

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146553.6 + phase: 0 /pseudo
(799 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
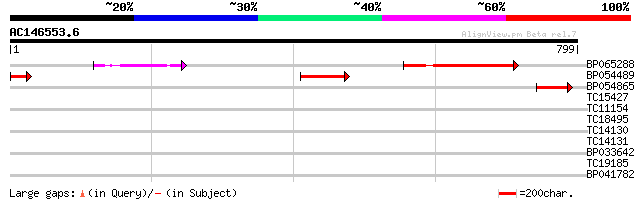
Score E
Sequences producing significant alignments: (bits) Value
BP065288 131 5e-31
BP054489 68 6e-12
BP054865 66 2e-11
TC15427 31 0.67
TC11154 weakly similar to UP|Q9LQ75 (Q9LQ75) T1N6.22 protein, pa... 30 2.0
TC18495 weakly similar to UP|Q40077 (Q40077) Seed imbibition pro... 29 2.6
TC14130 similar to UP|ATPG_PEA (P28552) ATP synthase gamma chain... 29 2.6
TC14131 similar to UP|ATPG_PEA (P28552) ATP synthase gamma chain... 29 2.6
BP033642 29 3.3
TC19185 similar to UP|Q9LJF4 (Q9LJF4) Emb|CAB87904.1, partial (14%) 28 4.4
BP041782 28 7.4
>BP065288
Length = 543
Score = 131 bits (329), Expect = 5e-31
Identities = 79/162 (48%), Positives = 102/162 (62%)
Frame = +3
Query: 555 WESILQTLKPGSKVTVLTNGPLTNLAKVVSIKNISSIIQA*QFQNVSGGLCNGRTYQQE* 614
WES+LQT +PGSK+TVLTNGPLTN + +S K+ + N GGLC+ T+Q E
Sbjct: 42 WESVLQTTEPGSKITVLTNGPLTNWQRCISRKH--------KI*N-QGGLCSWGTHQPEC 194
Query: 615 Q*QRKCIFCSFQQVCRIQYVLGSFSSKDSVSIRS*HHTHSTRYPAQSKFIFKYFKLA*SD 674
Q* RK +F SFQ +CRIQ+V SF +DSV I S*H+ S+ AQSKFIF + +
Sbjct: 195 Q**RKHLFSSFQPICRIQHVPRSFGCQDSV*I*S*HNIDSS*CTAQSKFIFNRHRSVTYE 374
Query: 675 RKNTRSSVFQASSVKATPFEENPPQISPYGAVVLANGHSSLL 716
N+ +FQAS+VK PFE NP Q+S YG ++ N S L
Sbjct: 375 INNS*GCIFQASAVKPIPFEANP*QVSSYGHILGGNSRCSNL 500
Score = 51.6 bits (122), Expect = 5e-07
Identities = 36/131 (27%), Positives = 61/131 (46%)
Frame = +1
Query: 119 HLKKNVEHVYIMGGVIRSKTCCTKNASSSCIPSKCGDTGNVLTNYNANPYAEYNIFGDPF 178
+++ ++ VY++GG I ++NA+ D GN+ + +N YAE+N+F DP
Sbjct: 136 NIRSRIKEVYVVGGHI------SQNAN---------DRGNIFS-VPSNQYAEFNMFLDPL 267
Query: 179 AAYKVIHSGIPITLVPLDATNTIPISEEFFDEFEKSQDTYEAQYCFKSLKMAHDTWFDNQ 238
AA V S + ITL+PL A + + T EA + S ++ + Q
Sbjct: 268 AAKTVFESEVNITLIPLSAQRKVSSFSTAIGQLRTKSTTPEAVF---SKRLLSSLYHLKQ 438
Query: 239 FYTSYFMWDSF 249
+ Y D+F
Sbjct: 439 IHNKYHHMDTF 471
>BP054489
Length = 487
Score = 67.8 bits (164), Expect = 6e-12
Identities = 31/31 (100%), Positives = 31/31 (100%)
Frame = +1
Query: 1 MMGRDDIAVGVGGEGGILPNGTILPNVGGYL 31
MMGRDDIAVGVGGEGGILPNGTILPNVGGYL
Sbjct: 394 MMGRDDIAVGVGGEGGILPNGTILPNVGGYL 486
Score = 60.5 bits (145), Expect = 1e-09
Identities = 29/69 (42%), Positives = 43/69 (62%), Gaps = 1/69 (1%)
Frame = +1
Query: 411 LGKPVVFDMDMSAGDFLALSYLLKVPVEVINLKAIIVSPTGWANAA-TIDVIYDLLHMMG 469
L + ++ D D+ DF AL YLLK+ L+ I ++ W +A ++ IYD+L+MMG
Sbjct: 223 LPRRILMDTDVDTDDFFALLYLLKLNRSQFQLEGITINANSWLDAGHAVNQIYDILYMMG 402
Query: 470 RDDIKVGIG 478
RDDI VG+G
Sbjct: 403 RDDIAVGVG 429
>BP054865
Length = 461
Score = 65.9 bits (159), Expect = 2e-11
Identities = 31/51 (60%), Positives = 37/51 (71%)
Frame = -3
Query: 743 EKYGKLVRILRHVDAKTYHEIYAKRLGDPNQSAKVGSFKEQKRKWSHPHDR 793
EK+GKLVRIL V A Y+ + A +LGD NQSAKVGSF+E RKW H D+
Sbjct: 447 EKHGKLVRILSRVSALAYYNLLADKLGDHNQSAKVGSFEEPPRKWRHSPDK 295
>TC15427
Length = 1079
Score = 31.2 bits (69), Expect = 0.67
Identities = 21/81 (25%), Positives = 34/81 (41%)
Frame = +2
