

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146566.8 + phase: 0
(465 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
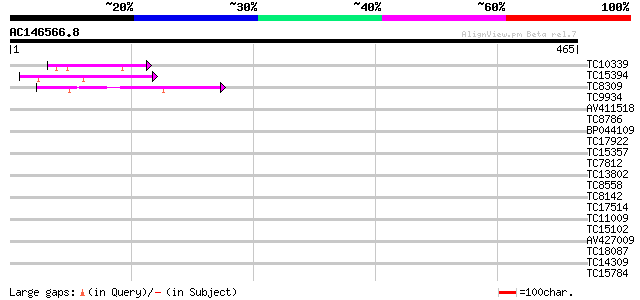
Score E
Sequences producing significant alignments: (bits) Value
TC10339 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protei... 55 3e-08
TC15394 similar to UP|Q9FFQ8 (Q9FFQ8) Emb|CAB62360.1, partial (16%) 42 3e-04
TC8309 UP|O22306 (O22306) Nlj21, complete 40 6e-04
TC9934 similar to UP|BAC98494 (BAC98494) AG-motif binding protei... 39 0.002
AV411518 36 0.015
TC8786 similar to UP|Q9ZTK7 (Q9ZTK7) CONSTANS-like protein 2, pa... 35 0.026
BP044109 35 0.034
TC17922 35 0.034
TC15357 similar to UP|Q9ZV06 (Q9ZV06) At2g39020 protein, partial... 35 0.034
TC7812 UP|Q9LKJ6 (Q9LKJ6) Water-selective transport intrinsic me... 34 0.045
TC13802 similar to UP|Q9M1T1 (Q9M1T1) Sugar-phosphate isomerase-... 34 0.045
TC8558 weakly similar to UP|Q9NDI0 (Q9NDI0) 200 kDa antigen p200... 34 0.059
TC8142 UP|BAD11133 (BAD11133) Glutamate-rich protein, complete 34 0.059
TC17514 weakly similar to UP|O48683 (O48683) F3I6.9 protein, par... 33 0.077
TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Ar... 33 0.10
TC15102 similar to UP|Q7X9B3 (Q7X9B3) 9/13 hydroperoxide lyase, ... 32 0.17
AV427009 32 0.17
TC18087 similar to UP|Q9LF46 (Q9LF46) 2-hydroxyphytanoyl-CoA lya... 32 0.17
TC14309 homologue to UP|H2B_GOSHI (O22582) Histone H2B, partial ... 32 0.17
TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precurso... 32 0.29
>TC10339 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protein L24,
complete
Length = 677
Score = 54.7 bits (130), Expect = 3e-08
Identities = 41/102 (40%), Positives = 59/102 (57%), Gaps = 17/102 (16%)
Frame = +2
Query: 32 KPSKRS-----RKAEKETVA--APKPKKGKTVKTES-STVSASEEVIQKKRNREPEVQDA 83
KPSK + RK K+ +A A K ++ T K S S V A+ EVIQKKR +PEV+DA
Sbjct: 212 KPSKLTWTAMYRKQHKKDIAQEAVKKRRRATKKPYSRSIVGATLEVIQKKRTEKPEVRDA 391
Query: 84 AREAALRE---------DATKREAALQAIRSKRAKGERSIQE 116
AREAALRE D K + A +S++++G+ ++ +
Sbjct: 392 AREAALREIKERIKKTKDEKKAKKAETTAKSQKSQGKGNVSK 517
>TC15394 similar to UP|Q9FFQ8 (Q9FFQ8) Emb|CAB62360.1, partial (16%)
Length = 1093
Score = 41.6 bits (96), Expect = 3e-04
Identities = 33/119 (27%), Positives = 52/119 (42%), Gaps = 6/119 (5%)
Frame = +2
