

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146564.3 - phase: 0 /pseudo
(281 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
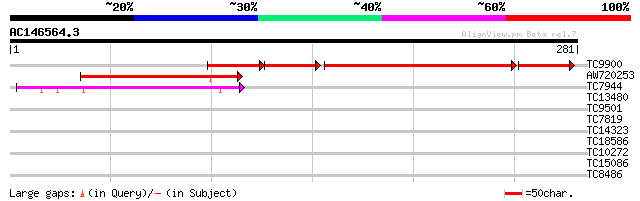
Score E
Sequences producing significant alignments: (bits) Value
TC9900 homologue to UP|EFGC_SOYBN (P34811) Elongation factor G, ... 110 7e-43
AW720253 60 3e-10
TC7944 homologue to UP|Q9ASR1 (Q9ASR1) At1g56070/T6H22_13 (Elong... 52 9e-08
TC13480 weakly similar to UP|Q9AA65 (Q9AA65) Elongation factor T... 28 1.3
TC9501 weakly similar to UP|Q9MBB6 (Q9MBB6) Pectin methylesteras... 27 3.0
TC7819 similar to UP|Q43817 (Q43817) Lipoxygenase , partial (97%) 27 3.0
TC14323 homologue to UP|O22124 (O22124) Proton pyrophosphatase ,... 27 3.0
TC18586 homologue to UP|Q9LTG4 (Q9LTG4) Acetyl-CoA synthetase, p... 26 8.7
TC10272 weakly similar to GB|AAB64934.1|927762|SCD8035 Ydr458cp ... 26 8.7
TC15086 similar to UP|Q8VZF6 (Q8VZF6) AT5g45560/MFC19_23, partia... 26 8.7
TC8486 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Ara... 26 8.7
>TC9900 homologue to UP|EFGC_SOYBN (P34811) Elongation factor G,
chloroplast precursor (EF-G), partial (34%)
Length = 1284
Score = 110 bits (276), Expect(4) = 7e-43
Identities = 58/95 (61%), Positives = 64/95 (67%)
Frame = +3
Query: 157 WGF*RIGKLHEQWCSSRLSGCWCTRSTCG*YLP*CGFKCEGIPVGSKPSFYGRNEKSQTI 216
W RIG++HEQWC +S C CT T G +LP C FK GIPVGS SF GR+ KS T
Sbjct: 492 WRDERIGRMHEQWCPCGISSC*CTSCTHGWFLPRCRFKRAGIPVGS*RSFSGRS*KSWTK 671
Query: 217 NA*TYNET*SCYS*RTL*RCTW*SQQKKRPNHKCW 251
NA*TYNE * CYS*RT C W*SQ KKRPN + W
Sbjct: 672 NA*TYNEG*GCYS*RTSR*CNW*SQLKKRPNQQFW 776
Score = 47.8 bits (112), Expect(4) = 7e-43
Identities = 22/28 (78%), Positives = 25/28 (88%)
Frame = +1
Query: 99 VEANVGAPQANYRESISEVTEVRYVFNK 126
VEANVGAPQ NYRESIS+V+EV+YV K
Sbjct: 286 VEANVGAPQVNYRESISKVSEVKYVHKK 369
Score = 36.6 bits (83), Expect(4) = 7e-43
Identities = 15/28 (53%), Positives = 22/28 (78%)
Frame = +2
Query: 127 PYILNPWTQVVDTNLRIKSKKKHFQKNS 154
PY LNPW+QVVD + R+KSK+ ++ N+
Sbjct: 404 PYGLNPWSQVVDMSSRVKSKEVLYRGNT 487
Score = 35.8 bits (81), Expect(4) = 7e-43
Identities = 15/28 (53%), Positives = 22/28 (78%)
Frame = +1
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELTT 280
+T+GRASY+MQ FD V ++IQN+L +
Sbjct: 859 MTKGRASYTMQLAMFDVVPQHIQNQLAS 942
>AW720253
Length = 481
Score = 60.5 bits (145), Expect = 3e-10
Identities = 29/82 (35%), Positives = 50/82 (60%), Gaps = 2/82 (2%)
Frame = +3
Query: 36 VIKIAIEPKTKADMEKMEAGLIKLAEEDPSFRVSQDKEDRTIIQGTGELHLETIVDRLKR 95
V+++A+EPK AD+ K+ GL +L++ DP + ++ I+ G GELHLE + L+
Sbjct: 69 VVRVAVEPKNPADLPKLVEGLKRLSKSDPCVQCFMEESGEHIVAGAGELHLEICLKDLQD 248
Query: 96 EFK--VEANVGAPQANYRESIS 115
+F VE + P +YRE+++
Sbjct: 249 DFMNGVELKITDPIVSYRETVT 314
>TC7944 homologue to UP|Q9ASR1 (Q9ASR1) At1g56070/T6H22_13 (Elongation factor
EF-2), complete
Length = 2971
Score = 52.4 bits (124), Expect = 9e-08
Identities = 38/119 (31%), Positives = 62/119 (51%), Gaps = 6/119 (5%)
Frame = +1
Query: 4 VTLVGLKDTIA-GETLCDPK--GPFVLDQMDFPVH-VIKIAIEPKTKADMEKMEAGLIKL 59
V LVGL I TL + K + M F V V+++A++ K +D+ K+ GL +L
Sbjct: 1561 VALVGLDQFITKNATLTNEKETDAHPIRAMKFSVSPVVRVAVQCKVASDLPKLVEGLKRL 1740
Query: 60 AEEDPSFRVSQDKEDRTIIQGTGELHLETIVDRLKREFKVEANV--GAPQANYRESISE 116
A+ DP + ++ I+ G GELHLE + L+ +F A + P ++RE++ E
Sbjct: 1741 AKSDPMVVCTIEESGEHIVAGAGELHLEICLKDLQDDFMGGAEIVKSDPVVSFRETVLE 1917
