

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146553.2 + phase: 0 /pseudo
(654 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
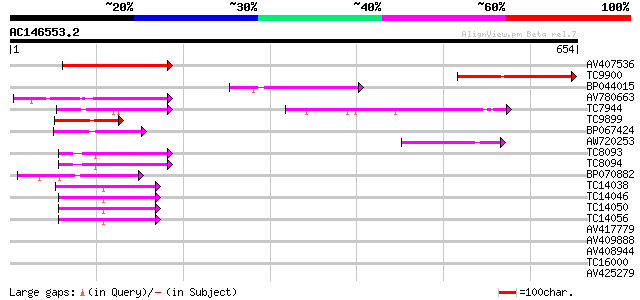
Score E
Sequences producing significant alignments: (bits) Value
AV407536 202 1e-52
TC9900 homologue to UP|EFGC_SOYBN (P34811) Elongation factor G, ... 132 2e-31
BP044015 113 1e-25
AV780663 87 1e-17
TC7944 homologue to UP|Q9ASR1 (Q9ASR1) At1g56070/T6H22_13 (Elong... 77 1e-14
TC9899 homologue to PIR|S35701|S35701 translation elongation fac... 69 2e-12
BP067424 66 2e-11
AW720253 62 3e-10
TC8093 homologue to UP|EFTU_TOBAC (P41342) Elongation factor Tu,... 54 6e-08
TC8094 homologue to UP|EFTU_PEA (O24310) Elongation factor Tu, c... 54 1e-07
BP070882 52 3e-07
TC14038 homologue to UP|Q9LN13 (Q9LN13) T6D22.2, partial (47%) 44 8e-05
TC14046 homologue to UP|P93769 (P93769) Elongation factor-1 alph... 44 1e-04
TC14050 homologue to UP|EF1A_TOBAC (P43643) Elongation factor 1-... 43 1e-04
TC14056 homologue to UP|P93769 (P93769) Elongation factor-1 alph... 43 1e-04
AV417779 35 0.038
AV409888 33 0.14
AV408944 30 0.94
TC16000 similar to UP|Q9FQD5 (Q9FQD5) Glutathione S-transferase ... 30 0.94
AV425279 29 2.7
>AV407536
Length = 382
Score = 202 bits (514), Expect = 1e-52
Identities = 100/126 (79%), Positives = 110/126 (86%)
Frame = +2
Query: 62 AHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITIKSAATYCNWK 121
AHIDSGKTTLTE +LFY G+IH ++EVR +DG+G KMD LE GITI+SAATYC+WK
Sbjct: 5 AHIDSGKTTLTERVLFYAGRIHEIHEVRGRDGVGAKMDSMDLEREKGITIQSAATYCSWK 184
Query: 122 GSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSITVDRQMRRYQVPRIAF 181
KI IIDTPGHVDFTIEVERALRVLDGA+LVLCSVGGVQ QSITVDRQMRRY VPR+AF
Sbjct: 185 DCKINIIDTPGHVDFTIEVERALRVLDGAILVLCSVGGVQSQSITVDRQMRRYDVPRLAF 364
Query: 182 INKLDR 187
INKLDR
Sbjct: 365 INKLDR 382
>TC9900 homologue to UP|EFGC_SOYBN (P34811) Elongation factor G,
chloroplast precursor (EF-G), partial (34%)
Length = 1284
Score = 132 bits (332), Expect = 2e-31
Identities = 65/137 (47%), Positives = 95/137 (68%)
Frame = +1
Query: 517 VDATVGKPRVNFRETVTQRADFDYLHKKQSGGQGQYGRVIGYIEPLPAGSGTKFEFDNML 576
V+A VG P+VN+RE++++ ++ Y+HKKQSGGQGQ+ + EP+ GSG +EF + +
Sbjct: 286 VEANVGAPQVNYRESISKVSEVKYVHKKQSGGQGQFADITVRFEPMEPGSG--YEFKSEI 459
Query: 577 VGQAIPSNFFPAIEKGFKEAANSGALIGHPVQNLRVVLTDGAAHDVDSSELAFKLASIYA 636
G A+P + P + KG +E ++G L G PV ++R VLTDG+ HDVDSS LAF+LA+ A
Sbjct: 460 KGGAVPREYIPGVMKGLEECMSNGVLAGFPVVDVRAVLTDGSYHDVDSSVLAFQLAARGA 639