Query: 9 AKSRKTKSKALTKE--DDLVEEATRKPSKRSRKAEKETVAAPKPKKGKTVKTE----SST 62
+K++K SK + KE E T+ K K+ K PK K E S++
Sbjct: 113 SKAQKKTSKKIVKEGSGSKAGERTKSAKKTPVKSSKNVDKTPKKSTPKQTAAEHDSASAS 292
Query: 63 VSASEEVIQKKRNREPEVQDAAREAALREDATKREAALQAIRSKRAKGERSIQERIREA 121
+S S++ KKR E E D +A+ ++ K AL + K G+++ REA
Sbjct: 293 LSKSKQPASKKRKTENEKPDTKGKASSKKQTDKSSKALVKDQGKSKSGKKAKAVPSREA 469
>TC8309 UP|O22306 (O22306) Nlj21, complete
Length = 1063
Score = 40.4 bits (93), Expect = 6e-04
Identities = 38/165 (23%), Positives = 70/165 (42%), Gaps = 10/165 (6%)
Frame = +3
Query: 23 DDLVEEATRKPSKRSRKAEKETVAA-----PKPKKGKTVKTESSTVSASEEVIQKKRNRE 77
+ +VE T +P S K ++E A P+P + E+ T EEV+ E
Sbjct: 276 EQVVETQTPEPEPVSEKTKEEDSAVTEASEPEPTEDNP-SIEAETTEVVEEVVTVTVTDE 452
Query: 78 PEVQDAAREAALREDATKREAALQAIRSKRAKGERS-IQERIREAAEQ----LRKEEADP 132
P+V E+ T EA +A +K K ++ + EA E+ KE+A
Sbjct: 453 PKV----------EEKTDGEAKKEATETKETKESADPVEVQAPEAGEEPETVTLKEDASK 602
Query: 133 SLKKAQKPVSLESPCFRLTPEQEERTREIAANGLAKRRQMTEIYK 177
+K +P + E P++++ ++ G ++++ E+ K
Sbjct: 603 EEEKVAEPEAKEEVVNTEAPDEKKEEEKLDTEGSDEKKEEEEVIK 737
>TC9934 similar to UP|BAC98494 (BAC98494) AG-motif binding protein-4,
partial (37%)
Length = 1050
Score = 38.5 bits (88), Expect = 0.002
Identities = 30/83 (36%), Positives = 40/83 (48%), Gaps = 1/83 (1%)
Frame = -1
Query: 226 HVSGATSSEPASEAP-KSEATRPEAHPSGIPSVPKASVNIQTPILPSSPSSSSSSSSTNS 284
H ATSS + +P K E+ P A + S S + +LP S SSSSSSSS +S
Sbjct: 261 HSRPATSSAGTANSPAKMES*SPAAVELELLSSSIRSCDPPETVLPFSSSSSSSSSS*SS 82
Query: 285 DDIPLSQHIKQCLPNFKPSTSTF 307
PL + K P+T+ F
Sbjct: 81 ISFPLEKSSKSSTEKSSPTTTLF 13
>AV411518
Length = 360
Score = 35.8 bits (81), Expect = 0.015
Identities = 23/60 (38%), Positives = 28/60 (46%), Gaps = 2/60 (3%)
Frame = +3
Query: 231 TSSEPASEAPKSEATRPEAHPSGI--PSVPKASVNIQTPILPSSPSSSSSSSSTNSDDIP 288
TSS PAS +P S A P +PS P+ P P SSPSS+ +S S P
Sbjct: 168 TSSGPASPSPSSTAPSPRPNPSSTSGPTSPTPHTPSPPPPTSSSPSSAIPPTSAPSSSTP 347
>TC8786 similar to UP|Q9ZTK7 (Q9ZTK7) CONSTANS-like protein 2, partial
(33%)
Length = 594
Score = 35.0 bits (79), Expect = 0.026
Identities = 30/122 (24%), Positives = 39/122 (31%), Gaps = 4/122 (3%)
Frame = +1
Query: 344 PLNIQHPETTPTPQKASEV----ALEAATSEDPQQHETSTLHNFEKHLGGEMQPTSTKAS 399
P + H P P A+ A A P +H P ++ A
Sbjct: 94 PSSATHANPPPPPSTAARTPPSSAPPATPKSTPPTSSPPAIHVSLSAKSASKPPLASHAR 273
