

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146566.2 + phase: 2 /partial
(358 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
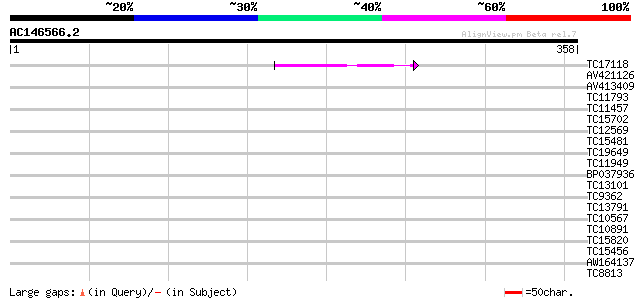
Score E
Sequences producing significant alignments: (bits) Value
TC17118 similar to UP|Q94CL1 (Q94CL1) RING finger-like protein (... 40 8e-04
AV421126 39 0.002
AV413409 38 0.002
TC11793 similar to UP|AAR99376 (AAR99376) Ring domain containing... 37 0.004
TC11457 37 0.004
TC15702 weakly similar to UP|Q8LFU7 (Q8LFU7) Zinc finger protein... 37 0.007
TC12569 UP|Q40221 (Q40221) Protein containing C-terminal RING-fi... 36 0.011
TC15481 similar to UP|Q9LUR1 (Q9LUR1) RING zinc finger protein-l... 35 0.025
TC19649 weakly similar to UP|RNF6_HUMAN (Q9Y252) RING finger pro... 35 0.025
TC11949 35 0.025
BP037936 34 0.033
TC13101 similar to PIR|E86378|E86378 protein F21J9.10 [imported]... 34 0.043
TC9362 similar to PIR|F86340|F86340 protein F2D10.34 [imported] ... 34 0.043
TC13791 weakly similar to UP|RO52_HUMAN (P19474) 52 kDa Ro prote... 33 0.074
TC10567 similar to UP|AAR99376 (AAR99376) Ring domain containing... 33 0.074
TC10891 similar to UP|Q941Q3 (Q941Q3) Floral homeotic protein HU... 33 0.096
TC15820 similar to UP|Q9SWF9 (Q9SWF9) Zinc finger protein, parti... 33 0.096
TC15456 weakly similar to UP|AAS49051 (AAS49051) At5g57740, part... 33 0.096
AW164137 32 0.13
TC8813 weakly similar to UP|Q9XF65 (Q9XF65) RING-H2 zinc finger ... 32 0.13
>TC17118 similar to UP|Q94CL1 (Q94CL1) RING finger-like protein
(AT5g19430/F7K24_180), partial (35%)
Length = 708
Score = 39.7 bits (91), Expect = 8e-04
Identities = 21/91 (23%), Positives = 43/91 (47%)
Frame = +2
Query: 168 HPFRPEEREEHMMSCRNKQKHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHP 227
H F E+H++ + ++ + + C++CL++++ + TA L+ C+H
Sbjct: 245 HDFHDLSIEDHLVEKGSDEETHKGGFGNHGGICAICLDKIVLQETA------LVKGCEHA 406
Query: 228 FCVSCIRNWRSSNPTLGMDVNSTLRACPICR 258
+CV+CI W + N + CP C+
Sbjct: 407 YCVTCILRWATYNNKV---------TCPQCK 472
>AV421126
Length = 416
Score = 38.5 bits (88), Expect = 0.002
Identities = 23/72 (31%), Positives = 33/72 (44%), Gaps = 18/72 (25%)
Frame = +2
Query: 9 GACLKGEHCEFSH--DWKAPPNNI---------------CTFY-QKGVCAYGSRCRYDHV 50
G C G+ C + H D A N+ CT Y Q+G+C +GS C++DH
Sbjct: 161 GECKFGQSCRYHHPPDMGASKANVNLSPVGLPLRPGAPPCTHYTQRGICKFGSACKFDHP 340
Query: 51 KASRAQSSTPSS 62
S + S + SS
Sbjct: 341 IGSLSYSPSASS 376
Score = 27.3 bits (59), Expect = 4.0
Identities = 11/25 (44%), Positives = 13/25 (52%), Gaps = 1/25 (4%)
Frame = +2
Query: 26 PPNNICTFYQK-GVCAYGSRCRYDH 49
P C +Y K G C +G CRY H
Sbjct: 125 PDQPECHYYMKTGECKFGQSCRYHH 199
>AV413409
Length = 374
Score = 38.1 bits (87), Expect = 0.002
Identities = 21/61 (34%), Positives = 27/61 (43%)
Frame = +1
