

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146556.9 + phase: 0
(2244 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
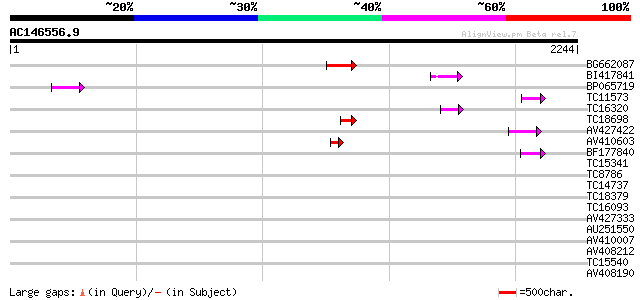
Score E
Sequences producing significant alignments: (bits) Value
BG662087 136 4e-32
BI417841 71 2e-12
BP065719 69 6e-12
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 63 6e-10
TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial ... 59 1e-08
TC18698 57 2e-08
AV427422 55 1e-07
AV410603 53 6e-07
BF177840 48 1e-05
TC15341 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich pro... 38 0.020
TC8786 similar to UP|Q9ZTK7 (Q9ZTK7) CONSTANS-like protein 2, pa... 37 0.034
TC14737 similar to UP|Q9LEB3 (Q9LEB3) RNA binding protein 47, pa... 36 0.058
TC18379 similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich glycop... 35 0.17
TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transc... 35 0.17
AV427333 34 0.22
AU251550 34 0.22
AV410007 34 0.22
AV408212 34 0.29
TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fra... 34 0.29
AV408190 33 0.64
>BG662087
Length = 373
Score = 136 bits (342), Expect = 4e-32
Identities = 64/119 (53%), Positives = 85/119 (70%)
Frame = +1
Query: 1252 GKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNQIKMSPEDREK 1311
GK RM VD+ DLNKA PKD++PLP ID LVD + +++ S MD +SGY+QIKM P D +K
Sbjct: 16 GKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDEDK 195
Query: 1312 TSFITPWGTFCYKVMPFGLINAGATYQRGMTTLFHDMIHKEVEVYVDDMIVKSTDEEQH 1370
T+F+T +CY+ +PFGL NAGATYQ M +F D + + +EVY+D+MIVKS H
Sbjct: 196 TAFMTARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKSALRANH 372
>BI417841
Length = 617
Score = 71.2 bits (173), Expect = 2e-12
Identities = 40/131 (30%), Positives = 65/131 (49%), Gaps = 2/131 (1%)
Frame = +1
Query: 1664 LVFDGAV--NAYGKGIGAVIVSPQGHHIPFTARILFECTNNMAEYEACIFGIEEAIDMRI 1721
L FDG+ N G GAV+ + G + + + TNN AEY I G++ A +
Sbjct: 142 LEFDGSSKGNPGSAGAGAVLRAEDGSKV-YLREGVGNQTNNQAEYRGLILGLKHAHEQGY 318
Query: 1722 KHLDIYGDSALVINQIKGEWETHHAKLIPYRDYARRLLTYFTKVELHHIPRDENQMADAL 1781
+H+++ GDS LV Q++G W+ + + + A+ L + F +++H+PR N AD
Sbjct: 319 QHINVKGDSQLVCKQVEGSWKARNPNIASLCNEAKELKSKFQSFDINHVPRQYNSEADVQ 498
Query: 1782 ATLSSMFRVNH 1792
A L H
Sbjct: 499 ANLGVNLPAGH 531
>BP065719
Length = 567
Score = 69.3 bits (168), Expect = 6e-12
Identities = 40/132 (30%), Positives = 72/132 (54%), Gaps = 2/132 (1%)
Frame = +3
Query: 166 VSKVDVPKKFKVPEFDKYNGLTCPQN--HIVKYVRKMGNYKDNDSLMIHYFQDSLMEDAA 223
V + ++P+ +KVP+F K++G + HI +Y + G+ N++L + YF SL ++A
Sbjct: 147 VLESELPRGWKVPKFTKFSGDSGESTVEHIARYQIEAGDLAINENLKMKYFPSSLTKNAF 326
Query: 224 EWYTSLSKDDVHTFDELATAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAA 283