Query: 637 FRECYTASRPVILEPVM 653
FRE + P +LEP+M
Sbjct: 640 FREGVRKAGPRMLEPIM 690
>BP044015
Length = 480
Score = 113 bits (282), Expect = 1e-25
Identities = 64/161 (39%), Positives = 93/161 (57%), Gaps = 6/161 (3%)
Frame = +3
Query: 254 LVAEKRRELIETVSEVDDVLAEAFLS----DDENISAADLEGAIRRATIARKFIPVFMGS 309
L + R +++E + E DD + E +L DDE I IR+ TI+ F+PV GS
Sbjct: 6 LAQDYRSQMMENIVEFDDQVMENYLEGIEPDDETIKKL-----IRKGTISASFVPVMCGS 170
Query: 310 AVKNTGVQPLLDGVVSYLPCPIEVSNY-ALDQSKNEEKVQLTGSPDGPLVALAFKLEQTK 368
A KN GVQPLLD VV YLP P+++ D E ++ + S D P LAFK+
Sbjct: 171 AFKNKGVQPLLDAVVDYLPSPLDLPPMKGTDPENPEVTMERSASDDEPFSGLAFKIMSDP 350
Query: 369 F-GQLTYLRVYEGVIRKGDFIVNVSTGKKIKVPRLVQMHSN 408
F G LT++RVY G + G +++N + GKK ++ RL++MH+N
Sbjct: 351 FVGTLTFVRVYAGKLSAGSYVLNANKGKKERIGRLLEMHAN 473
>AV780663
Length = 568
Score = 86.7 bits (213), Expect = 1e-17
Identities = 65/190 (34%), Positives = 96/190 (50%), Gaps = 6/190 (3%)
Frame = +1
Query: 5 GYALCSSTLTTEGSLIAGT------FHIRHFSAGNVARATAATIDKDPWWKESMEKVRNI 58
G L S +T SL + + F R FSA + A A +AT P ++RN+
Sbjct: 7 GSLLLRSLWSTRKSLSSSSPSPPSRFFSRPFSAASAAAAASATA---PAGSLDPNRLRNV 177
Query: 59 GISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITIKSAATYC 118
+ AH+D GKTTL + IL G + +E MD LE GITI S T
Sbjct: 178 AVIAHVDHGKTTLMDRILRQCGA-DIPHE--------RAMDSNSLERERGITIASKVTTV 330
Query: 119 NWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSITVDRQMRRYQVPR 178
+WK +++ ++DTPGH D EVER + +++GAV V+ + G Q+ V + +Y +
Sbjct: 331 SWKENELNMVDTPGHADLGGEVERVVGMVEGAV*VVDAGEGPLAQTKLVLAKALKYGLRP 510
Query: 179 IAFINKLDRP 188
+ NK+DRP
Sbjct: 511 LLLKNKVDRP 540
>TC7944 homologue to UP|Q9ASR1 (Q9ASR1) At1g56070/T6H22_13 (Elongation
factor EF-2), complete
Length = 2971
Score = 76.6 bits (187), Expect = 1e-14
Identities = 53/149 (35%), Positives = 79/149 (52%), Gaps = 16/149 (10%)
Frame = +1
Query: 55 VRNIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITIKSA 114
+RN+ + AH+D GK+TLT+ ++ G I + G D + E GITIKS
Sbjct: 262 IRNMSVIAHVDHGKSTLTDSLVAAAGII-----AQEVAGDVRMTDTRADEAERGITIKST 426
Query: 115 ATYC----------NWKGSK------ITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVG 158
++KG + I +ID+PGHVDF+ EV ALR+ DGA++V+ V
Sbjct: 427 GISLYYEMTDVALKSFKGERMGNEYLINLIDSPGHVDFSSEVTAALRITDGALVVVDCVE 606
Query: 159 GVQCQSITVDRQMRRYQVPRIAFINKLDR 187
GV Q+ TV RQ ++ + +NK+DR
Sbjct: 607 GVCVQTETVLRQALGERIKPVLTVNKMDR 693
Score = 63.9 bits (154), Expect = 8e-11
