

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
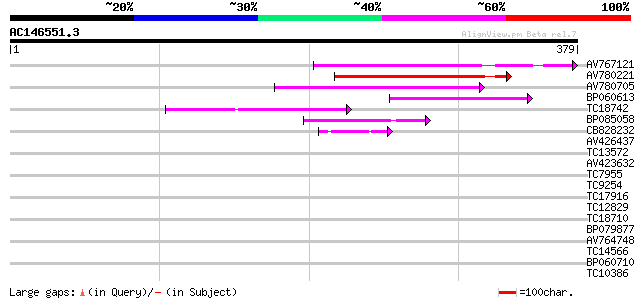
Query= AC146551.3 - phase: 1 /pseudo
(379 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV767121 127 3e-30
AV780221 108 1e-24
AV780705 72 2e-13
BP060613 64 3e-11
TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%) 49 1e-06
BP085058 43 8e-05
CB828232 40 8e-04
AV426437 39 0.001
TC13572 26 0.24
AV423632 30 0.51
TC7955 similar to UP|WR11_ARATH (Q9SV15) Probable WRKY transcrip... 30 0.51
TC9254 UP|CAE45592 (CAE45592) Transcription factor homolog BTF3-... 30 0.66
TC17916 weakly similar to PIR|B97853|B97853 NADH2 dehydrogenase ... 29 1.1
TC12829 29 1.1
TC18710 29 1.1
BP079877 29 1.5
AV764748 27 4.3
TC14566 similar to UP|SRP_PHAVU (Q41112) Stress-related protein ... 27 5.6
BP060710 27 7.3
TC10386 similar to UP|Q9LU67 (Q9LU67) Phosphatidylserine decarbo... 27 7.3
>AV767121
Length = 525
Score = 127 bits (319), Expect = 3e-30
Identities = 68/176 (38%), Positives = 98/176 (55%)
Frame = +2
Query: 204 LSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIISSIESIFKSFFWGGGEENRK 263
LS W K +S GGR+ L++ VL++LP++FLSFFK P G+ S + ++F WGG E K
Sbjct: 2 LSKWKQKTVSFGGRIFLIQSVLTALPLFFLSFFKLPIGVGKSCVRLMRNFLWGGSENENK 181
Query: 264 IAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTERDELWYRVLKAKYGEEGGR 323
IA +KW +C K GGLG++ + FN +LLGK WR LTE D LW RV++A+
Sbjct: 182 IA*VKWTDVCKPKELGGLGIKDLFTFNKALLGK*RWRYLTEPDSLWRRVIEAQ------- 340
Query: 324 IREGGRLSSAWWKMVCHIREGLVDWGFTVTEEVSSSFQFDDGKGEYGEGYGEAWLG 379
+ S+WW + + D F SS + G+G+ + + E WLG
Sbjct: 341 -PDHYSCGSSWWNDILSLCPEDADGWF------SSGLKKLVGEGDQTKFWSEDWLG 487
>AV780221
Length = 461
Score = 108 bits (270), Expect = 1e-24
Identities = 55/119 (46%), Positives = 74/119 (61%), Gaps = 1/119 (0%)
Frame = +3
Query: 218 LVLLKVVLSSLPVYFLSFFKAPAG-IISSIESIFKSFFWGGGEENRKIAWIKWDTICLSK 276
L L++ VL +LP++ L FFK P G +++ + FF E +KIAW+KW T+C K
Sbjct: 3 LCLIRSVLIALPLFHLPFFKLPTGGFYTNVNN**DPFFGREVEGGKKIAWVKWSTVCRPK 182
Query: 277 AEGGLGVRRMGAFNVSLLGKWCWRMLTERDELWYRVLKAKYGEEGGRIREGGRLSSAWW 335
EGGLGV+ +G FN +LLGKW WRML ERD LWY+VL KY + +S+WW
Sbjct: 183 DEGGLGVQNLGLFNKALLGKWRWRMLKERDGLWYKVLLIKYK------NAIPQSASSWW 341
>AV780705
Length = 524
Score = 71.6 bits (174), Expect = 2e-13
Identities = 40/141 (28%), Positives = 71/141 (49%), Gaps = 1/141 (0%)
Frame = -2
Query: 178 YLGVPIGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFK 237
