

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
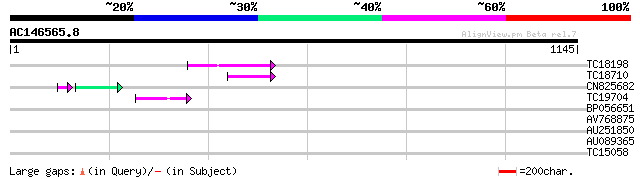
Query= AC146565.8 - phase: 0 /pseudo
(1145 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, p... 98 8e-21
TC18710 63 3e-10
CN825682 39 1e-04
TC19704 weakly similar to UP|Q9SZ87 (Q9SZ87) RNA-directed DNA po... 42 7e-04
BP056651 38 0.010
AV768875 32 0.57
AU251850 31 1.3
AU089365 30 2.2
TC15058 30 2.8
>TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, partial
(14%)
Length = 592
Score = 97.8 bits (242), Expect = 8e-21
Identities = 63/179 (35%), Positives = 91/179 (50%), Gaps = 1/179 (0%)
Frame = -3
Query: 359 GDRNTAYFHRVARIKASSKNISLLYDGATV-LSDAGDIAAHILNYFQGIFDVVNNCIPND 417
GD NT +FH + I + D + + + +NYF I+ +
Sbjct: 569 GDHNTRFFHASTIQRRDFNRILKIKDAHGIWVEGQKKVNEAAVNYFSDIYSSSPLQYLRE 390
Query: 418 LVARSIHALVTEDENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSD 477
+A I LVT+ N+ L DEIK AVF + APG DG G FYQ +IV +D
Sbjct: 389 CLA-PIPKLVTDRLNHKLCAQVSTDEIKEAVFSMGDLKAPGMDGINGLFYQQNREIVKND 213
Query: 478 VVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRL 536
V ++ DFF G +N L+ LIPKIP A+A+ FRPI+ +F +K+++KI+ RL
Sbjct: 212 VNNAILDFFDHGELPVELNETLVTLIPKIPHAEAIQQFRPISCCSFIYKVISKIIVARL 36
>TC18710
Length = 843
Score = 62.8 bits (151), Expect = 3e-10
Identities = 32/97 (32%), Positives = 51/97 (51%)
Frame = +2
Query: 440 LRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADNINSNL 499
L+ +++ F L G DG G FYQ D+V V ++ F + + IN L
Sbjct: 134 LQKKLRLLFFLLETSQPRGMDGMNGLFYQKNGDVVKDSVT*AILAFLERAEIPNEINETL 313
Query: 500 LVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADRL 536
+ LIPK+P +++ FRPI+ F +K+++KI RL
Sbjct: 314 VTLIPKVPHPESINQFRPISGCTFLYKVISKIFVARL 424
>CN825682
Length = 626
Score = 38.9 bits (89), Expect(2) = 1e-04
Identities = 26/95 (27%), Positives = 33/95 (34%)
Frame = -3
Query: 134 WLFIGDFNAVMGAHEKRGRWPPTAASCSDFGSWSNANLLTHLPSSGPLLTWSNGRLGSAY 193
W GDFN ++ EK G F + + L L G TW +
Sbjct: 300 WTCYGDFNEILHHQEKEGARDQDPTRIQVFRDFLDGANLMDLELKGCRFTWLSNLRNGLV 121
Query: 194 VALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHP 228
RLDR + N W Y + L SDH P
Sbjct: 120 TKERLDRVLVNWTWRLDYPNATAMALPIISSDHSP 16
Score = 24.6 bits (52), Expect(2) = 1e-04
Identities = 12/31 (38%), Positives = 16/31 (50%), Gaps = 1/31 (3%)
Frame = -1
Query: 97 WQHSSVFLAA-IYANTSYLFRRLLWADLTHL 126
W + + L IYAN + RR LW +T L
Sbjct: 413 WPNKEIVLGTFIYANPIFRQRRNLWPQITSL 321
>TC19704 weakly similar to UP|Q9SZ87 (Q9SZ87) RNA-directed DNA
polymerase-like protein, partial (3%)
Length = 530
Score = 41.6 bits (96), Expect = 7e-04
Identities = 22/115 (19%), Positives = 51/115 (44%), Gaps = 2/115 (1%)
Frame = -3
Query: 255 DCRRLVLENW--AKNVRGVGMVRLQGKLRILKTVFKQWNRSVFGDVDRQVRLATDEVSRI 312
+C+ + + W L GK K +W+ F D+++ +++ +
Sbjct: 528 ECKETIKKAWDNTSETASSHCSNLSGKTTNTKLALTEWHHQNFSRADKEIAELKNQLEHL 349