Query: 400 KTVPEKTVLENQPETSTVPEQSVPEHTVPEKVVSDLPQPNTEQQTENLNHSPTQNQTIPE 459
P T TST P S P VP + S P P T T + QNQ +
Sbjct: 274 PMPP--TSASPATMTSTPPTPSPPATNVPPSLPSSTPPPPTSSTTNTCSQMQNQNQKQKQ 447
Query: 460 QQ 461
Q+
Sbjct: 448 QK 453
>BP044109
Length = 497
Score = 34.7 bits (78), Expect = 0.034
Identities = 25/83 (30%), Positives = 35/83 (42%), Gaps = 13/83 (15%)
Frame = +2
Query: 219 KQALKAKHVSGATS-SEPASEAPKSEATRPEAHPSGIPSV------------PKASVNIQ 265
K H S ATS S P ++ P + P + P PS P +S +
Sbjct: 137 KNKTTPSHASNATSKSTPRTQTPTTPPPSPTSSPKPPPSTFATKPSTSPPETPSSSSHGP 316
Query: 266 TPILPSSPSSSSSSSSTNSDDIP 288
P LPS PSSS+ +S+ + P
Sbjct: 317 APTLPSPPSSSTPTSTPSPPSPP 385
Score = 32.0 bits (71), Expect = 0.22
Identities = 33/149 (22%), Positives = 53/149 (35%), Gaps = 2/149 (1%)
Frame = +2
Query: 312 EQMQIHFSEQRIKICENLNLSADHFFQPPLVEPLNIQHPETTPTPQKASEVALEAATSED 371
+Q H + + + +LS+ HFF P P P K + ATS+
Sbjct: 26 QQQPPHTATTSLSHSSSTSLSSSHFFPSP-------PPPPPPPNKNKTTPSHASNATSKS 184
Query: 372 PQQHETSTLHNFEKHLGGEMQPT--STKASKTVPEKTVLENQPETSTVPEQSVPEHTVPE 429
+ +T T + P+ +TK S + PE + T+P S P + P
Sbjct: 185 TPRTQTPTTPPPSPTSSPKPPPSTFATKPSTSPPETPSSSSHGPAPTLP--SPPSSSTPT 358
Query: 430 KVVSDLPQPNTEQQTENLNHSPTQNQTIP 458
S PN +PT ++P
Sbjct: 359 STPSPPSPPNGSTLPSPPRATPTPAPSLP 445
>TC17922
Length = 470
Score = 34.7 bits (78), Expect = 0.034
Identities = 21/57 (36%), Positives = 28/57 (48%)
Frame = +1
Query: 228 SGATSSEPASEAPKSEATRPEAHPSGIPSVPKASVNIQTPILPSSPSSSSSSSSTNS 284
S TSS P+S +S +A + P P S QTP PSS +S+S S + S
Sbjct: 139 SPPTSSPPSSPPSRSTTPTSQATANAPPYSPNVSTRHQTPSGPSSAASTSHSPTNTS 309
>TC15357 similar to UP|Q9ZV06 (Q9ZV06) At2g39020 protein, partial (69%)
Length = 1578
Score = 34.7 bits (78), Expect = 0.034
Identities = 27/74 (36%), Positives = 35/74 (46%)
Frame = +1
Query: 231 TSSEPASEAPKSEATRPEAHPSGIPSVPKASVNIQTPILPSSPSSSSSSSSTNSDDIPLS 290
TS+ ++ P S A+ + P PS P++S+ P PS PSSSS S T S P S
Sbjct: 487 TSTR*HTKWPSSNASPTSSPPPNPPSPPRSSLRTTNPSTPS-PSSSSKSPRTPS-PTPTS 660
Query: 291 QHIKQCLPNFKPST 304
P KP T
Sbjct: 661 TTTLSTSP*RKPCT 702
>TC7812 UP|Q9LKJ6 (Q9LKJ6) Water-selective transport intrinsic membrane
protein 1, complete
Length = 1149
Score = 34.3 bits (77), Expect = 0.045
Identities = 24/76 (31%), Positives = 35/76 (45%)
Frame = +1
Query: 235 PASEAPKSEATRPEAHPSGIPSVPKASVNIQTPILPSSPSSSSSSSSTNSDDIPLSQHIK 294