Identities = 71/284 (25%), Positives = 122/284 (42%), Gaps = 24/284 (8%)
Frame = +1
Query: 319 LLDGVVSYLPCPIEVSNYALDQ------SKNEEKVQLTGSPDGPLVALAFKL--EQTKFG 370
LL+ ++ +LP P Y ++ P+GPL+ K+ K
Sbjct: 1201 LLEMMIFHLPSPSTAQRYRVENLYEGPLDDQYAAAIRNCDPEGPLMLYVSKMIPASDKGR 1380
Query: 371 QLTYLRVYEGVIRKGDFI----VNVSTGKKI-----KVPRLVQMHSNEMNDIEEAHAGQI 421
+ RV+ G + G + N G+K V R V + +E+ G
Sbjct: 1381 FFAFGRVFSGKVSTGLKVRIMGPNYVPGEKKDLYTKSVQRTVIWMGKKQETVEDVPCGNT 1560
Query: 422 VAVFGVDCASSDTFTDGSVKYT----MTSMNVP-EPVMSLAVQPVSKDSGGKFSKALNRF 476
VA+ G+D + T + K T + +M PV+ +AVQ K + L R
Sbjct: 1561 VALVGLDQFITKNATLTNEKETDAHPIRAMKFSVSPVVRVAVQCKVASDLPKLVEGLKRL 1740
Query: 477 QREDPTFRVSLDPESGQTIISGMGELHLDIYVKRIKMEY--GVDATVGKPRVNFRETVTQ 534
+ DP +++ ESG+ I++G GELHL+I +K ++ ++ G + P V+FRETV +
Sbjct: 1741 AKSDPMVVCTIE-ESGEHIVAGAGELHLEICLKDLQDDFMGGAEIVKSDPVVSFRETVLE 1917
Query: 535 RADFDYLHKKQSGGQGQYGRVIGYIEPLPAGSGTKFEFDNMLVG 578
R+ + K + ++ R+ Y+E P G D+ +G
Sbjct: 1918 RSCRTVMSKSPN----KHNRL--YMEARPLEDGLAEAIDDGKIG 2031
>TC9899 homologue to PIR|S35701|S35701 translation elongation factor EF-G,
chloroplast - soybean {Glycine max;} , partial (15%)
Length = 624
Score = 69.3 bits (168), Expect = 2e-12
Identities = 41/80 (51%), Positives = 49/80 (61%)
Frame = +1
Query: 52 MEKVRNIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITI 111
++ RNIGI AHID+GKTT TE ILFYTG+ + + EV MD+ E GITI
Sbjct: 394 LKDYRNIGIMAHIDAGKTTTTERILFYTGRNYKIGEVHEGTA---TMDWMEQEQERGITI 564
Query: 112 KSAATYCNWKGSKITIIDTP 131
SAAT W +I IIDTP
Sbjct: 565 TSAATTTFWNKHRIHIIDTP 624
>BP067424
Length = 473
Score = 65.9 bits (159), Expect = 2e-11
Identities = 40/109 (36%), Positives = 61/109 (55%), Gaps = 2/109 (1%)
Frame = +1
Query: 51 SMEKVRNIGISAHIDSGKTTLTEWILFYT--GKIHLMYEVRSKDGMGPKMDFKPLEIIMG 108
S K+RNI I AH+D GKTTL + ++ G +H R + MD+ E
Sbjct: 157 SRHKIRNICILAHVDHGKTTLADQLIAVASGGMVHPKVAGRVRF-----MDYLDEEQRRA 321
Query: 109 ITIKSAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSV 157
IT+KS++ + + +ID+PGH+DF EV A R+ DGA+L++ +V
Sbjct: 322 ITMKSSSIALRYNHHTVNLIDSPGHIDFCGEVSTAARLSDGALLLVDAV 468
>AW720253
Length = 481
Score = 62.0 bits (149), Expect = 3e-10
Identities = 39/122 (31%), Positives = 66/122 (53%), Gaps = 2/122 (1%)
Frame = +3
Query: 452 PVMSLAVQPVSKDSGGKFSKALNRFQREDPTFRVSLDPESGQTIISGMGELHLDIYVKRI 511
PV+ +AV+P + K + L R + DP + ++ ESG+ I++G GELHL+I +K +
Sbjct: 66 PVVRVAVEPKNPADLPKLVEGLKRLSKSDPCVQCFME-ESGEHIVAGAGELHLEICLKDL 242