Query: 198 IECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPIC 257
+EC+VCL T L+ +CDH F CI W SS+ T CP+C
Sbjct: 43 LECAVCLNEFEDTETLR-----LIPKCDHVFHPECIDEWLSSHST-----------CPVC 174
Query: 258 R 258
R
Sbjct: 175 R 177
>TC11793 similar to UP|AAR99376 (AAR99376) Ring domain containing protein,
partial (23%)
Length = 527
Score = 37.4 bits (85), Expect = 0.004
Identities = 16/60 (26%), Positives = 28/60 (46%)
Frame = +3
Query: 199 ECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICR 258
+C++CL+ V +++ C H +C CI W +SN +V CP+C+
Sbjct: 348 DCNICLDCVQDP---------VVTLCGHLYCWPCIYKWLNSNSASSENVEQNKPQCPVCK 500
>TC11457
Length = 629
Score = 37.4 bits (85), Expect = 0.004
Identities = 23/70 (32%), Positives = 32/70 (44%), Gaps = 1/70 (1%)
Frame = +1
Query: 199 ECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICR 258
EC+VCL+ + S A +L C+H F + C W S +P CP+CR
Sbjct: 379 ECAVCLDEIDSDQPAR-----VLPGCNHGFHLECADTWLSKHP-----------LCPVCR 510
Query: 259 -KLSYFVVPS 267
KL F P+
Sbjct: 511 AKLDLFSNPA 540
>TC15702 weakly similar to UP|Q8LFU7 (Q8LFU7) Zinc finger protein 3,
partial (17%)
Length = 620
Score = 36.6 bits (83), Expect = 0.007
Identities = 18/59 (30%), Positives = 25/59 (41%), Gaps = 18/59 (30%)
Frame = +3
Query: 9 GACLKGEHCEFSHDWKA-----------------PPNNICTFYQK-GVCAYGSRCRYDH 49
GAC +C+F H P N+C+ Y + G+C +G CRYDH
Sbjct: 6 GACKFKSNCKFHHPRNRVARLPPCILSDKGLPLRPDQNVCSHYSRYGICKFGPACRYDH 182
>TC12569 UP|Q40221 (Q40221) Protein containing C-terminal RING-finger,
complete
Length = 2092
Score = 35.8 bits (81), Expect = 0.011
Identities = 20/83 (24%), Positives = 32/83 (38%)
Frame = +2
Query: 176 EEHMMSCRNKQKHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRN 235
E+ C + + + + E C +CLE + G L C H + V+CI+
Sbjct: 1712 EDSFSQCMTETIYCSSEQGQDEGSCVICLEEYKNMDDV-----GTLKTCGHDYHVNCIKK 1876
Query: 236 WRSSNPTLGMDVNSTLRACPICR 258
W S + CPIC+
Sbjct: 1877 WLSMK-----------KLCPICK 1912
>TC15481 similar to UP|Q9LUR1 (Q9LUR1) RING zinc finger protein-like,
partial (32%)
Length = 706
Score = 34.7 bits (78), Expect = 0.025
Identities = 21/62 (33%), Positives = 26/62 (41%)
Frame = +1
Query: 199 ECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICR 258
EC+VCL S T +L +C H F CI W S+ T CP+CR
Sbjct: 421 ECAVCLSEFESGETGR-----VLPKCKHAFHTDCIDMWFHSHST-----------CPLCR 552
Query: 259 KL 260
L
Sbjct: 553 TL 558
>TC19649 weakly similar to UP|RNF6_HUMAN (Q9Y252) RING finger protein 6
(RING-H2 protein), partial (4%)
Length = 523
Score = 34.7 bits (78), Expect = 0.025
Identities = 23/80 (28%), Positives = 31/80 (38%), Gaps = 4/80 (5%)
Frame = +2
Query: 192 LKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTL 251
L + + C +CL + K T + EC+H F CI W N
Sbjct: 134 LPKPDDKACPICLSEYMPKETVKT-----IPECEHCFHAQCIDEWLPLN----------- 265