W+T+L+ VHT+ +L F + F K + + L S+ +K ES +Y R+R
Sbjct: 327 TWFTTLAPRSVHTWAQLERIFHEQF-FRGECKVSXKDLASVKRKPAESIDDYLNRFRMLK 503
Query: 284 ARITPALDEEEM 295
+R + E E+
Sbjct: 504 SRCFTHVSEHEL 539
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 62.8 bits (151), Expect = 6e-10
Identities = 36/96 (37%), Positives = 49/96 (50%), Gaps = 1/96 (1%)
Frame = +2
Query: 2026 DNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEAANKNI-KRIVQKMVTTYKDWHE 2084
+NG + Q C+ I+ SS PQ NG EAANK I K I +++ W +
Sbjct: 8 ENGIQFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWID 187
Query: 2085 MLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEV 2120
LP L Y T +SS TPF L YG + +LP+E+
Sbjct: 188 ELPIVLWSYNTMPQSSIKETPF*LTYGADTMLPVEI 295
>TC16320 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (11%)
Length = 632
Score = 58.5 bits (140), Expect = 1e-08
Identities = 29/91 (31%), Positives = 44/91 (47%)
Frame = +2
Query: 1706 YEACIFGIEEAIDMRIKHLDIYGDSALVINQIKGEWETHHAKLIPYRDYARRLLTYFTKV 1765
Y I G++ AI KH+ + GDS LV NQ++G W+ + + A+ L F
Sbjct: 2 YRGLILGLKHAIKEGYKHIQVKGDSMLVCNQVQGLWKIKNQNIASLCSEAKELKNKFLSF 181
Query: 1766 ELHHIPRDENQMADALATLSSMFRVNHWNDV 1796
+++HIPR+ N AD A R +V
Sbjct: 182 KINHIPREYNSEADVQANFGISLRAGQVEEV 274
>TC18698
Length = 808
Score = 57.4 bits (137), Expect = 2e-08
Identities = 31/62 (50%), Positives = 39/62 (62%)
Frame = -2
Query: 1309 REKTSFITPWGTFCYKVMPFGLINAGATYQRGMTTLFHDMIHKEVEVYVDDMIVKSTDEE 1368
++KT+ + Y+VMP GL N TYQR M +FH I K VEVYV+DMIVKS+ E
Sbjct: 807 KKKTTLKINRVNYYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKSSQE* 628
Query: 1369 QH 1370
H
Sbjct: 627 FH 622
>AV427422
Length = 417
Score = 55.1 bits (131), Expect = 1e-07
Identities = 33/133 (24%), Positives = 63/133 (46%), Gaps = 2/133 (1%)
Frame = +1
Query: 1975 SNGHRFILVAIDYFTKWVEAASYTN-VTKQVVAKFIKNNIICRYGVPSKIITDNGTNLNN 2033
S G+ +LV +D +K+ + T +V+A ++ +GVP I++D +
Sbjct: 19 SKGYEAVLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDPLFMS 198
Query: 2034 NVVQALCEEFKIEHHNSSPYRPQMNGAVEAANKNIKRIVQKMVTTY-KDWHEMLPYALHG 2092
N + L + + S+ Y P+ +G E N+ ++ ++ + K W +P+A +
Sbjct: 199 NFWKELFKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPWAEYW 378
Query: 2093 YRTTVRSSTGATP 2105
Y T+ STG TP
Sbjct: 379 YNTSYHVSTGQTP 417
>AV410603
Length = 162
Score = 52.8 bits (125), Expect = 6e-07
Identities = 26/51 (50%), Positives = 34/51 (65%)
Frame = +1
Query: 1269 KDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNQIKMSPEDREKTSFITPWG 1319
KD+FP+P +D L+D S+ FS +D SGY+QI + PEDR KT F T G
Sbjct: 10 KDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTHHG 162
>BF177840
Length = 410
