

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146564.14 - phase: 0 /pseudo
(312 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
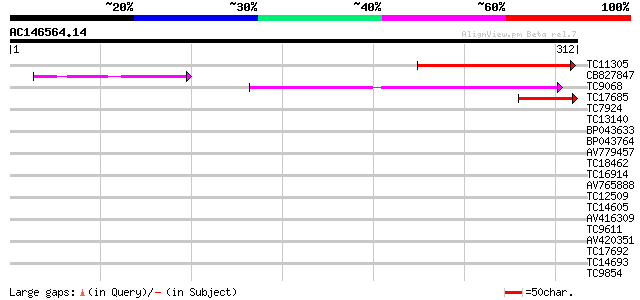
TC11305 weakly similar to UP|AAQ62869 (AAQ62869) At1g12450, part... 117 3e-27
CB827847 73 5e-14
TC9068 similar to UP|SCWA_YEAST (Q04951) Probable family 17 gluc... 70 4e-13
TC17685 weakly similar to UP|AAQ62869 (AAQ62869) At1g12450, part... 51 3e-07
TC7924 homologue to UP|CAE53883 (CAE53883) Aquaporin, partial (95%) 31 0.31
TC13140 weakly similar to UP|RN12_HUMAN (Q9NVW2) RING finger pro... 30 0.53
BP043633 27 0.68
BP043764 29 0.90
AV779457 29 0.90
TC18462 weakly similar to UP|Q945M3 (Q945M3) At1g23890/T23E23_13... 29 1.2
TC16914 similar to UP|RL9_PEA (P30707) 60S ribosomal protein L9 ... 29 1.2
AV765888 29 1.2
TC12509 similar to UP|ACRO_HUMAN (P10323) Acrosin precursor , p... 28 1.5
TC14605 similar to UP|Q9ZV06 (Q9ZV06) At2g39020 protein, partial... 28 2.0
AV416309 28 2.0
TC9611 homologue to UP|Q9FEZ7 (Q9FEZ7) Ribosomal protein L2, par... 28 2.0
AV420351 27 3.4
TC17692 weakly similar to UP|Q8LLW2 (Q8LLW2) Receptor-like kinas... 27 3.4
TC14693 similar to GB|BAB01116.1|11994113|AB026658 CND41, chloro... 27 3.4
TC9854 weakly similar to UP|Q9LSN7 (Q9LSN7) Gb|AAC24081.1 (AT3g1... 27 3.4
>TC11305 weakly similar to UP|AAQ62869 (AAQ62869) At1g12450, partial (20%)
Length = 568
Score = 117 bits (292), Expect = 3e-27
Identities = 56/87 (64%), Positives = 71/87 (81%)
Frame = +1
Query: 225 AVATNVKYGPYIIGSLVGMVPEIFVAIYTGILIKTLADASNERQSLSAQQIILNALGFCL 284
AVAT VKYGPY+ GSLVGMVPE+ V++YTGILI+TLA AS++ ++S QI+LN +G C+
Sbjct: 1 AVATTVKYGPYLCGSLVGMVPEVLVSLYTGILIRTLAHASHKEHTISVVQIVLNVVGLCI 180
Query: 285 TVVTTVIITVYAKRRLKELQEEDKMLL 311
TV T ITVYAKRRLKELQ+ ++LL
Sbjct: 181 TVATVYFITVYAKRRLKELQKGGELLL 261
>CB827847
Length = 503
Score = 73.2 bits (178), Expect = 5e-14
Identities = 39/87 (44%), Positives = 52/87 (58%)
Frame = +2
Query: 14 GGGWKKEVVSDVTITIEADDDNKGDYIKLIPGSDECLPLTAVEMVEECSLPSRRSVVWYW 73
GGG + EVV + D++ GDY+KL PG+ + P P RRS +W+W
Sbjct: 278 GGGRRVEVVCQIE-----DNNINGDYVKLGPGAVDDSPPAP-------PAPGRRSALWFW 421
Query: 74 VKMVLLFLSLGFLAVAVLKWVGPYLID 100
VK+V+L L +G LAV V+KWVGPY ID
Sbjct: 422 VKLVVLILFVGSLAVVVIKWVGPYFID 502
>TC9068 similar to UP|SCWA_YEAST (Q04951) Probable family 17 glucosidase