PAS P S P + PS PS P + TP SPS+SSS+ ++ S + + +
Sbjct: 241 PASSPPPSPTHSPSSSPS--PSAPTSPAATSTP---PSPSASSSAVTSPSSAVSSTSSLS 405
Query: 295 QCLPNFKPSTSTFQDD 310
P+ P +S D
Sbjct: 406 FSDPSSPPCSSPSSPD 453
>TC13802 similar to UP|Q9M1T1 (Q9M1T1) Sugar-phosphate isomerase-like
protein, partial (41%)
Length = 522
Score = 34.3 bits (77), Expect = 0.045
Identities = 24/60 (40%), Positives = 32/60 (53%), Gaps = 6/60 (10%)
Frame = +2
Query: 230 ATSSEPASEAPKSEATRPEAHPSGIPSVPKASVNIQTPILPS------SPSSSSSSSSTN 283
A SS PAS +P S T+ S S P +S TP+ PS SP+++SSSSS +
Sbjct: 194 APSSSPASASPASSPTKSPKPSSPSASAPPSS----TPLTPSTATSESSPTATSSSSSAS 361
Score = 28.9 bits (63), Expect = 1.9
Identities = 21/60 (35%), Positives = 27/60 (45%), Gaps = 7/60 (11%)
Frame = +2
Query: 230 ATSSEPASEAPKSEATRPEAHPSGIPSVPKASVNIQTPIL-------PSSPSSSSSSSST 282
+T+S P S PK P PS P +S +P PSSPS+S+ SST
Sbjct: 122 STTSSPTSTTPKPS---PSPAPSSTLRAPSSSPASASPASSPTKSPKPSSPSASAPPSST 292
>TC8558 weakly similar to UP|Q9NDI0 (Q9NDI0) 200 kDa antigen p200
(Fragment), partial (4%)
Length = 585
Score = 33.9 bits (76), Expect = 0.059
Identities = 34/111 (30%), Positives = 52/111 (46%), Gaps = 5/111 (4%)
Frame = +1
Query: 60 SSTVSASEEVIQKKRN--REPEVQDAAREAALREDATKREAALQAIRSKRAKGERSIQER 117
S T+ A+ +++RN R E QDAA AAL D R + R QER
Sbjct: 10 SPTLVAARLDAEERRNNIRLREEQDAAYRAALEADQA------------RERQRREEQER 153
Query: 118 I-REAAEQ--LRKEEADPSLKKAQKPVSLESPCFRLTPEQEERTREIAANG 165
+ REAAE RKEE + ++A++ ++ ++ ++ E E A G
Sbjct: 154 LAREAAEAERKRKEEEEAREREAREAAEKQAALAKIRQQKAESLGEEPAKG 306
Score = 26.6 bits (57), Expect = 9.4
Identities = 22/83 (26%), Positives = 39/83 (46%), Gaps = 7/83 (8%)
Frame = +1
Query: 82 DAAREAALREDATKREAALQAIRSKRAKGERSIQERIREAAE------QLRKEEADPSLK 135
D ARE RE+ + KR + E + + REAAE ++R+++A+ +
Sbjct: 112 DQARERQRREEQERLAREAAEAERKRKEEEEAREREAREAAEKQAALAKIRQQKAESLGE 291
Query: 136 KAQKPVSLESPCFRL-TPEQEER 157
+ K ++ R T E++ER
Sbjct: 292 EPAKGPNVTQVLVRFPTGERKER 360
>TC8142 UP|BAD11133 (BAD11133) Glutamate-rich protein, complete
Length = 999
Score = 33.9 bits (76), Expect = 0.059
Identities = 20/79 (25%), Positives = 36/79 (45%), Gaps = 11/79 (13%)
Frame = +1
Query: 392 QPTSTKASKTVPEKTVLENQPETSTVPEQSVPEH--------TVPEKVVSDLP---QPNT 440
Q S + ++ P V EN+P T ++++PE PE+ +++P +P T