Score = 48.1 bits (113), Expect = 1e-05
Identities = 26/101 (25%), Positives = 49/101 (47%), Gaps = 1/101 (0%)
Frame = +2
Query: 2021 SKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEAANKNIKRIVQKMVT-TY 2079
+ I++D T ++ + L + + S+ PQ +G E NK + +++ ++
Sbjct: 11 TSIVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNL 190
Query: 2080 KDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEV 2120
K W LP+ Y V S+T +PF +VYG + PL++
Sbjct: 191 KMWETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDL 313
>TC15341 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich protein,
partial (33%)
Length = 1002
Score = 37.7 bits (86), Expect = 0.020
Identities = 37/114 (32%), Positives = 45/114 (39%), Gaps = 32/114 (28%)
Frame = +1
Query: 372 AGQAMATVAPINAAQMPPSYPYMPYSQHPFFPPFYHQYP----LPPGQ----PQV----- 418
A Q A + A P+Y Y+P + P Q P LPP Q PQ+
Sbjct: 607 AAQPQAPAPNVTQAPQQPAY-YIPSTPLPGNTQTVAQLPQNQYLPPDQQYRTPQMQDMSR 783
Query: 419 -------------PVNAIAQQMKQQLPVQQQQQQQQQQQ------QYTRPTFPP 453
P + Q + Q P QQQQQQQQQQQ Q +PT PP
Sbjct: 784 VAPQPCTNXSGXSPPPPVQQYPQYQQPQQQQQQQQQQQQWPQQLPQQVQPTQPP 945
>TC8786 similar to UP|Q9ZTK7 (Q9ZTK7) CONSTANS-like protein 2, partial
(33%)
Length = 594
Score = 37.0 bits (84), Expect = 0.034
Identities = 26/119 (21%), Positives = 52/119 (42%), Gaps = 2/119 (1%)
Frame = +1
Query: 31 ATQEQPVPVVSTAANVTPTVT--TDPRSIMPSGYPYGLPLYYTANTGAGTSGTTNNGPIP 88
AT P P STAA P+ P+S P+ P + + +A + + ++ P+P
Sbjct: 103 ATHANPPPPPSTAARTPPSSAPPATPKSTPPTSSPPAIHVSLSAKSASKPPLASHARPMP 282
Query: 89 GANLTPVSTALTQAATTVTEPIVNAVPLFVHANAHHGSTATTGTIEERMAELAKELRRE 147
+ +P + +T T + P N P + +++TT T + + K+ +++
Sbjct: 283 PTSASPAT--MTSTPPTPSPPATNVPPSLPSSTPPPPTSSTTNTCSQMQNQNQKQKQQK 453
>TC14737 similar to UP|Q9LEB3 (Q9LEB3) RNA binding protein 47, partial (32%)
Length = 803
Score = 36.2 bits (82), Expect = 0.058
Identities = 27/72 (37%), Positives = 32/72 (43%)
Frame = +3
Query: 374 QAMATVAPINAAQMPPSYPYMPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPV 433
Q MA P A M M Y QH + P+ H + P Q Q A A + QQ
Sbjct: 213 QWMAMQYPATAMAMMQQQMMM-YPQH--YMPYVHHHYQPHHQQQQAAAAAAGAVPQQQHQ 383
Query: 434 QQQQQQQQQQQQ 445
Q QQQQ+QQQ
Sbjct: 384 QHHHQQQQKQQQ 419
>TC18379 similar to UP|Q9SBM1 (Q9SBM1) Hydroxyproline-rich glycoprotein
DZ-HRGP precursor, partial (8%)
Length = 584
Score = 34.7 bits (78), Expect = 0.17
Identities = 17/40 (42%), Positives = 20/40 (49%)
Frame = +1
Query: 380 APINAAQMPPSYPYMPYSQHPFFPPFYHQYPLPPGQPQVP 419
AP + P YP +PY QH F+PP P PP P P
Sbjct: 172 APPYGMEFSP-YPSLPYPQHQFYPPPMAPPPPPPPAPAPP 288
>TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transcription
factor PosF21 (AtbZIP59), partial (17%)
