

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146553.10 - phase: 0
(301 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
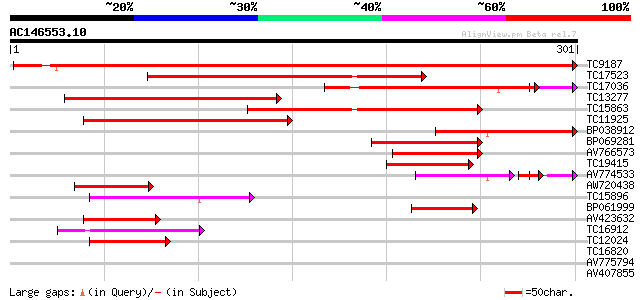
Sequences producing significant alignments: (bits) Value
TC9187 similar to UP|Q8LAN3 (Q8LAN3) Prolyl 4-hydroxylase alpha ... 487 e-138
TC17523 similar to UP|Q9FKX6 (Q9FKX6) Prolyl 4-hydroxylase, alph... 166 6e-42
TC17036 similar to UP|Q8VZD7 (Q8VZD7) AT3g28480/MFJ20_16, partia... 150 4e-40
TC13277 similar to UP|Q9LSI6 (Q9LSI6) Prolyl 4-hydroxylase alpha... 158 9e-40
TC15863 153 4e-38
TC11925 similar to UP|Q9FKX6 (Q9FKX6) Prolyl 4-hydroxylase, alph... 132 9e-32
BP038912 91 2e-19
BP069281 84 2e-17
AV766573 71 3e-13
TC19415 similar to UP|BAD07294 (BAD07294) Prolyl 4-hydroxylase, ... 60 6e-10
AV774533 46 8e-10
AW720438 59 1e-09
TC15896 similar to UP|BAD07294 (BAD07294) Prolyl 4-hydroxylase, ... 57 5e-09
BP061999 52 2e-07
AV423632 51 2e-07
TC16912 similar to UP|BAD07294 (BAD07294) Prolyl 4-hydroxylase, ... 51 3e-07
TC12024 50 4e-07
TC16820 similar to UP|Q8L8T9 (Q8L8T9) Prolyl 4-hydroxylase alpha... 39 0.001
AV775794 30 0.65
AV407855 27 5.5
>TC9187 similar to UP|Q8LAN3 (Q8LAN3) Prolyl 4-hydroxylase alpha
subunit-like protein, partial (89%)
Length = 1198
Score = 487 bits (1253), Expect = e-138
Identities = 235/302 (77%), Positives = 263/302 (86%), Gaps = 3/302 (0%)
Frame = +2
Query: 3 VICRVWCSIIVPLLLICKIHF--ALGSYAGT-SAIIDPTKVKQVSWKPRAFVYKGFLTDL 59
++ +W + +PL+ H+ SYAG+ S+II+P+KVKQVSWKPRAFVY+GFLT L
Sbjct: 62 IMSSLWFLLFLPLIF----HWNEVSSSYAGSASSIINPSKVKQVSWKPRAFVYEGFLTGL 229
Query: 60 ECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSGMFISKNKDAIVSGIEDKISSWTFL 119
ECDHLIS+AKSELKRSAVADNLSG+SKLSEVRTSSGMFISK KD IV+GIEDKIS+WTFL
Sbjct: 230 ECDHLISLAKSELKRSAVADNLSGDSKLSEVRTSSGMFISKKKDPIVAGIEDKISAWTFL 409
Query: 120 PKENGEDIQVLRYEHGQKYDPHYDYFADKVNIARGGHRVATVLMYLTNVTKGGETVFPNA 179
PKENGED+QVLRYEHGQKYDPHYDYF DKVNI RGGHR+ATVL+YLTNVT+GGETVFP A
Sbjct: 410 PKENGEDMQVLRYEHGQKYDPHYDYFTDKVNIVRGGHRMATVLLYLTNVTRGGETVFPVA 589
Query: 180 EESPRHKLSETDEDLSECGKKGVAVNPRRGDALLFCSLHPNAIPDTLSLHAGCPVIEGEK 239
EE PR + ET+ DLSEC KKG+AV PRRGDALLF SLH A PD SLHAGCPVIEGEK
Sbjct: 590 EEPPRRRGLETNSDLSECAKKGIAVKPRRGDALLFFSLHTTATPDPDSLHAGCPVIEGEK 769
Query: 240 WSATKWIHVDSFDKTVGAGGDCTDQHESCERWAALGECTKNPEYMVGTSGLPGYCRKSCK 299
WSATKWIHVDSFDKTVGAGGDC+DQHESC+RWA+LGECT NPEYMVG+S LPG CR+SCK