Sbjct: 82 QMASVEVAQQAPTTVVQENEPTEVTKIQETIPEPEQPAATEVAAPEQPAAEVPAPAEPAT 261
Query: 441 EQQTENLNHSPTQNQTIPE 459
E+ E P + +T E
Sbjct: 262 EEPKEETTEEPKETETETE 318
>TC17514 weakly similar to UP|O48683 (O48683) F3I6.9 protein, partial (9%)
Length = 582
Score = 33.5 bits (75), Expect = 0.077
Identities = 32/139 (23%), Positives = 53/139 (38%), Gaps = 7/139 (5%)
Frame = +2
Query: 9 AKSRKTKSKALTKEDDLVEEATRKPSKRSRKAEKETVAAPKPKKGKTVKTESSTVSASEE 68
+ + KTK K++ + + +T + SK + K +A KKG + +++SEE
Sbjct: 5 SNAAKTKKKSMLPKSKASQISTPRNSKPASTPSKPLTSASSTKKGNSPSLSRKPITSSEE 184
Query: 69 -------VIQKKRNREPEVQDAAREAALREDATKREAALQAIRSKRAKGERSIQERIREA 121
+ K N EP D A LR+ + I + K ++ Q
Sbjct: 185 SKKVANKPLHKSLNLEPSNPDPAPLPTLRKSFIMESMGDKDIVKRAFKTFKTFQNNY--- 355
Query: 122 AEQLRKEEADPSLKKAQKP 140
Q + D S K Q P
Sbjct: 356 -SQPKSSGEDRSSVKKQVP 409
>TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;}, partial (44%)
Length = 700
Score = 33.1 bits (74), Expect = 0.10
Identities = 21/62 (33%), Positives = 33/62 (52%)
Frame = -2
Query: 223 KAKHVSGATSSEPASEAPKSEATRPEAHPSGIPSVPKASVNIQTPILPSSPSSSSSSSST 282
K+ S ++SS +S + S + P PS PS +S + + P + SSSSSS S+
Sbjct: 234 KSSSSSSSSSSSSSSSSSSSGSPPPPRPPSTPPSSSSSSSSSSSSPPPGASSSSSSSPSS 55
Query: 283 NS 284
+S
Sbjct: 54 SS 49
Score = 32.0 bits (71), Expect = 0.22
Identities = 23/75 (30%), Positives = 35/75 (46%)
Frame = -2
Query: 210 TVREGAALLKQALKAKHVSGATSSEPASEAPKSEATRPEAHPSGIPSVPKASVNIQTPIL 269
TVR + ++ + S ++SS +S + RP + P S +S + P
Sbjct: 264 TVRRTRYSVPKSSSSSSSSSSSSSSSSSSSGSPPPPRPPSTPPSSSSSSSSSSSSPPPGA 85
Query: 270 PSSPSSSSSSSSTNS 284
SS SSS SSSS +S
Sbjct: 84 SSSSSSSPSSSSPSS 40
Score = 31.2 bits (69), Expect = 0.38
Identities = 20/63 (31%), Positives = 30/63 (46%)
Frame = -2
Query: 228 SGATSSEPASEAPKSEATRPEAHPSGIPSVPKASVNIQTPILPSSPSSSSSSSSTNSDDI 287
S ++SS +S + S + P P P P +S + + P +SSSSSS+ S
Sbjct: 222 SSSSSSSSSSSSSSSSGSPPPPRPPSTP--PSSSSSSSSSSSSPPPGASSSSSSSPSSSS 49
Query: 288 PLS 290
P S
Sbjct: 48 PSS 40
Score = 29.6 bits (65), Expect = 1.1
Identities = 21/65 (32%), Positives = 34/65 (52%), Gaps = 6/65 (9%)
Frame = -2
Query: 230 ATSSEPASEAPKSEATRPEAHPSGIPSVPKASVNIQT------PILPSSPSSSSSSSSTN 283
A+SS A+ ++ + P++ S S +S + + P PS+P SSSSSSS++
Sbjct: 288 ASSSSAAATVRRTRYSVPKSSSSSSSSSSSSSSSSSSSGSPPPPRPPSTPPSSSSSSSSS 109
Query: 284 SDDIP 288
S P
Sbjct: 108 SSSPP 94