Length = 570
Score = 34.7 bits (78), Expect = 0.17
Identities = 15/21 (71%), Positives = 17/21 (80%)
Frame = +1
Query: 425 QQMKQQLPVQQQQQQQQQQQQ 445
QQ +QQ P+QQ QQQ QQQQQ
Sbjct: 163 QQQQQQQPIQQPQQQMQQQQQ 225
Score = 32.0 bits (71), Expect = 1.1
Identities = 15/32 (46%), Positives = 20/32 (61%)
Frame = +1
Query: 415 QPQVPVNAIAQQMKQQLPVQQQQQQQQQQQQY 446
Q Q P+ QQ +Q + QQQ QQQQQ+Q+
Sbjct: 139 QFQQPIQQQQQQQQQPIQQPQQQMQQQQQEQH 234
Score = 30.0 bits (66), Expect = 4.1
Identities = 28/95 (29%), Positives = 35/95 (36%), Gaps = 4/95 (4%)
Frame = +1
Query: 392 PYMPYSQ----HPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQQQQQQQQQYT 447
P M Y+ F+P + + L Q Q I Q QQ Q QQ QQQQQQ
Sbjct: 4 PMMNYASFGGGQQFYPHNHSMHTLLAAQ-QFQQLQIHPQKHQQHQHQFQQPIQQQQQQQQ 180
Query: 448 RPTFPPIPMLYAELLPTLLQRGHCTTRQGKPPPDP 482
+P P + + Q G R P P
Sbjct: 181 QPIQQPQQQMQQQQQEQHQQSGDLKMRSATSSPSP 285
>AV427333
Length = 387
Score = 34.3 bits (77), Expect = 0.22
Identities = 30/104 (28%), Positives = 36/104 (33%), Gaps = 6/104 (5%)
Frame = +3
Query: 388 PPSYPYMPYS--QHPFFPP----FYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQQQQQ 441
PPS+PY P + HPF PP Y + P PP P P
Sbjct: 87 PPSHPYNPPTPPSHPFNPPPTPSHYFKSPPPPSHSFAPPPRGHPSPPPSSPPPPSSHPFT 266
Query: 442 QQQQYTRPTFPPIPMLYAELLPTLLQRGHCTTRQGKPPPDPLPP 485
+ RP PP P P + +PPP PLPP
Sbjct: 267 PPPPHVRP--PPSPHHPITPPPHV-----------RPPPPPLPP 359
>AU251550
Length = 329
Score = 34.3 bits (77), Expect = 0.22
Identities = 18/40 (45%), Positives = 21/40 (52%), Gaps = 1/40 (2%)
Frame = +2
Query: 388 PPSYPYMPYSQHPFFPPFYHQYPLPP-GQPQVPVNAIAQQ 426
PP P PY+ P PPF H +P PP G P P A+ Q
Sbjct: 140 PPGPPRDPYAPAPPPPPFGHHHPPPPHGPPSPPPPAMFYQ 259
>AV410007
Length = 425
Score = 34.3 bits (77), Expect = 0.22
Identities = 18/31 (58%), Positives = 22/31 (70%)
Frame = +3
Query: 421 NAIAQQMKQQLPVQQQQQQQQQQQQYTRPTF 451
N AQQ +QQ QQQ+QQ +QQQ +RPTF
Sbjct: 210 NTKAQQQRQQ---QQQRQQAKQQQNGSRPTF 293
>AV408212
Length = 362
Score = 33.9 bits (76), Expect = 0.29
Identities = 16/24 (66%), Positives = 18/24 (74%)
Frame = +3
Query: 422 AIAQQMKQQLPVQQQQQQQQQQQQ 445
A+ QQ +QQ QQQQQQQQ QQQ
Sbjct: 84 ALQQQQQQQQQQQQQQQQQQSQQQ 155
Score = 33.1 bits (74), Expect = 0.49
Identities = 16/25 (64%), Positives = 18/25 (72%)
Frame = +3
Query: 421 NAIAQQMKQQLPVQQQQQQQQQQQQ 445
+A QQ +QQ QQQQQQQQQ QQ
Sbjct: 78 HAALQQQQQQQQQQQQQQQQQQSQQ 152
Score = 33.1 bits (74), Expect = 0.49
Identities = 16/26 (61%), Positives = 18/26 (68%)
Frame = +3
Query: 424 AQQMKQQLPVQQQQQQQQQQQQYTRP 449
A Q +QQ QQQQQQQQQQ Q +P
Sbjct: 84 ALQQQQQQQQQQQQQQQQQQSQQQQP 161
Score = 30.8 bits (68), Expect = 2.4
Identities = 15/24 (62%), Positives = 17/24 (70%)
Frame = +3