Sbjct: 770 WSATKWIHVDSFDKTVGAGGDCSDQHESCQRWASLGECTNNPEYMVGSSDLPGSCRRSCK 949
Query: 300 TC 301
C
Sbjct: 950 AC 955
>TC17523 similar to UP|Q9FKX6 (Q9FKX6) Prolyl 4-hydroxylase, alpha
subunit-like protein, partial (55%)
Length = 448
Score = 166 bits (419), Expect = 6e-42
Identities = 83/148 (56%), Positives = 108/148 (72%)
Frame = +1
Query: 74 RSAVADNLSGESKLSEVRTSSGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYE 133
RS V D+ +G+SK S VRTSSG F+ + +D IV IE KI+ +TF+P E+GE +QVL YE
Sbjct: 7 RSTVVDSETGKSKDSRVRTSSGTFLPRGRDKIVRNIEKKIADFTFIPVEHGEGLQVLHYE 186
Query: 134 HGQKYDPHYDYFADKVNIARGGHRVATVLMYLTNVTKGGETVFPNAEESPRHKLSETDED 193
GQKY+PHYDYF D+ N GG R+ATVLMYLT+V GGETVFP+A+ + +
Sbjct: 187 VGQKYEPHYDYFLDEFNTKNGGQRIATVLMYLTDVEXGGETVFPSAKGN--FXSVPWWNE 360
Query: 194 LSECGKKGVAVNPRRGDALLFCSLHPNA 221
LS+CGKKG+++NP GDALLF S++P A
Sbjct: 361 LSDCGKKGLSINPXXGDALLFWSMNPXA 444
>TC17036 similar to UP|Q8VZD7 (Q8VZD7) AT3g28480/MFJ20_16, partial (21%)
Length = 697
Score = 150 bits (380), Expect(2) = 4e-40
Identities = 75/117 (64%), Positives = 85/117 (72%), Gaps = 3/117 (2%)
Frame = +2
Query: 168 VTKGGETVFPNAEESPRHKLSET-DEDLSECGKKGVAVNPRRGDALLFCSLHPNAIPDTL 226
V KGGET+FP+AE KLS+ DE SEC KG AV PR+GDALLF SLH NA D+
Sbjct: 2 VGKGGETIFPHAES----KLSQPKDESWSECAHKGYAVKPRKGDALLFFSLHLNATTDSN 169
Query: 227 SLHAGCPVIEGEKWSATKWIHVDSFDKTVGAG--GDCTDQHESCERWAALGECTKNP 281
SLH CPVIEGEKWSATKWIHV F+K + GDCTD++E+C RWA LGEC KNP
Sbjct: 170 SLHGSCPVIEGEKWSATKWIHVSDFEKAIKPDDHGDCTDENENCSRWAKLGECVKNP 340
Score = 30.4 bits (67), Expect(2) = 4e-40
Identities = 11/25 (44%), Positives = 14/25 (56%)
Frame = +1
Query: 277 CTKNPEYMVGTSGLPGYCRKSCKTC 301
C + M+G G+ GYC KSC C
Sbjct: 328 CEEPTLTMIGGKGVKGYCMKSCNVC 402
>TC13277 similar to UP|Q9LSI6 (Q9LSI6) Prolyl 4-hydroxylase alpha
subunit-like protein, partial (34%)
Length = 582
Score = 158 bits (400), Expect = 9e-40
Identities = 72/115 (62%), Positives = 95/115 (82%)
Frame = +1
Query: 30 GTSAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSE 89
G+S DPT+V Q+SW PRAF++ GFL+D ECDHL+++AK +L+ S VADN SG+S SE
Sbjct: 238 GSSVKFDPTRVTQLSWSPRAFLHTGFLSDKECDHLVNLAKDKLEVSMVADNESGKSIKSE 417
Query: 90 VRTSSGMFISKNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDY 144
VRTSSGMF++K +D +V+ IE +I++WTFLP ENGE IQ+L YE+GQKY+PHYDY
Sbjct: 418 VRTSSGMFLNKAQDEVVADIEARIATWTFLPIENGESIQILHYENGQKYEPHYDY 582
>TC15863
Length = 607