Score = 29.3 bits (64), Expect = 1.4
Identities = 26/93 (27%), Positives = 42/93 (44%), Gaps = 1/93 (1%)
Frame = -2
Query: 215 AALLKQALKAKHVSGATSSEPASEAPKSEATRPEAHPSGIPSVPKASVNIQTPILPSSPS 274
AA +++ + S ++SS +S + S ++ P PS P PS
Sbjct: 270 AATVRRTRYSVPKSSSSSSSSSSSSSSSSSSSGSPPPPRPPSTP--------------PS 133
Query: 275 SSSSSSSTNSDDIPLSQHIKQCLP-NFKPSTST 306
SSSSSSS++S P + P + PS+S+
Sbjct: 132 SSSSSSSSSSSPPPGASSSSSSSPSSSSPSSSS 34
Score = 29.3 bits (64), Expect = 1.4
Identities = 23/92 (25%), Positives = 39/92 (42%)
Frame = -2
Query: 191 LDAEKAAVIASLAAEIEEETVREGAALLKQALKAKHVSGATSSEPASEAPKSEATRPEAH 250
LDA ++ A++ +V + ++ + + S ++S P P S +
Sbjct: 294 LDASSSSAAATVRRT--RYSVPKSSSSSSSSSSSSSSSSSSSGSPPPPRPPSTPPSSSSS 121
Query: 251 PSGIPSVPKASVNIQTPILPSSPSSSSSSSST 282
S S P + + PSS S SSSSSS+
Sbjct: 120 SSSSSSSPPPGASSSSSSSPSSSSPSSSSSSS 25
Score = 26.9 bits (58), Expect = 7.2
Identities = 25/79 (31%), Positives = 37/79 (46%)
Frame = -2
Query: 228 SGATSSEPASEAPKSEATRPEAHPSGIPSVPKASVNIQTPILPSSPSSSSSSSSTNSDDI 287
SG+ S + +S + + R SVPK+S + SS SSSSSSSS++S
Sbjct: 309 SGSKSLDASSSSAAATVRRTRY------SVPKSSSS-------SSSSSSSSSSSSSSSGS 169
Query: 288 PLSQHIKQCLPNFKPSTST 306
P P+ S+S+
Sbjct: 168 PPPPRPPSTPPSSSSSSSS 112
>TC15102 similar to UP|Q7X9B3 (Q7X9B3) 9/13 hydroperoxide lyase, partial
(68%)
Length = 1097
Score = 32.3 bits (72), Expect = 0.17
Identities = 35/128 (27%), Positives = 51/128 (39%), Gaps = 5/128 (3%)
Frame = +1
Query: 339 PPLVEPLNIQHPETTPTPQK--ASEVALEAATSEDPQQHETSTLHNFEKHLGGEMQPTST 396
PP EP + P + PTP+ +S ++S P+ ++L P S
Sbjct: 247 PPSSEPTCLLAPASPPTPKSSHSSTPPPSRSSSTTPKSKNATSL-----TAPSCPPPASP 411
Query: 397 KASKTVPEKTVLENQPETSTVP---EQSVPEHTVPEKVVSDLPQPNTEQQTENLNHSPTQ 453
A+ P KT N P S+ P + S+P +T P + S P P T SP
Sbjct: 412 AATAPAPSKTP-PNPPTNSSKPSSCKSSLP-NTTPSSLSSAPPSPTT---------SPIS 558
Query: 454 NQTIPEQQ 461
+ PE Q
Sbjct: 559 TPSSPENQ 582
>AV427009
Length = 404
Score = 32.3 bits (72), Expect = 0.17
Identities = 17/41 (41%), Positives = 26/41 (62%)
Frame = +2
Query: 267 PILPSSPSSSSSSSSTNSDDIPLSQHIKQCLPNFKPSTSTF 307
P LP++P+SSS+ SST+S+ P S C N + + S+F
Sbjct: 281 PPLPTTPASSSTISSTSSEKTPSSTRWSSC--NHRNNASSF 397
>TC18087 similar to UP|Q9LF46 (Q9LF46) 2-hydroxyphytanoyl-CoA lyase-like
protein, partial (30%)
Length = 571
Score = 32.3 bits (72), Expect = 0.17