Query: 512 KMEY--GVDATVGKPRVNFRETVTQRADFDYLHKKQSGGQGQYGRVIGYIEPLPAGSGTK 569
+ ++ GV+ + P V++RETVT +D H S ++ R+ EP+ G
Sbjct: 243 QDDFMNGVELKITDPIVSYRETVTAPSD----HVVLSKSPNKHNRIYLKAEPMQDGLAEA 410
Query: 570 FE 571
E
Sbjct: 411 IE 416
>TC8093 homologue to UP|EFTU_TOBAC (P41342) Elongation factor Tu,
chloroplast precursor (EF-Tu), partial (58%)
Length = 1146
Score = 54.3 bits (129), Expect = 6e-08
Identities = 41/136 (30%), Positives = 58/136 (42%), Gaps = 5/136 (3%)
Frame = +1
Query: 57 NIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPK----MDFKPLEIIMGITIK 112
NIG H+D GKTTLT L + S PK +D P E GITI
Sbjct: 358 NIGTIGHVDHGKTTLTA---------ALTMALASLGNSAPKKYDEIDAAPEERARGITIN 510
Query: 113 SAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSITVDRQMR 172
+A + +D PGH D+ + +DGA+LV+ G Q+ +
Sbjct: 511 TATVEYETENRHYAHVDCPGHADYVKNMITGAAQMDGAILVVSGADGPMPQTKEHILLAK 690
Query: 173 RYQVPR-IAFINKLDR 187
+ VP + F+NK D+
Sbjct: 691 QVGVPNMVVFLNKQDQ 738
>TC8094 homologue to UP|EFTU_PEA (O24310) Elongation factor Tu, chloroplast
precursor (EF-Tu), partial (60%)
Length = 1092
Score = 53.5 bits (127), Expect = 1e-07
Identities = 41/136 (30%), Positives = 58/136 (42%), Gaps = 5/136 (3%)
Frame = +1
Query: 57 NIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPK----MDFKPLEIIMGITIK 112
NIG H+D GKTTLT L + S PK +D P E GITI
Sbjct: 331 NIGTIGHVDHGKTTLTA---------ALTMALASLGNSAPKKYDEIDAAPEERARGITIN 483
Query: 113 SAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSITVDRQMR 172
+A + +D PGH D+ + +DGA+LV+ G Q+ +
Sbjct: 484 TATVEYETETRHYAHVDCPGHADYVKNMITGAAQMDGAILVVSGADGPMPQTKEHILLAK 663
Query: 173 RYQVPR-IAFINKLDR 187
+ VP + F+NK D+
Sbjct: 664 QVGVPNVVVFLNKQDQ 711
>BP070882
Length = 496
Score = 52.0 bits (123), Expect = 3e-07
Identities = 44/156 (28%), Positives = 66/156 (42%), Gaps = 11/156 (7%)
Frame = +2
Query: 10 SSTLTTEGSLIAGTFHIRHFSAGN------VARATAATIDKDPWWKESMEKVR-----NI 58
SST S + + HI H S + + ++ +PWW+ R N+
Sbjct: 35 SSTCRRHLSYLPFSSHILHSSLSSSPLFNDTPSSFSSPSSSNPWWRSMATFTRTKPHVNV 214
Query: 59 GISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITIKSAATYC 118
G H+D GKTTLT I K+ L E ++K ++D P E GITI +A
Sbjct: 215 GTMGHVDHGKTTLTAAIT----KV-LADEGKAKAIAFDEIDKAPEEKKRGITIATAHVEY 379
Query: 119 NWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVL 154
+D PGH D+ + +DG +LV+
Sbjct: 380 ETVKRHYAHVDCPGHADYVKNMITGAAQMDGGILVV 487
>TC14038 homologue to UP|Q9LN13 (Q9LN13) T6D22.2, partial (47%)
Length = 1844
Score = 43.9 bits (102), Expect = 8e-05