SCW10 precursor (Soluble cell wall protein 10) ,
partial (6%)
Length = 1209
Score = 70.5 bits (171), Expect = 4e-13
Identities = 45/175 (25%), Positives = 87/175 (49%), Gaps = 3/175 (1%)
Frame = +2
Query: 133 ILLPSTPSMWVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSIF-HHKIEGWLEKYPKKAS 191
+ +P+ P AG+ G G +++ + + S+ F+I F +I +E K +
Sbjct: 374 LAIPAIPLTMSAGLLFGSVTGTVIVSISGTVAASVAFLIARYFARERILKLVEGNKKFLA 553
Query: 192 ILKSAGAGNWFHQFRAVALIRISPF-PYMVFNYCAVATNVKYGPYIIGSLVGMVPEIFVA 250
+ K+ G F+ V L+R+SP P+ + NY T+VK+ PY++GS +GM+P +
Sbjct: 554 VDKAIGENG----FKVVTLLRLSPLLPFSLGNYLYGLTSVKFLPYVLGSWLGMLPGTWAY 721
Query: 251 IYTGILIKTLADASNERQSLSA-QQIILNALGFCLTVVTTVIITVYAKRRLKELQ 304
+ G + + +E + L Q++ LG +T + +T AK +K+++
Sbjct: 722 VSAGAFGRAIIQEESELKVLGGNSQLLTLGLGLLVTALAATYVTRLAKDAIKDIE 886
>TC17685 weakly similar to UP|AAQ62869 (AAQ62869) At1g12450, partial (10%)
Length = 601
Score = 50.8 bits (120), Expect = 3e-07
Identities = 23/32 (71%), Positives = 27/32 (83%)
Frame = +2
Query: 281 GFCLTVVTTVIITVYAKRRLKELQEEDKMLLL 312
GFC+TV TT+++TVYAKRRLKELQ ED LL
Sbjct: 2 GFCVTVATTLVVTVYAKRRLKELQSEDDESLL 97
>TC7924 homologue to UP|CAE53883 (CAE53883) Aquaporin, partial (95%)
Length = 1078
Score = 30.8 bits (68), Expect = 0.31
Identities = 25/81 (30%), Positives = 36/81 (43%)
Frame = -1
Query: 3 DNTDDCGGGGGGGGWKKEVVSDVTITIEADDDNKGDYIKLIPGSDECLPLTAVEMVEECS 62
D +D G G + + V+ + + ADD GD ++ G DE +VE E
Sbjct: 277 DEDEDHAAEGPGNAEEADTVAWSGLLLVADDGGDGD-VEEEEGGDELGNEGSVEG-PELD 104
Query: 63 LPSRRSVVWYWVKMVLLFLSL 83
L W+WV +VL LSL
Sbjct: 103 LSKVDERCWWWVHIVLRHLSL 41
>TC13140 weakly similar to UP|RN12_HUMAN (Q9NVW2) RING finger protein 12
(LIM domain interacting RING finger protein) (RING
finger LIM domain-binding protein) (R-LIM) (NY-REN-43
antigen), partial (4%)
Length = 526
Score = 30.0 bits (66), Expect = 0.53
Identities = 11/14 (78%), Positives = 11/14 (78%)
Frame = +3
Query: 3 DNTDDCGGGGGGGG 16
D DCGGGGGGGG
Sbjct: 63 DRGGDCGGGGGGGG 104
>BP043633
Length = 542
Score = 26.6 bits (57), Expect(2) = 0.68
Identities = 8/21 (38%), Positives = 13/21 (61%)
Frame = -2
Query: 10 GGGGGGGWKKEVVSDVTITIE 30
GG GGGGW + ++ + +E
Sbjct: 304 GGNGGGGWDQAAEAEAEVVVE 242
Score = 21.6 bits (44), Expect(2) = 0.68
Identities = 8/10 (80%), Positives = 8/10 (80%)
Frame = -1
Query: 6 DDCGGGGGGG 15
D GGGGGGG