YLG+P K + +++ +L W LL+ G+ VL+K ++ ++P Y ++
Sbjct: 484 YLGLPAIWGRNKSHSLVWIEEKVKEKLEGWKETLLNQAGKEVLIKAIIQAIPSYAMTIVH 305
Query: 238 APAGIISSIESIFKSFFWGG-GEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGK 296
P +S+ ++ F+W GE + LSK +GG+G R + N++ L +
Sbjct: 304 FPKTFCNSLYALVADFWWKSQGEGAVEYTGAVGTK*PLSKEKGGVGFRDLRTQNLAFLAR 125
Query: 297 WCWRMLTERDELWYRVLKAKY 317
WR+LT + LW RV+K+ Y
Sbjct: 124 QAWRVLTNPEALWVRVMKSLY 62
>BP060613
Length = 378
Score = 64.3 bits (155), Expect = 3e-11
Identities = 34/96 (35%), Positives = 48/96 (49%), Gaps = 1/96 (1%)
Frame = +1
Query: 255 WGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTERDELWYRVLK 314
W + R + W KWD + K EGGLG + N +LL K WR+L D LW ++LK
Sbjct: 46 WWSSKGTRGVHWRKWDLLTELKDEGGLGFKDFEIQNQALLAKQAWRILHNPDALWVQILK 225
Query: 315 AKYGEEGGRIREGGRLSSAW-WKMVCHIREGLVDWG 349
A Y ++ R ++W W + H RE L+ G
Sbjct: 226 ALYFPHHDFLQTTKRTGASWVWSSLLHGRELLLKQG 333
>TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%)
Length = 912
Score = 49.3 bits (116), Expect = 1e-06
Identities = 34/124 (27%), Positives = 61/124 (48%)
Frame = -3
Query: 105 VTHLQFADDTLIIGEKCWLNVRTMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWLVEAS 164
++HL FAD+ ++ ++ + VL +F+ +SGL N KS + + + ++ + +
Sbjct: 373 ISHLCFADNLMVFSNGYLESIAIINNVLHIFQHLSGLTPNPAKSEVF-ILVKETTKNQIT 197
Query: 165 ALMNCCRGNFTFVYLGVPIGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVV 224
++ G LGVP K S K ++DR +L W + LS GRL L+ +
Sbjct: 196 NMLGYKEGKLPVR*LGVPQLTTKLKASDCKVLVDRKPSKLQHWTGRSLSYAGRLQLINSI 17
Query: 225 LSSL 228
L S+
Sbjct: 16 LFSM 5
>BP085058
Length = 452
Score = 43.1 bits (100), Expect = 8e-05
Identities = 27/85 (31%), Positives = 42/85 (48%)
Frame = +2
Query: 197 IDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIISSIESIFKSFFWG 256
+DR++ RL+ LL+ RL L K VLS++ +Y + P I I ++ + F W
Sbjct: 194 LDRVISRLAC*KAYLLNKPARLTLAKPVLSAISMYVMQLNWLPQSICDQICTVVRHFVW- 370
Query: 257 GGEENRKIAWIKWDTICLSKAEGGL 281
E I + W+TI K+ GL
Sbjct: 371 --ENVNGIPLVAWETIQSRKSMVGL 439
>CB828232
Length = 532
Score = 39.7 bits (91), Expect = 8e-04
Identities = 22/50 (44%), Positives = 28/50 (56%)
Frame = +1
Query: 207 WNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIISSIESIFKSFFWG 256
W LS G R+ K+VLSSLP++FLSFFK + + I F WG
Sbjct: 388 WKTLCLS-GARVCCFKMVLSSLPLFFLSFFKIKC-VAKVCKQIQSRFLWG 531
>AV426437
Length = 303
Score = 39.3 bits (90), Expect = 0.001