Score = 153 bits (386), Expect = 4e-38
Identities = 71/125 (56%), Positives = 91/125 (72%)
Frame = +2
Query: 127 IQVLRYEHGQKYDPHYDYFADKVNIARGGHRVATVLMYLTNVTKGGETVFPNAEESPRHK 186
+QVL YE GQKY+PHYDYF D+ N GG R+ATVLMYL++V +GGET+FP A+ +
Sbjct: 5 LQVLHYEVGQKYEPHYDYFLDEFNTKNGGQRIATVLMYLSDVEEGGETIFPAAKAN--FS 178
Query: 187 LSETDEDLSECGKKGVAVNPRRGDALLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWI 246
DLS C KKG++V P+RGDALLF S+ P+A D SLH GCPVI G KWS+TKW+
Sbjct: 179 SVPWYNDLSVCAKKGLSVKPKRGDALLFWSIRPDATLDPSSLHGGCPVIRGNKWSSTKWM 358
Query: 247 HVDSF 251
H++ +
Sbjct: 359 HLEEY 373
>TC11925 similar to UP|Q9FKX6 (Q9FKX6) Prolyl 4-hydroxylase, alpha
subunit-like protein, partial (43%)
Length = 613
Score = 132 bits (331), Expect = 9e-32
Identities = 60/111 (54%), Positives = 84/111 (75%)
Frame = +1
Query: 40 VKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGESKLSEVRTSSGMFIS 99
V+ +SW+PRAF+Y FLT EC++LI+IAK +++S V D+ +G+SK S RTSSG F++
Sbjct: 277 VELISWEPRAFLYHNFLTKEECEYLINIAKPHMRKSTVVDSETGKSKDSRARTSSGTFLA 456
Query: 100 KNKDAIVSGIEDKISSWTFLPKENGEDIQVLRYEHGQKYDPHYDYFADKVN 150
+ +D IV IE +I+ +TF+P E+GE +QVL YE GQKYD HYDYF D+ N
Sbjct: 457 RGRDKIVRNIEKRIADYTFIPIEHGEGLQVLHYEVGQKYDSHYDYFMDEFN 609
>BP038912
Length = 481
Score = 90.9 bits (224), Expect = 2e-19
Identities = 38/79 (48%), Positives = 51/79 (64%), Gaps = 4/79 (5%)
Frame = -2
Query: 227 SLHAGCPVIEGEKWSATKWIHVDSFD----KTVGAGGDCTDQHESCERWAALGECTKNPE 282
S HA CPV++G+ WSA K+ + S T+ GG+CTD+ +SC WAA+GEC +NP
Sbjct: 477 SYHARCPVLKGDMWSAIKFFYARSISGGKLSTISDGGECTDEDDSCPAWAAMGECQRNPV 298
Query: 283 YMVGTSGLPGYCRKSCKTC 301
YM+G+ G CRKSC C
Sbjct: 297 YMIGSPDYYGTCRKSCNAC 241
>BP069281
Length = 468
Score = 84.3 bits (207), Expect = 2e-17
Identities = 33/59 (55%), Positives = 47/59 (78%)
Frame = -3
Query: 193 DLSECGKKGVAVNPRRGDALLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSF 251
+LS+CGKKG+++ P+RGDALLF S+ P+ DT SLH GCPV +G+KWS TKW+ ++ +
Sbjct: 445 ELSDCGKKGLSIKPKRGDALLFWSMKPDTTADTSSLHGGCPVTKGDKWSCTKWMRLNEY 269
>AV766573
Length = 338
Score = 70.9 bits (172), Expect = 3e-13
Identities = 29/48 (60%), Positives = 37/48 (76%)
Frame = -3
Query: 204 VNPRRGDALLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSF 251
V P+RGDALLF S+ P+A D SLH GCPVI G KWS+TKW+H++ +
Sbjct: 336 VKPKRGDALLFWSIRPDATLDPSSLHGGCPVIRGNKWSSTKWMHLEEY 193
>TC19415 similar to UP|BAD07294 (BAD07294) Prolyl 4-hydroxylase, partial