Query: 429 QQLPVQQQQQQQQQQQQYTRPTFP 452
QQ QQQQQQQQQQQQ ++ P
Sbjct: 90 QQQQQQQQQQQQQQQQQQSQQQQP 161
Score = 30.4 bits (67), Expect = 3.2
Identities = 16/31 (51%), Positives = 18/31 (57%)
Frame = +3
Query: 415 QPQVPVNAIAQQMKQQLPVQQQQQQQQQQQQ 445
Q + + I Q L QQQQQQQQQQQQ
Sbjct: 39 QQHMQMQQILLQRHAALQQQQQQQQQQQQQQ 131
Score = 29.6 bits (65), Expect = 5.4
Identities = 14/21 (66%), Positives = 15/21 (70%)
Frame = +3
Query: 425 QQMKQQLPVQQQQQQQQQQQQ 445
QQ +QQ QQQQQ QQQQ Q
Sbjct: 102 QQQQQQQQQQQQQQSQQQQPQ 164
>TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fragment),
partial (6%)
Length = 806
Score = 33.9 bits (76), Expect = 0.29
Identities = 27/70 (38%), Positives = 33/70 (46%), Gaps = 2/70 (2%)
Frame = +2
Query: 403 PPFYHQYPLPPGQPQVPVNAIAQQMKQQLP-VQQQQQQQQQQQQYTRPTFPPIPMLY-AE 460
P Q L PQ+ + QQ +QQLP VQQQQ Q QQQQ + + L +
Sbjct: 11 PQLQQQQQLQQQPPQL--TQLQQQQQQQLPQVQQQQLSQMQQQQLPQLQQQQLSQLQPQQ 184
Query: 461 LLPTLLQRGH 470
LP L Q H
Sbjct: 185 QLPQLQQLQH 214
Score = 32.3 bits (72), Expect = 0.83
Identities = 20/53 (37%), Positives = 26/53 (48%)
Frame = +2
Query: 416 PQVPVNAIAQQMKQQLPVQQQQQQQQQQQQYTRPTFPPIPMLYAELLPTLLQR 468
PQV ++Q +QQLP QQQQ Q Q Q P + L + LP Q+
Sbjct: 92 PQVQQQQLSQMQQQQLPQLQQQQLSQLQPQ---QQLPQLQQLQHQQLPQQQQQ 241
Score = 29.6 bits (65), Expect = 5.4
Identities = 22/56 (39%), Positives = 30/56 (53%), Gaps = 7/56 (12%)
Frame = +2
Query: 417 QVPVNAIAQQMKQQLP----VQQQQQQ---QQQQQQYTRPTFPPIPMLYAELLPTL 465
Q+P QQ++QQ P +QQQQQQ Q QQQQ ++ +P L + L L
Sbjct: 5 QLPQLQQQQQLQQQPPQLTQLQQQQQQQLPQVQQQQLSQMQQQQLPQLQQQQLSQL 172
>AV408190
Length = 382
Score = 32.7 bits (73), Expect = 0.64
Identities = 27/97 (27%), Positives = 43/97 (43%), Gaps = 4/97 (4%)
Frame = +1
Query: 382 INAAQMPPSYPYMPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQQQQQ 441
+ A+ PPS P P + +F H PLPP Q ++ Q+ P+ ++ +++
Sbjct: 100 LQQAEAPPS-PSKPLLRRHYF----HLLPLPPWQDRLHRRCHWPQVHLPTPLDRRPRRRL 264
Query: 442 QQQQYTRPTF----PPIPMLYAELLPTLLQRGHCTTR 474
++ P PP P L+ LLP L R H R
Sbjct: 265 LSLRHGHPKGQRRPPPFPQLH--LLPRRLPRRHVPRR 369
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,816,580
Number of Sequences: 28460
Number of extensions: 544155
Number of successful extensions: 3919
Number of sequences better than 10.0: 83
Number of HSP's better than 10.0 without gapping: 3224
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3605
length of query: 2244
length of database: 4,897,600
effective HSP length: 105
effective length of query: 2139
effective length of database: 1,909,300
effective search space: 4083992700
effective search space used: 4083992700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146556.9