Query: 252 RACPIC----RKLSYFVVPS 267
+CPIC RKL + PS
Sbjct: 266 ASCPICRNSPRKLPQRLAPS 325
>TC11949
Length = 780
Score = 34.7 bits (78), Expect = 0.025
Identities = 17/70 (24%), Positives = 30/70 (42%)
Frame = +3
Query: 190 EALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNS 249
+++K ++ CS+C++ V + C H FC SC + S G+
Sbjct: 99 DSIKIDIDLTCSICMDTVFDPVSLT---------CGHIFCYSCACSAASVTIVDGLKAAX 251
Query: 250 TLRACPICRK 259
CP+CR+
Sbjct: 252 PKEKCPLCRE 281
>BP037936
Length = 518
Score = 34.3 bits (77), Expect = 0.033
Identities = 18/77 (23%), Positives = 27/77 (34%), Gaps = 26/77 (33%)
Frame = +3
Query: 2 LCKFFAHGACLKGEHCEFSHDWKAPPNNICTF--------------------------YQ 35
+C+ + C+KG+ C F H + +C F Y+
Sbjct: 282 VCRHWLRSLCMKGDACGFLHQYDKSRMPVCRFFRLYGECREQDCVYKHTNEDIKECNMYK 461
Query: 36 KGVCAYGSRCRYDHVKA 52
G C G CRY H K+
Sbjct: 462 LGFCPNGPDCRYRHAKS 512
>TC13101 similar to PIR|E86378|E86378 protein F21J9.10 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(47%)
Length = 588
Score = 33.9 bits (76), Expect = 0.043
Identities = 25/101 (24%), Positives = 34/101 (32%)
Frame = +3
Query: 196 QEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACP 255
+E EC +CLE +L C H C+ C W N +CP
Sbjct: 144 REDECGICLEPCTKI---------VLPYCCHAMCIKCYCKW-----------NRKSESCP 263
Query: 256 ICRKLSYFVVPSVIWYATSEEKMEIIDTYKAKLKSIDCKHF 296
CR V +W T E + +T + D HF
Sbjct: 264 FCRSSLRRVKSEDLWVLTCNEDVVDAETVSRE----DLLHF 374
>TC9362 similar to PIR|F86340|F86340 protein F2D10.34 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(58%)
Length = 591
Score = 33.9 bits (76), Expect = 0.043
Identities = 19/62 (30%), Positives = 29/62 (46%)
Frame = +1
Query: 199 ECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICR 258
+C++CL AA + +L +C H F VSCI W S+ +CP CR
Sbjct: 325 DCAICLTEF-----AAGDEIRVLPQCGHGFHVSCIDAWLKSH-----------SSCPSCR 456
Query: 259 KL 260
++
Sbjct: 457 QI 462
>TC13791 weakly similar to UP|RO52_HUMAN (P19474) 52 kDa Ro protein (Sjogren
syndrome type A antigen) (SS-A) (Ro(SS-A)) (52 kDa
ribonucleoprotein autoantigen Ro/SS-A), partial (5%)
Length = 545
Score = 33.1 bits (74), Expect = 0.074
Identities = 17/70 (24%), Positives = 31/70 (44%)
Frame = +3
Query: 190 EALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNS 249
+++K ++ C +CL+ V + C H FC C + S + G+
Sbjct: 30 DSIKIDIDLTCPICLDTVFDPVSLT---------CGHIFCYMCACSAASVSIVDGLKSAV 182
Query: 250 TLRACPICRK 259
T + CP+CR+
Sbjct: 183 TKQKCPLCRE 212
>TC10567 similar to UP|AAR99376 (AAR99376) Ring domain containing protein,
partial (27%)
Length = 423
Score = 33.1 bits (74), Expect = 0.074
Identities = 15/60 (25%), Positives = 26/60 (43%)
Frame = +2
Query: 199 ECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICR 258
+CS+CL+RV +++ C H +C CI W + + CP+C+