Query: 313 QQIIDSVGFSDDLYRQDLEAQLLLTNALNVQDQFWKEKSRHQSFIHGDRNTAYFH 367
Q +S + +++ + ++ L + ++ FW ++SR + GD+NT +FH
Sbjct: 348 QNYSNS----GEDWQEIKQRKMRLKDLWRQEELFWSQRSRIKWLKSGDKNTKFFH 196
>BP056651
Length = 564
Score = 37.7 bits (86), Expect = 0.010
Identities = 26/87 (29%), Positives = 39/87 (43%), Gaps = 2/87 (2%)
Frame = +1
Query: 67 RLPNLWALWNDEVLVVV-IFNSDQCIALEVTWQHSSVFLAAIYANTSYLFRRLLWADLTH 125
R L +W V V F + + L+ W+ S + +Y+ S +R LW DL
Sbjct: 103 RAGGLLCVWKPSVFTVEECFLGEGFLGLKGKWKGVSCVIINVYSPCSRGEKRQLWEDLKA 282
Query: 126 LQGCFLGP-WLFIGDFNAVMGAHEKRG 151
L+ W GDFN+V + E+RG
Sbjct: 283 LRRRLEEDIWCMAGDFNSVRRSEERRG 363
>AV768875
Length = 232
Score = 32.0 bits (71), Expect = 0.57
Identities = 13/36 (36%), Positives = 20/36 (55%)
Frame = +1
Query: 116 RRLLWADLTHLQGCFLGPWLFIGDFNAVMGAHEKRG 151
R +LW L+ ++ PWL IGD N ++ + RG
Sbjct: 4 RDILWQHLSTIRASISIPWLLIGDMNEILFPSQVRG 111
>AU251850
Length = 343
Score = 30.8 bits (68), Expect = 1.3
Identities = 23/88 (26%), Positives = 35/88 (39%), Gaps = 4/88 (4%)
Frame = +2
Query: 1 SSMIVLYWNVRGFGNLDTKIALQNFYLSNKPTFIFLAEPMISFSYVPSWYWHSIGVKKC- 59
++M +L WN RG G+ ALQ P +FL E + + G+ C
Sbjct: 32 TAMKLLSWNCRGLGSPRAVEALQRLVRKEIPQLVFLMETKLK-KFETEKISLGFGLSNCL 208
Query: 60 ---CLNNRGPRLPNLWALWNDEVLVVVI 84
C+ + R L WN V V ++
Sbjct: 209 IVDCVGDEKERKGGLAMWWNASVDVSLV 292
>AU089365
Length = 583
Score = 30.0 bits (66), Expect = 2.2
Identities = 26/93 (27%), Positives = 40/93 (42%), Gaps = 13/93 (13%)
Frame = +2
Query: 972 EFAG*VELDP*CCFSTAICCASFFGAPGPPFLAACSRWEVNCQAGL---LFSHVAHSLCS 1028
E A +E D +TA+ GA + C W VNCQ L F+++ H L
Sbjct: 167 EIAEKIEKDLILLGATAVVGLEVLGAT----MYTCVVWVVNCQMALSISYFTYIQHLLIW 334
Query: 1029 MGVNYLEVFH----------SSISFFYFLEACS 1051
GV + +F S+ ++ F+EAC+
Sbjct: 335 GGVLFWYIFFLXYGAMDPTLSTTAYKVFVEACA 433
>TC15058
Length = 716
Score = 29.6 bits (65), Expect = 2.8
Identities = 16/64 (25%), Positives = 32/64 (50%)
Frame = +1
Query: 56 VKKCCLNNRGPRLPNLWALWNDEVLVVVIFNSDQCIALEVTWQHSSVFLAAIYANTSYLF 115
++ CC+ + L W L+ +++++ N + + L +WQ S LA +S L+
Sbjct: 193 IEVCCMWRQLIALDYCWILYRLLLILILQLNPEN-LTLRDSWQRQSFMLATRVKPSSSLY 369
Query: 116 RRLL 119
+R L
Sbjct: 370 KRFL 381
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.346 0.153 0.561
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,196,558
Number of Sequences: 28460
Number of extensions: 445495
Number of successful extensions: 4593
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 4464
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4590
length of query: 1145
length of database: 4,897,600
effective HSP length: 100
effective length of query: 1045
effective length of database: 2,051,600
effective search space: 2143922000
effective search space used: 2143922000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146565.8