Identities = 17/29 (58%), Positives = 19/29 (64%)
Frame = +1
Query: 259 EENRKIAWIKWDTICLSKAEGGLGVRRMG 287
E RKIAWI W+TIC K GLGV+ G
Sbjct: 214 ENQRKIAWINWETICKPKWLRGLGVKNWG 300
>TC13572
Length = 534
Score = 26.2 bits (56), Expect(2) = 0.24
Identities = 10/19 (52%), Positives = 13/19 (67%)
Frame = -1
Query: 290 NVSLLGKWCWRMLTERDEL 308
N +LLGKW WR+ T + L
Sbjct: 534 NKALLGKWLWRLKTVPNSL 478
Score = 23.9 bits (50), Expect(2) = 0.24
Identities = 13/31 (41%), Positives = 18/31 (57%), Gaps = 1/31 (3%)
Frame = -3
Query: 309 WYRVLKAKYGEEGGRIREGGRLS-SAWWKMV 338
W RV+ AKYG +GG + S+WW+ V
Sbjct: 478 WCRVVLAKYG-------DGGCTNISSWWRDV 407
>AV423632
Length = 484
Score = 30.4 bits (67), Expect = 0.51
Identities = 20/70 (28%), Positives = 31/70 (43%), Gaps = 2/70 (2%)
Frame = -3
Query: 296 KWCWRMLTERDELWYRVLKAKYGEEGGRIREGGRLSSAWWKMVCHIREG--LVDWGFTVT 353
+W W E L +R + + G GG+ R G R AW K + EG ++ +
Sbjct: 290 RWVWGTCHEMRSLEWRGCRGRRG*GGGK*RRGTRRR*AWKKTMISEMEGESILVSPLLMV 111
Query: 354 EEVSSSFQFD 363
EE S +F+
Sbjct: 110 EEARSDLRFN 81
>TC7955 similar to UP|WR11_ARATH (Q9SV15) Probable WRKY transcription
factor 11 (WRKY DNA-binding protein 11), partial (27%)
Length = 579
Score = 30.4 bits (67), Expect = 0.51
Identities = 16/49 (32%), Positives = 23/49 (46%), Gaps = 2/49 (4%)
Frame = +1
Query: 276 KAEGGLGVRRMGAFNVSLLGKWCWRMLTERDELWYR--VLKAKYGEEGG 322
+ G G R +LLG W W ++R+ W+ +LK K G GG
Sbjct: 241 RTRGSTGTRFRPRCTRTLLGLWVW-FFSQRERRWWNTIILKKKVGGNGG 384
>TC9254 UP|CAE45592 (CAE45592) Transcription factor homolog BTF3-like
protein, complete
Length = 758
Score = 30.0 bits (66), Expect = 0.66
Identities = 11/21 (52%), Positives = 15/21 (71%)
Frame = +3
Query: 316 KYGEEGGRIREGGRLSSAWWK 336
++ +E G+I E GR SS WWK
Sbjct: 36 QHKDESGKIDEDGRFSSNWWK 98
>TC17916 weakly similar to PIR|B97853|B97853 NADH2 dehydrogenase
(ubiquinone) - Rickettsia conorii
(strain Malish 7) {Rickettsia conorii;} , partial (3%)
Length = 520
Score = 29.3 bits (64), Expect = 1.1
Identities = 14/45 (31%), Positives = 24/45 (53%)
Frame = -2
Query: 231 YFLSFFKAPAGIISSIESIFKSFFWGGGEENRKIAWIKWDTICLS 275
+FLSF +G+ ++ S+ K+ WG + + AWI + LS
Sbjct: 357 FFLSFKFLLSGVEITLVSLIKTSIWGKEKNEEQFAWICLELDALS 223
>TC12829
Length = 448
Score = 29.3 bits (64), Expect = 1.1
Identities = 14/48 (29%), Positives = 21/48 (43%)
Frame = +1
Query: 226 SSLPVYFLSFFKAPAGIISSIESIFKSFFWGGGEENRKIAWIKWDTIC 273
S L + FL F + +++ SF W G R I W+ W +C