Sbjct: 431 DGGGGGGGGG 402
>BP043764
Length = 479
Score = 29.3 bits (64), Expect = 0.90
Identities = 12/29 (41%), Positives = 19/29 (65%)
Frame = -2
Query: 9 GGGGGGGGWKKEVVSDVTITIEADDDNKG 37
GGGGGGGG + ++ ++ + A +D KG
Sbjct: 340 GGGGGGGGGGESLLVSQSLLVRA*EDLKG 254
>AV779457
Length = 501
Score = 29.3 bits (64), Expect = 0.90
Identities = 17/48 (35%), Positives = 24/48 (49%), Gaps = 3/48 (6%)
Frame = -3
Query: 135 LPSTPSMWVAGVTLGYGFGFLLII---TAAAIGVSLPFIIGSIFHHKI 179
LP W AG+ + GF FL + + + I +SLP I +HH I
Sbjct: 298 LPLGNLRWKAGLIVRVGFCFLFFLLECSISCIALSLPSFISCNWHHLI 155
>TC18462 weakly similar to UP|Q945M3 (Q945M3) At1g23890/T23E23_13, partial
(26%)
Length = 755
Score = 28.9 bits (63), Expect = 1.2
Identities = 21/64 (32%), Positives = 30/64 (46%)
Frame = +3
Query: 44 PGSDECLPLTAVEMVEECSLPSRRSVVWYWVKMVLLFLSLGFLAVAVLKWVGPYLIDKEV 103
P S +P TA+ CS P + V FL +G L + ++ WV LI EV
Sbjct: 225 PISSFSIPSTALST--PCSSPYPKKV------FSPSFLEMGLLGILMVMWV*LNLISPEV 380
Query: 104 IPII 107
+P+I
Sbjct: 381 LPLI 392
>TC16914 similar to UP|RL9_PEA (P30707) 60S ribosomal protein L9
(Gibberellin-regulated protein GA), complete
Length = 787
Score = 28.9 bits (63), Expect = 1.2
Identities = 21/71 (29%), Positives = 38/71 (52%), Gaps = 1/71 (1%)
Frame = -1
Query: 211 IRISPFPYMVFNYCAVATNVKYGPYIIGSLVGMVPEIFVAIYTGILIKTLADASNERQSL 270
I+I F +++FNYC++ N+ + +P+IFV T +LI + ++ +
Sbjct: 667 IKILFFNHLLFNYCSLLANI--------DTIKELPDIFVLNMT-LLINECTRSRHKLNVI 515
Query: 271 SAQ-QIILNAL 280
S Q Q+IL+ L
Sbjct: 514 SIQNQLILDLL 482
>AV765888
Length = 581
Score = 28.9 bits (63), Expect = 1.2
Identities = 19/60 (31%), Positives = 29/60 (47%), Gaps = 1/60 (1%)
Frame = +2
Query: 78 LLFLSLGFLAVAVLKWVGPYLIDKEVIPIINWETETFS-PPVLTILLFASVAIFPTILLP 136
L+ +S+G + P+L+ +V I W E+ S PP L+ ASV I P + P
Sbjct: 200 LMTVSMGIAKLTPDDTPVPFLVKVDVFTPITWPRESISGPPEFPALIAASV*IAPLMAPP 379
>TC12509 similar to UP|ACRO_HUMAN (P10323) Acrosin precursor , partial (5%)
Length = 710
Score = 28.5 bits (62), Expect = 1.5
Identities = 11/14 (78%), Positives = 11/14 (78%)
Frame = -3
Query: 9 GGGGGGGGWKKEVV 22
GGGGGGGGW VV
Sbjct: 237 GGGGGGGGWSWVVV 196
>TC14605 similar to UP|Q9ZV06 (Q9ZV06) At2g39020 protein, partial (59%)
Length = 1323
Score = 28.1 bits (61), Expect = 2.0
Identities = 15/45 (33%), Positives = 21/45 (46%)
Frame = +2
