCFVEFDNARD 408
Query: 366 MQSALEVWSISINSCYQQGCKLNV 389
+ A+ + GC+L V
Sbjct: 409 AEDAIR----GRDGYNFDGCRLRV 468
>TC15623 similar to UP|P93486 (P93486) Glycine-rich RNA-binding protein
PsGRBP, partial (88%)
Length = 683
Score = 32.3 bits (72), Expect = 0.14
Identities = 19/57 (33%), Positives = 27/57 (47%), Gaps = 2/57 (3%)
Frame = +1
Query: 316 IFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGSC--FGFVEFESAASMQSAL 370
+F+ L + L+ AF FG + + V + G FGFV F+S S SAL
Sbjct: 190 LFIGGLSYGVDDKSLEDAFSSFGTVAEARVIVDRDTGRSRGFGFVSFDSEESASSAL 360
>AV417593
Length = 423
Score = 32.3 bits (72), Expect = 0.14
Identities = 15/49 (30%), Positives = 27/49 (54%)
Frame = +2
Query: 294 APTPNMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKR 342
APT + P ++ + + A++V NLP + T EQ+ + F+ G I +
Sbjct: 194 APTVSWADPKNADSSASSQVKAVYVKNLPKNVTQEQVRKLFEHHGKITK 340
>BP036473
Length = 510
Score = 31.6 bits (70), Expect = 0.24
Identities = 16/58 (27%), Positives = 29/58 (49%)
Frame = +3
Query: 313 AHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGSCFGFVEFESAASMQSAL 370
+ I+V NLP + +++ F KFGP +++ + FV+FE A + A+
Sbjct: 69 SRTIYVGNLPGDIRLREVEDIFYKFGPXVDIDLKIPPRPPG-YAFVQFEDARDAEDAI 239
>TC8350 similar to UP|Q9LEB4 (Q9LEB4) RNA binding protein 45, partial (38%)
Length = 735
Score = 31.6 bits (70), Expect = 0.24
Identities = 29/119 (24%), Positives = 53/119 (44%), Gaps = 6/119 (5%)
Frame = +1
Query: 276 PAKP--AVHPPRVHSVPAPEAPTPNMDIPLEKNNENAGRAHAIFVANLPMSATVEQLDRA 333
P +P A+ PP+ + P P PN+ P ++ A +++ +L L
Sbjct: 184 PQQPPYAMMPPQAQA-PPPHMWAPNVQTP-QQQPATAEEVRTLWIGDLQYWMDENYLYTC 357
Query: 334 FKKFGPIKRDGIQVRSNKGSC----FGFVEFESAASMQSALEVWSISINSCYQQGCKLN 388
F G + ++V NK + +GF+EF S A+ + L+ ++ +I Q +LN
Sbjct: 358 FAHTGEVTT--VKVIRNKQTSQSEGYGFIEFNSRAAAERVLQTFNGAIMPNGGQNFRLN 528
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.129 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,331,045
Number of Sequences: 28460
Number of extensions: 91553
Number of successful extensions: 856
Number of sequences better than 10.0: 92
Number of HSP's better than 10.0 without gapping: 777
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 840
length of query: 393
length of database: 4,897,600
effective HSP length: 92
effective length of query: 301
effective length of database: 2,279,280
effective search space: 686063280
effective search space used: 686063280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146567.10