Identities = 25/72 (34%), Positives = 33/72 (45%), Gaps = 4/72 (5%)
Frame = +2
Query: 240 PKSEATRPEAHPSGIPSVPKASVNIQTPILPS----SPSSSSSSSSTNSDDIPLSQHIKQ 295
PKS AT +P + S + ++P PS SPS S+SS ST S + +
Sbjct: 47 PKSTATPSPPNP*PDSASNTCSASSESP*PPSPPAPSPSESASSPSTTSSPLATPPPLTA 226
Query: 296 CLPNFKPSTSTF 307
LP STS F
Sbjct: 227 TLPPAPASTSLF 262
>TC14309 homologue to UP|H2B_GOSHI (O22582) Histone H2B, partial (93%)
Length = 682
Score = 32.3 bits (72), Expect = 0.17
Identities = 18/51 (35%), Positives = 29/51 (56%), Gaps = 2/51 (3%)
Frame = +1
Query: 29 ATRKPS--KRSRKAEKETVAAPKPKKGKTVKTESSTVSASEEVIQKKRNRE 77
A +KP+ K+S AEK A KPK GK + E ++ + ++ + K+N E
Sbjct: 94 AEKKPAEEKKSTAAEKAPPAEKKPKAGKKLPKEGASAAGDKKKKRSKKNVE 246
>TC15784 similar to UP|PE31_ARATH (Q9LHA7) Peroxidase 31 precursor (Atperox
P31) (ATP41) , partial (38%)
Length = 475
Score = 31.6 bits (70), Expect = 0.29
Identities = 23/86 (26%), Positives = 36/86 (41%), Gaps = 6/86 (6%)
Frame = +1
Query: 226 HVSGATSSEPASEAPK------SEATRPEAHPSGIPSVPKASVNIQTPILPSSPSSSSSS 279
H S ++S S P S +T AH S S + N P P+ P S+SSS
Sbjct: 49 HHSSSSSQHSHSSPPSHTRDSHSTSTNTRAHNSPKSSATPSPTNKSPPPPPAPPRSASSS 228
Query: 280 SSTNSDDIPLSQHIKQCLPNFKPSTS 305
++ ++ LP+ KP+ +
Sbjct: 229 TTASTPTAATPPSSSPQLPSTKPNAT 306
Score = 28.1 bits (61), Expect = 3.2
Identities = 18/69 (26%), Positives = 32/69 (46%)
Frame = +1
Query: 219 KQALKAKHVSGATSSEPASEAPKSEATRPEAHPSGIPSVPKASVNIQTPILPSSPSSSSS 278
+ A + S T++ P S +P+ +T+P A P+ T P++PS+SSS
Sbjct: 211 RSASSSTTASTPTAATPPSSSPQLPSTKPNATPTS------------TSPSPATPSTSSS 354
Query: 279 SSSTNSDDI 287
S+ +
Sbjct: 355 ELKPPSNSL 381
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.303 0.119 0.314
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,361,539
Number of Sequences: 28460
Number of extensions: 96508
Number of successful extensions: 1586
Number of sequences better than 10.0: 181
Number of HSP's better than 10.0 without gapping: 1081
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1355
length of query: 465
length of database: 4,897,600
effective HSP length: 93
effective length of query: 372
effective length of database: 2,250,820
effective search space: 837305040
effective search space used: 837305040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 17 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146566.8