Identities = 40/133 (30%), Positives = 57/133 (42%), Gaps = 11/133 (8%)
Frame = +3
Query: 53 EKVR-NIGISAHIDSGKTTLTEWILFYTGKI--HLMYEVRSKDGMGPKMDFKPLEII--- 106
EKV NI + H+DSGK+T T +++ G I ++ + K FK ++
Sbjct: 99 EKVHINIVVIGHVDSGKSTTTGHLIYKLGGIDKRVIERFEKEAAEMNKRSFKYAWVLDKL 278
Query: 107 -----MGITIKSAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQ 161
GITI A T+ID PGH DF + D AVL++ S G
Sbjct: 279 KAERERGITIDIALWKFETTKYYCTVIDAPGHRDFIKNMITGTSQADCAVLIIDSTTGGF 458
Query: 162 CQSITVDRQMRRY 174
I+ D Q R +
Sbjct: 459 EAGISKDGQTREH 497
>TC14046 homologue to UP|P93769 (P93769) Elongation factor-1 alpha, partial
(84%)
Length = 1506
Score = 43.5 bits (101), Expect = 1e-04
Identities = 37/128 (28%), Positives = 54/128 (41%), Gaps = 10/128 (7%)
Frame = +1
Query: 57 NIGISAHIDSGKTTLTEWILFYTGKI--HLMYEVRSKDGMGPKMDFKPLEII-------- 106
NI + H+DSGK+T T +++ G I ++ + K FK ++
Sbjct: 106 NIVVIGHVDSGKSTTTGHLIYKLGGIDKRVIERFEKEAAEMNKRSFKYAWVLDKLKAERE 285
Query: 107 MGITIKSAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSIT 166
GITI A T+ID PGH DF + D AVL++ S G I+
Sbjct: 286 RGITIDIALWKFETNKYYCTVIDAPGHRDFIKNMITGTSQADCAVLIIDSTTGGFEAGIS 465
Query: 167 VDRQMRRY 174
D Q R +
Sbjct: 466 KDGQTREH 489
>TC14050 homologue to UP|EF1A_TOBAC (P43643) Elongation factor 1-alpha
(EF-1-alpha) (Vitronectin-like adhesion protein 1)
(PVN1), partial (38%)
Length = 583
Score = 43.1 bits (100), Expect = 1e-04
Identities = 37/128 (28%), Positives = 54/128 (41%), Gaps = 10/128 (7%)
Frame = +2
Query: 57 NIGISAHIDSGKTTLTEWILFYTGKI--HLMYEVRSKDGMGPKMDFKPLEII-------- 106
NI + H+DSGK+T T +++ G I ++ + K FK ++
Sbjct: 95 NIVVIGHVDSGKSTTTGHLIYKLGGIDKRVIERFEKEAAEMNKRSFKYAWVLDKLKAERE 274
Query: 107 MGITIKSAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSIT 166
GITI A T+ID PGH DF + D AVL++ S G I+
Sbjct: 275 RGITIDIALWKFETTKYYCTVIDAPGHRDFIKNMITGTSQADCAVLIIDSTTGGFEAGIS 454
Query: 167 VDRQMRRY 174
D Q R +
Sbjct: 455 KDGQTREH 478
>TC14056 homologue to UP|P93769 (P93769) Elongation factor-1 alpha, complete
Length = 1897
Score = 43.1 bits (100), Expect = 1e-04
Identities = 37/128 (28%), Positives = 54/128 (41%), Gaps = 10/128 (7%)
Frame = +1
Query: 57 NIGISAHIDSGKTTLTEWILFYTGKI--HLMYEVRSKDGMGPKMDFKPLEII-------- 106
NI + H+DSGK+T T +++ G I ++ + K FK ++
Sbjct: 133 NIVVIGHVDSGKSTTTGHLIYKLGGIDKRVIERFEKEAAEMNKRSFKYAWVLDKLKAERE 312
Query: 107 MGITIKSAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSIT 166
GITI A T+ID PGH DF + D AVL++ S G I+