Query: 8 CGGGGGGGGWKKEVVSDVTITIEADDDNKGDYIKLIPGSDECLPL 52
C GG + V + ADD+NKG + + P S+ C PL
Sbjct: 815 CTFRGGSNSGSFPLRRKVASVLPADDNNKGSFFPMAP-SNPCRPL 946
>AV416309
Length = 277
Score = 28.1 bits (61), Expect = 2.0
Identities = 11/18 (61%), Positives = 12/18 (66%)
Frame = -2
Query: 10 GGGGGGGWKKEVVSDVTI 27
GGGGGGGW VV V +
Sbjct: 141 GGGGGGGWDSTVVVVVVV 88
>TC9611 homologue to UP|Q9FEZ7 (Q9FEZ7) Ribosomal protein L2, partial
(42%)
Length = 394
Score = 28.1 bits (61), Expect = 2.0
Identities = 10/13 (76%), Positives = 11/13 (83%)
Frame = -2
Query: 3 DNTDDCGGGGGGG 15
D+T CGGGGGGG
Sbjct: 75 DHTTHCGGGGGGG 37
>AV420351
Length = 188
Score = 27.3 bits (59), Expect = 3.4
Identities = 10/12 (83%), Positives = 10/12 (83%)
Frame = -2
Query: 8 CGGGGGGGGWKK 19
CGGGGGGGG K
Sbjct: 91 CGGGGGGGGGMK 56
>TC17692 weakly similar to UP|Q8LLW2 (Q8LLW2) Receptor-like kinase
Xa21-binding protein 3, partial (19%)
Length = 482
Score = 27.3 bits (59), Expect = 3.4
Identities = 11/14 (78%), Positives = 12/14 (85%)
Frame = -1
Query: 9 GGGGGGGGWKKEVV 22
GGGGGGGG + EVV
Sbjct: 170 GGGGGGGGGRGEVV 129
>TC14693 similar to GB|BAB01116.1|11994113|AB026658 CND41, chloroplast
nucleoid DNA binding protein-like {Arabidopsis
thaliana;} , partial (58%)
Length = 2067
Score = 27.3 bits (59), Expect = 3.4
Identities = 10/15 (66%), Positives = 11/15 (72%)
Frame = -1
Query: 5 TDDCGGGGGGGGWKK 19
T + GG GGGGGW K
Sbjct: 132 TVNWGGAGGGGGWSK 88
>TC9854 weakly similar to UP|Q9LSN7 (Q9LSN7) Gb|AAC24081.1
(AT3g17100/K14A17_22_), partial (43%)
Length = 1108
Score = 27.3 bits (59), Expect = 3.4
Identities = 9/9 (100%), Positives = 9/9 (100%)
Frame = -3
Query: 9 GGGGGGGGW 17
GGGGGGGGW
Sbjct: 32 GGGGGGGGW 6
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.142 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,879,310
Number of Sequences: 28460
Number of extensions: 90488
Number of successful extensions: 1741
Number of sequences better than 10.0: 87
Number of HSP's better than 10.0 without gapping: 1081
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1464
length of query: 312
length of database: 4,897,600
effective HSP length: 90
effective length of query: 222
effective length of database: 2,336,200
effective search space: 518636400
effective search space used: 518636400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146564.14