(24%)
Length = 443
Score = 59.7 bits (143), Expect = 6e-10
Identities = 28/46 (60%), Positives = 30/46 (64%)
Frame = +3
Query: 201 GVAVNPRRGDALLFCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWI 246
G+ V PR GD LLF SL PN D SL CPVI+GEKW A KWI
Sbjct: 51 GLRVRPRPGDGLLFYSLFPNGTIDPTSLPGSCPVIKGEKWVAPKWI 188
>AV774533
Length = 499
Score = 46.2 bits (108), Expect(3) = 8e-10
Identities = 21/57 (36%), Positives = 32/57 (55%), Gaps = 4/57 (7%)
Frame = -2
Query: 216 SLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHVDSFD----KTVGAGGDCTDQHESC 268
SLHP + ++ CP +G+ WSA K+ + DS T+ GG+CTD+ +SC
Sbjct: 498 SLHPVHLRTRVATLLRCPAPKGDMWSAIKFFYADSITGGKLSTISDGGECTDEDDSC 328
Score = 28.5 bits (62), Expect(3) = 8e-10
Identities = 10/13 (76%), Positives = 11/13 (83%)
Frame = -3
Query: 271 WAALGECTKNPEY 283
WAALGEC +NP Y
Sbjct: 320 WAALGECQRNPVY 282
Score = 23.5 bits (49), Expect(3) = 8e-10
Identities = 11/25 (44%), Positives = 12/25 (48%)
Frame = -1
Query: 277 CTKNPEYMVGTSGLPGYCRKSCKTC 301
CT N G+ G CRKSC C
Sbjct: 286 CTMN-----GSPDYYGTCRKSCNAC 227
>AW720438
Length = 463
Score = 58.5 bits (140), Expect = 1e-09
Identities = 23/42 (54%), Positives = 36/42 (84%)
Frame = +3
Query: 35 IDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSA 76
+DP++V Q+SW+PR F+YKGFL+D ECD+LIS+A + ++S+
Sbjct: 333 VDPSRVVQISWQPRVFLYKGFLSDKECDYLISLAYAVKEKSS 458
>TC15896 similar to UP|BAD07294 (BAD07294) Prolyl 4-hydroxylase, partial
(45%)
Length = 613
Score = 56.6 bits (135), Expect = 5e-09
Identities = 32/91 (35%), Positives = 52/91 (56%), Gaps = 3/91 (3%)
Frame = +3
Query: 43 VSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNL-SGESKLSEVRTSSGMFIS-- 99
+SW P A + F T +C+ +I AK LK S + + + + +RTSSG+FIS
Sbjct: 339 LSWNPHALYFPNFATAEQCESIIETAKEGLKPSTLVLRVGETDESTTGIRTSSGVFISAF 518
Query: 100 KNKDAIVSGIEDKISSWTFLPKENGEDIQVL 130
++K ++ IE+KI+ T +P+ +GE VL
Sbjct: 519 EDKTGVLDVIEEKIARATKIPRTHGEAFNVL 611
>BP061999
Length = 564
Score = 51.6 bits (122), Expect = 2e-07
Identities = 20/35 (57%), Positives = 26/35 (74%)
Frame = -2
Query: 214 FCSLHPNAIPDTLSLHAGCPVIEGEKWSATKWIHV 248
F S+ P+ DT SLH GCPVI+G+KWS TKW+ +
Sbjct: 563 FWSMKPDTTADTSSLHGGCPVIKGDKWSCTKWMRL 459
>AV423632
Length = 484
Score = 51.2 bits (121), Expect = 2e-07
Identities = 22/41 (53%), Positives = 30/41 (72%)
Frame = +2
Query: 40 VKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADN 80