Sbjct: 313 RGITIDIALWKFETTKYYCTVIDAPGHRDFIKNMITGTSQADCAVLIIDSTTGGFEAGIS 492
Query: 167 VDRQMRRY 174
D Q R +
Sbjct: 493 KDGQTREH 516
>AV417779
Length = 401
Score = 35.0 bits (79), Expect = 0.038
Identities = 29/90 (32%), Positives = 41/90 (45%), Gaps = 10/90 (11%)
Frame = +1
Query: 57 NIGISAHIDSGKTTLTEWILFYTGKIHLM----YEVRSKD-GMGP-----KMDFKPLEII 106
+I + H+DSGK+T T +++ G I +E + + G G +D E
Sbjct: 103 SIVVIGHVDSGKSTTTGHLIYKCGGIEKRAIEKFEKEAAEMGKGSFKYAWVLDKLKAERE 282
Query: 107 MGITIKSAATYCNWKGSKITIIDTPGHVDF 136
GITI A + TIID PGH DF
Sbjct: 283 RGITIDIALWKFETEKYSFTIIDAPGHRDF 372
>AV409888
Length = 426
Score = 33.1 bits (74), Expect = 0.14
Identities = 19/47 (40%), Positives = 27/47 (57%)
Frame = +2
Query: 33 NVARATAATIDKDPWWKESMEKVRNIGISAHIDSGKTTLTEWILFYT 79
+V+ AT +K + + VRNI I AH+D GKTTL + +L T
Sbjct: 275 SVSEATEPKTEKKQFTRRG--DVRNIAIVAHVDHGKTTLVDAMLRQT 409
>AV408944
Length = 295
Score = 30.4 bits (67), Expect = 0.94
Identities = 12/28 (42%), Positives = 20/28 (70%)
Frame = +1
Query: 55 VRNIGISAHIDSGKTTLTEWILFYTGKI 82
+RN+ + AH+D GK+TLT+ ++ G I
Sbjct: 208 IRNMSVIAHVDHGKSTLTDSLVAAAGII 291
>TC16000 similar to UP|Q9FQD5 (Q9FQD5) Glutathione S-transferase GST 23 ,
partial (73%)
Length = 627
Score = 30.4 bits (67), Expect = 0.94
Identities = 24/76 (31%), Positives = 32/76 (41%)
Frame = +2
Query: 150 AVLVLCSVGGVQCQSITVDRQMRRYQVPRIAFINKLDRPGADPWKVITQARSKLRHHCAA 209
AVL+ C V G+ + I ++ R++ P IN + A I R KL A
Sbjct: 122 AVLIFCKVNGIDFEEIKIELSKRQHLSPEFHAINPFKKVPA-----IVDGRFKLFESHAI 286
Query: 210 LQVPIGLESDFKGVVD 225
L I L F GV D
Sbjct: 287 L---IYLACAFPGVAD 325
>AV425279
Length = 426
Score = 28.9 bits (63), Expect = 2.7
Identities = 16/52 (30%), Positives = 26/52 (49%)
Frame = -1
Query: 434 TFTDGSVKYTMTSMNVPEPVMSLAVQPVSKDSGGKFSKALNRFQREDPTFRV 485
+FTD + N ++S V +K+ GGK K++N +R DP F +
Sbjct: 411 SFTDKPGHTRLGRRNSGSMLVSSFVSTKAKNFGGKSFKSMNASERTDPQFEL 256
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,064,666
Number of Sequences: 28460
Number of extensions: 102288
Number of successful extensions: 453
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 435
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 437
length of query: 654
length of database: 4,897,600
effective HSP length: 96
effective length of query: 558
effective length of database: 2,165,440
effective search space: 1208315520
effective search space used: 1208315520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146553.2