>TC13480 weakly similar to UP|Q9AA65 (Q9AA65) Elongation factor Tu family
protein, partial (21%)
Length = 607
Score = 28.5 bits (62), Expect = 1.3
Identities = 16/56 (28%), Positives = 29/56 (51%), Gaps = 4/56 (7%)
Frame = +1
Query: 78 IQGTGELHLETIVDRLKREFKVEANVGAPQANYR----ESISEVTEVRYVFNKPYI 129
+QG GEL L +++ ++RE E +V P+ Y+ + + V EV N ++
Sbjct: 160 VQGRGELQLGILIENMRRE-GFELSVSPPKVMYKTEKGQKLEPVEEVTIEVNDEHV 324
>TC9501 weakly similar to UP|Q9MBB6 (Q9MBB6) Pectin methylesterase ,
partial (29%)
Length = 934
Score = 27.3 bits (59), Expect = 3.0
Identities = 21/72 (29%), Positives = 31/72 (42%)
Frame = -1
Query: 114 ISEVTEVRYVFNKPYILNPWTQVVDTNLRIKSKKKHFQKNSLYWGF*RIGKLHEQWCSSR 173
++EVT+ R+ + P +L PW V +L ++ F N + L C
Sbjct: 544 MNEVTQSRFHHHGP*VLLPWPSEVRLDLEPHREQCGFWDN--------LTILDHNTCGHV 389
Query: 174 LSGCWCTRSTCG 185
L CWCT CG
Sbjct: 388 L--CWCT--VCG 365
>TC7819 similar to UP|Q43817 (Q43817) Lipoxygenase , partial (97%)
Length = 2842
Score = 27.3 bits (59), Expect = 3.0
Identities = 13/28 (46%), Positives = 20/28 (71%)
Frame = +3
Query: 114 ISEVTEVRYVFNKPYILNPWTQVVDTNL 141
IS T +R+VF +P+IL ++QVV + L
Sbjct: 393 ISNGTSMRWVFPEPFILRIFSQVVSSCL 476
>TC14323 homologue to UP|O22124 (O22124) Proton pyrophosphatase , partial
(74%)
Length = 2664
Score = 27.3 bits (59), Expect = 3.0
Identities = 8/12 (66%), Positives = 9/12 (74%)
Frame = +3
Query: 169 WCSSRLSGCWCT 180
WC R+SG WCT
Sbjct: 1413 WCPRRISGVWCT 1448
>TC18586 homologue to UP|Q9LTG4 (Q9LTG4) Acetyl-CoA synthetase, partial
(27%)
Length = 561
Score = 25.8 bits (55), Expect = 8.7
Identities = 12/36 (33%), Positives = 16/36 (44%), Gaps = 7/36 (19%)
Frame = +2
Query: 156 YWGF*RIGKLHEQWCS-------SRLSGCWCTRSTC 184
YW + H QWC+ ++LSGCW C
Sbjct: 11 YWAQLCNLRTHAQWCN*YCV*RGTQLSGCWSFLGHC 118
>TC10272 weakly similar to GB|AAB64934.1|927762|SCD8035 Ydr458cp
{Saccharomyces cerevisiae;} , partial (3%)
Length = 435
Score = 25.8 bits (55), Expect = 8.7
Identities = 11/23 (47%), Positives = 13/23 (55%), Gaps = 4/23 (17%)
Frame = +3
Query: 166 HEQWCSS----RLSGCWCTRSTC 184
HE+WCSS + S CTR C
Sbjct: 147 HEKWCSSNSSRQASSSLCTRKPC 215
>TC15086 similar to UP|Q8VZF6 (Q8VZF6) AT5g45560/MFC19_23, partial (38%)
Length = 1542
Score = 25.8 bits (55), Expect = 8.7
Identities = 11/23 (47%), Positives = 13/23 (55%)
Frame = +1
Query: 175 SGCWCTRSTCG*YLP*CGFKCEG 197
+GCWC + G*Y GF C G
Sbjct: 874 AGCWCDYNIGG*Y----GFSCTG 930
>TC8486 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Arabidopsis
thaliana;}, partial (25%)
Length = 856
Score = 25.8 bits (55), Expect = 8.7
Identities = 8/13 (61%), Positives = 8/13 (61%)
Frame = -1
Query: 169 WCSSRLSGCWCTR 181
WCS R G WC R
Sbjct: 262 WCSGRRRGSWCRR 224
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.334 0.145 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,541,894
Number of Sequences: 28460
Number of extensions: 64563
Number of successful extensions: 524
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 522
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 524
length of query: 281
length of database: 4,897,600
effective HSP length: 89
effective length of query: 192
effective length of database: 2,364,660
effective search space: 454014720
effective search space used: 454014720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146564.3