V+ +SW+PRAFVY FLT EC++LI IAK + +S V D+
Sbjct: 359 VEVISWEPRAFVYHNFLTKEECEYLIDIAKPNMHKSTVVDS 481
>TC16912 similar to UP|BAD07294 (BAD07294) Prolyl 4-hydroxylase, partial
(30%)
Length = 518
Score = 50.8 bits (120), Expect = 3e-07
Identities = 29/79 (36%), Positives = 44/79 (54%), Gaps = 1/79 (1%)
Frame = +1
Query: 26 GSYAGTSAIIDPTKVKQVSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVA-DNLSGE 84
G + S P +V +SWKPRA + F T +C ++ +AK+ LK SA+A E
Sbjct: 256 GEFGDDSITSIPFQV--LSWKPRALYFPNFATAEQCKSIVGVAKAGLKPSALALREGETE 429
Query: 85 SKLSEVRTSSGMFISKNKD 103
+RTSSG+F+S ++D
Sbjct: 430 QNTKGIRTSSGVFVSSSED 486
>TC12024
Length = 469
Score = 50.4 bits (119), Expect = 4e-07
Identities = 18/43 (41%), Positives = 35/43 (80%)
Frame = +1
Query: 43 VSWKPRAFVYKGFLTDLECDHLISIAKSELKRSAVADNLSGES 85
+SW+PRAF++ FL+ +EC+++I++AK + +S+V D+ +G+S
Sbjct: 340 LSWEPRAFIFHNFLSKVECEYMINLAKPHMAKSSVVDSQTGKS 468
>TC16820 similar to UP|Q8L8T9 (Q8L8T9) Prolyl 4-hydroxylase alpha
subunit-like protein, partial (7%)
Length = 571
Score = 38.9 bits (89), Expect = 0.001
Identities = 14/20 (70%), Positives = 17/20 (85%)
Frame = +3
Query: 227 SLHAGCPVIEGEKWSATKWI 246
S+H GC V+ GEKWSATKW+
Sbjct: 369 SVHGGCEVLAGEKWSATKWM 428
>AV775794
Length = 366
Score = 29.6 bits (65), Expect = 0.65
Identities = 13/27 (48%), Positives = 18/27 (66%)
Frame = +1
Query: 140 PHYDYFADKVNIARGGHRVATVLMYLT 166
PH+ +F VNIAR G RVA++ +T
Sbjct: 61 PHHSFFKHNVNIARRG*RVASLSQSIT 141
>AV407855
Length = 420
Score = 26.6 bits (57), Expect = 5.5
Identities = 16/56 (28%), Positives = 25/56 (44%)
Frame = +1
Query: 2 SVICRVWCSIIVPLLLICKIHFALGSYAGTSAIIDPTKVKQVSWKPRAFVYKGFLT 57
+V+C C+ + LLL I GS G + + P+ F+ KGFL+
Sbjct: 97 AVVCSALCAPSLLLLLYFSIRLPSGSNVGLRITYFTSL*RCSQQMPKVFLKKGFLS 264
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.422
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,333,284
Number of Sequences: 28460
Number of extensions: 95713
Number of successful extensions: 437
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 428
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 429
length of query: 301
length of database: 4,897,600
effective HSP length: 90
effective length of query: 211
effective length of database: 2,336,200
effective search space: 492938200
effective search space used: 492938200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146553.10


