

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
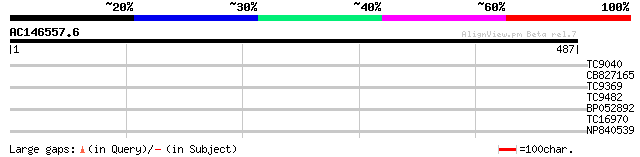
Query= AC146557.6 - phase: 0 /pseudo
(487 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC9040 similar to UP|RL71_ARATH (P60040) 60S ribosomal protein L... 30 1.2
CB827165 29 1.5
TC9369 similar to UP|Q93X72 (Q93X72) Leucine-rich repeat resista... 28 2.6
TC9482 similar to UP|Q9FI73 (Q9FI73) Mitochondrial carrier prote... 27 5.7
BP052892 27 5.7
TC16970 homologue to UP|ZDS_TARER (Q9FV46) Zeta-carotene desatur... 27 9.8
NP840539 hypothetical protein [Lotus corniculatus var. japonicus] 27 9.8
>TC9040 similar to UP|RL71_ARATH (P60040) 60S ribosomal protein L7-1,
complete
Length = 941
Score = 29.6 bits (65), Expect = 1.2
Identities = 19/60 (31%), Positives = 31/60 (51%), Gaps = 1/60 (1%)
Frame = +1
Query: 282 EELIR-KVFLEIKGVELPNPFPRLTYAEAMNRYGSDHPDTRFDLELKDVSDIFSGTSFKI 340
+EL+R K ++KG NP +L + + + HP TR L+L + IF+G K+
Sbjct: 235 KELVRLKREAKLKGGFYVNPEAKLLFIVRIRGINAMHPKTRKILQLLRLRQIFNGVFLKV 414
>CB827165
Length = 550
Score = 29.3 bits (64), Expect = 1.5
Identities = 16/31 (51%), Positives = 21/31 (67%), Gaps = 2/31 (6%)
Frame = -3
Query: 333 FSGTSFKIFS--DSLECGGVIKVLCVPSGAT 361
FSG +FK FS D+LE GG+ LC+P A+
Sbjct: 389 FSG-AFKFFSWSDALEPGGIAGALCIPFSAS 300
>TC9369 similar to UP|Q93X72 (Q93X72) Leucine-rich repeat resistance
protein-like protein, partial (70%)
Length = 859
Score = 28.5 bits (62), Expect = 2.6
Identities = 17/55 (30%), Positives = 29/55 (51%), Gaps = 3/55 (5%)
Frame = -3
Query: 180 SPNEEIRLRYRCLDLRKQQMNSNIL---LRHNVVKLIRRYLEDIHGFVEPCHKAL 231
SP + ++LRYRC+ L ++ I L H + ++RR+ I G + P + L
Sbjct: 197 SPLKLLKLRYRCVKLLSSPISGGIASCSLFHLKLSILRRFNRPI*GGIGPVNLLL 33
>TC9482 similar to UP|Q9FI73 (Q9FI73) Mitochondrial carrier protein-like
(At5g48970), partial (42%)
Length = 529
Score = 27.3 bits (59), Expect = 5.7
Identities = 13/46 (28%), Positives = 23/46 (49%)
Frame = +3
Query: 334 SGTSFKIFSDSLECGGVIKVLCVPSGATKDSYILDIYNEAEKYGAK 379
S +SF++F L G K++C P K + ++ +YGA+
Sbjct: 156 SPSSFQLFLCGLAAGTCAKLVCHPLDVVKKRFQIEGLQRHPRYGAR 293
>BP052892
Length = 501
Score = 27.3 bits (59), Expect = 5.7
Identities = 12/21 (57%), Positives = 16/21 (76%)
Frame = +1
Query: 278 LSLNEELIRKVFLEIKGVELP 298
LS++ +L+ KVFLE G ELP
Sbjct: 205 LSISRKLVSKVFLEDXGFELP 267
>TC16970 homologue to UP|ZDS_TARER (Q9FV46) Zeta-carotene desaturase,
chloroplast precursor (Carotene 7,8-desaturase) ,
partial (24%)
Length = 686
Score = 26.6 bits (57), Expect = 9.8
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = +1
Query: 422 LFHAGHHASVNKTLGHLRVYVAHKLGLIDHAK 453
L+H HH V K LGHL + +A+ + D K
Sbjct: 154 LYHCSHHLKV*K*LGHLLLKLANLCTVRDLVK 249
>NP840539 hypothetical protein [Lotus corniculatus var. japonicus]
Length = 3654
Score = 26.6 bits (57), Expect = 9.8
Identities = 16/34 (47%), Positives = 18/34 (52%)
Frame = +1
Query: 22 KTKIPLPYYEKLKPKTKKAITFKKPKPLNQSLQW 55
K K LPYYEK+ K ITFK +LQW
Sbjct: 961 KLKEALPYYEKVMDK----ITFKSELHGLAALQW 1050
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,208,197
Number of Sequences: 28460
Number of extensions: 111263
Number of successful extensions: 670
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 667
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 670
length of query: 487
length of database: 4,897,600
effective HSP length: 94
effective length of query: 393
effective length of database: 2,222,360
effective search space: 873387480
effective search space used: 873387480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC146557.6