Sbjct: 83 DCSICLDRVQDP---------VVTLCGHLYCWPCIYKWLNFHSVSSDHEEKKEPQCPVCK 235
>TC10891 similar to UP|Q941Q3 (Q941Q3) Floral homeotic protein HUA1, partial
(15%)
Length = 581
Score = 32.7 bits (73), Expect = 0.096
Identities = 22/73 (30%), Positives = 31/73 (42%), Gaps = 26/73 (35%)
Frame = +3
Query: 3 CKFFA-HGACLKGEHCEFSH--DWKAP------------------PNN----ICTFYQK- 36
C F+ +G C GE C+F H D AP P +C +Y K
Sbjct: 111 CDFYMKNGECKFGERCKFHHPIDRSAPLLSKQASQQTVKLTPAGLPRREGAAMCPYYLKT 290
Query: 37 GVCAYGSRCRYDH 49
G C +G+ C++DH
Sbjct: 291 GTCKFGATCKFDH 329
>TC15820 similar to UP|Q9SWF9 (Q9SWF9) Zinc finger protein, partial (18%)
Length = 663
Score = 32.7 bits (73), Expect = 0.096
Identities = 17/53 (32%), Positives = 28/53 (52%), Gaps = 3/53 (5%)
Frame = +2
Query: 26 PPNNICTFYQK-GVCAYGSRCRYDHVKASRAQ--SSTPSSSITEHQPLVSESA 75
P +C FY + G+C +G C++DH A S+ PS+ + + L S S+
Sbjct: 80 PGEPLCVFYSRYGICKFGPSCKFDHPMGIFAYNISAPPSADASGRRLLGSSSS 238
>TC15456 weakly similar to UP|AAS49051 (AAS49051) At5g57740, partial (24%)
Length = 1013
Score = 32.7 bits (73), Expect = 0.096
Identities = 20/61 (32%), Positives = 26/61 (41%), Gaps = 2/61 (3%)
Frame = +2
Query: 200 CSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNST--LRACPIC 257
C+VCLER + + CDH FC C S+N T + ACP+C
Sbjct: 152 CAVCLER---------KCMVSVEGCDHEFCTQCALYLCSTNSTSTTTTHGPPGSIACPLC 304
Query: 258 R 258
R
Sbjct: 305 R 307
>AW164137
Length = 385
Score = 32.3 bits (72), Expect = 0.13
Identities = 18/66 (27%), Positives = 26/66 (39%), Gaps = 19/66 (28%)
Frame = +3
Query: 3 CKFFAH-GACLKGEHCEFSHDWKA-----------------PPNNICTFYQK-GVCAYGS 43
C+F+ G C G C F H P +C FY + G+C +G
Sbjct: 129 CQFYMKTGDCKFGAVCRFHHPRDRQIPAPDCILSPLGLPLRPGEPLCVFYSRYGICKFGP 308
Query: 44 RCRYDH 49
C++DH
Sbjct: 309 SCKFDH 326
>TC8813 weakly similar to UP|Q9XF65 (Q9XF65) RING-H2 zinc finger protein
ATL6, partial (24%)
Length = 495
Score = 32.3 bits (72), Expect = 0.13
Identities = 16/45 (35%), Positives = 21/45 (46%)
Frame = +1
Query: 198 IECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPT 242
+EC+VCL T L+ +CDH F CI W S+ T
Sbjct: 370 LECAVCLSEFEDAETLR-----LIPKCDHVFHTECIDEWLVSHTT 489
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.135 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,397,983
Number of Sequences: 28460
Number of extensions: 147590
Number of successful extensions: 847
Number of sequences better than 10.0: 97
Number of HSP's better than 10.0 without gapping: 797
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 842
length of query: 358
length of database: 4,897,600
effective HSP length: 91
effective length of query: 267
effective length of database: 2,307,740
effective search space: 616166580
effective search space used: 616166580
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146566.2