Sbjct: 322 SCLALLFLILF------VLRLKAWLVSFIWSGDVSRRSIHWLGWKKLC 447
>TC18710
Length = 843
Score = 29.3 bits (64), Expect = 1.1
Identities = 17/52 (32%), Positives = 26/52 (49%), Gaps = 6/52 (11%)
Frame = -3
Query: 290 NVSLLGKWCWRMLTERDELWYRVLKAKYGE------EGGRIREGGRLSSAWW 335
N+S L K L RDE+W+ +LK+ Y + E G +G +L +W
Sbjct: 487 NLSSLNK---ATLHRRDEIWHGILKSSYKDF*DNFIEEGATTDGPKLIDGFW 341
>BP079877
Length = 381
Score = 28.9 bits (63), Expect = 1.5
Identities = 15/30 (50%), Positives = 22/30 (73%)
Frame = -2
Query: 225 LSSLPVYFLSFFKAPAGIISSIESIFKSFF 254
+ ++P+YFLSFF + +ISS+ S FKS F
Sbjct: 101 VKTIPLYFLSFFLLISFVISSLFS-FKSHF 15
>AV764748
Length = 260
Score = 27.3 bits (59), Expect = 4.3
Identities = 11/36 (30%), Positives = 21/36 (57%)
Frame = +3
Query: 240 AGIISSIESIFKSFFWGGGEENRKIAWIKWDTICLS 275
+G+ ++ S+ K+ FWG + ++ AWI + LS
Sbjct: 12 SGVEITLVSLIKTSFWGKEKNEKQFAWICLELDALS 119
>TC14566 similar to UP|SRP_PHAVU (Q41112) Stress-related protein (PvSRP),
partial (97%)
Length = 1198
Score = 26.9 bits (58), Expect = 5.6
Identities = 12/34 (35%), Positives = 17/34 (49%)
Frame = -3
Query: 323 RIREGGRLSSAWWKMVCHIREGLVDWGFTVTEEV 356
R R GG W M +R+GLV+ V E++
Sbjct: 485 RSRLGGHALDVGWNMAVDLRDGLVNLAIDVAEKL 384
>BP060710
Length = 534
Score = 26.6 bits (57), Expect = 7.3
Identities = 17/39 (43%), Positives = 20/39 (50%), Gaps = 1/39 (2%)
Frame = +2
Query: 226 SSLPVYFLSFFKAPAGI-ISSIESIFKSFFWGGGEENRK 263
SSL YFLS F P G S+ F FF G EN++
Sbjct: 122 SSLDSYFLSLFPFPLGASFFSLG*KFS*FFLGASLENKR 238
>TC10386 similar to UP|Q9LU67 (Q9LU67) Phosphatidylserine decarboxylase,
partial (42%)
Length = 788
Score = 26.6 bits (57), Expect = 7.3
Identities = 10/21 (47%), Positives = 16/21 (75%)
Frame = +3
Query: 114 TLIIGEKCWLNVRTMMAVLLL 134
+L+ G+ WLNV T++ VLL+
Sbjct: 570 SLVQGQLLWLNVTTLLCVLLI 632
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.331 0.146 0.494
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,081,647
Number of Sequences: 28460
Number of extensions: 129111
Number of successful extensions: 848
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 841
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 846
length of query: 379
length of database: 4,897,600
effective HSP length: 92
effective length of query: 287
effective length of database: 2,279,280
effective search space: 654153360
effective search space used: 654153360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146551.3