Query: 97 GPITLIVTGAHTNLAIFLMNNPHLKKNVEHVYIMGGVIRSKTCCTKNASSSCIPSKCGDT 156
G +T++ G TN+A+ + + V+ + ++GG +
Sbjct: 5 GEVTVLALGPLTNVALAVKRDSSFASKVKRIVVLGGSFFA-------------------L 127
Query: 157 GNVLTNYNANPYAEYNIFGDP 177
GNV NP AE NI+GDP
Sbjct: 128 GNV------NPAAEANIYGDP 172
>TC11154 weakly similar to UP|Q9LQ75 (Q9LQ75) T1N6.22 protein, partial (68%)
Length = 833
Score = 29.6 bits (65), Expect = 2.0
Identities = 17/42 (40%), Positives = 24/42 (56%)
Frame = +3
Query: 549 PQAMEIWESILQTLKPGSKVTVLTNGPLTNLAKVVSIKNISS 590
PQ E+ E LQT G+K+T + PL +L+ I N+SS
Sbjct: 288 PQTYELAEECLQTNYYGAKITTESLLPLLHLSDSPRIVNVSS 413
>TC18495 weakly similar to UP|Q40077 (Q40077) Seed imbibition protein,
partial (5%)
Length = 534
Score = 29.3 bits (64), Expect = 2.6
Identities = 12/41 (29%), Positives = 19/41 (46%)
Frame = -3
Query: 127 VYIMGGVIRSKTCCTKNASSSCIPSKCGDTGNVLTNYNANP 167
+Y GG + + CCT++ + I +KC G N P
Sbjct: 370 MYNSGGAVEALDCCTRDVAECRIKTKCRGCGRFGAYSNVRP 248
>TC14130 similar to UP|ATPG_PEA (P28552) ATP synthase gamma chain,
chloroplast precursor , partial (94%)
Length = 1527
Score = 29.3 bits (64), Expect = 2.6
Identities = 15/47 (31%), Positives = 23/47 (48%)
Frame = +3
Query: 261 SNRKKGENEFAEMEYINITVITSNKPYGISDGSNPLFDGLKVPKFNL 307
SN+ M N+T++ S+KP +SD SN F + + F L
Sbjct: 60 SNQNSIATPTTTMSCSNLTMLVSSKPSSLSDASNLSFRSIHLNPFQL 200
>TC14131 similar to UP|ATPG_PEA (P28552) ATP synthase gamma chain,
chloroplast precursor , partial (40%)
Length = 655
Score = 29.3 bits (64), Expect = 2.6
Identities = 15/47 (31%), Positives = 23/47 (48%)
Frame = +3
Query: 261 SNRKKGENEFAEMEYINITVITSNKPYGISDGSNPLFDGLKVPKFNL 307
SN+ M N+T++ S+KP +SD SN F + + F L
Sbjct: 27 SNQNSIATPTTTMSCSNLTMLVSSKPSSLSDASNLSFRSIHLNPFQL 167
>BP033642
Length = 448
Score = 28.9 bits (63), Expect = 3.3
Identities = 13/29 (44%), Positives = 19/29 (64%)
Frame = -3
Query: 24 LPNVGGYLPIIEQGMTTAGYCRYRQAIPV 52
LP +GG+LP I + +TT G +R AI +
Sbjct: 362 LPAMGGFLPAIWRFITTTGGSTHRSAIAI 276
>TC19185 similar to UP|Q9LJF4 (Q9LJF4) Emb|CAB87904.1, partial (14%)
Length = 533
Score = 28.5 bits (62), Expect = 4.4
Identities = 12/31 (38%), Positives = 19/31 (60%)
Frame = +2
Query: 545 ELRQPQAMEIWESILQTLKPGSKVTVLTNGP 575
E+ +ME WE++++ + GS TVL N P
Sbjct: 422 EVAAAVSMECWENVVEDIDDGSSGTVLANLP 514
>BP041782
Length = 538
Score = 27.7 bits (60), Expect = 7.4
Identities = 14/26 (53%), Positives = 18/26 (68%)
Frame = -1
Query: 664 IFKYFKLA*SDRKNTRSSVFQASSVK 689
IF+ F LA*S KNT ++VF SV+
Sbjct: 520 IFRIFVLA*SPEKNTTNTVFSIISVE 443
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,132,459
Number of Sequences: 28460
Number of extensions: 177070
Number of successful extensions: 805
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 794
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 804
length of query: 799
length of database: 4,897,600
effective HSP length: 98
effective length of query: 701
effective length of database: 2,108,520
effective search space: 1478072520
effective search space used: 1478072520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146553.6


