

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
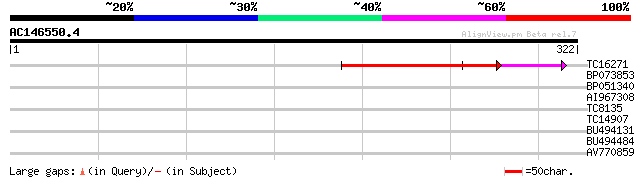
Query= AC146550.4 - phase: 0 /pseudo
(322 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC16271 similar to GB|AAM15909.1|20257479|AF492660 purple acid p... 115 1e-26
BP073853 29 1.2
BP051340 28 2.1
AI967308 27 4.6
TC8135 similar to UP|GUN4_ARATH (Q9LX31) Tetrapyrrole-binding pr... 27 6.1
TC14907 similar to GB|BAB10858.1|10177467|AB009053 permease 1 {A... 27 6.1
BU494131 27 6.1
BU494484 26 7.9
AV770859 26 7.9
>TC16271 similar to GB|AAM15909.1|20257479|AF492660 purple acid phosphatase
{Arabidopsis thaliana;} , partial (32%)
Length = 653
Score = 115 bits (287), Expect = 1e-26
Identities = 52/91 (57%), Positives = 69/91 (75%)
Frame = +1
Query: 189 VLPRESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNI 248
VLPR+ Y + LL+++++AL S AKWKIVVGHH I+S+GHHG+T+EL +QLLPIL+ NN+
Sbjct: 1 VLPRDKYLSNLLRDLEIALKVSTAKWKIVVGHHPIRSIGHHGDTKELIRQLLPILEENNV 180
Query: 249 DAYINGHDHCLQHIIDNERHGEAISSHGTRK 279
D YINGHDHCL+HI ++S G K
Sbjct: 181 DMYINGHDHCLEHITSTNSQILFLTSGGGSK 273
Score = 43.9 bits (102), Expect = 4e-05
Identities = 25/59 (42%), Positives = 31/59 (52%)
Frame = +2
Query: 258 CLQHIIDNERHGEAISSHGTRK*SFTMTDKDSCLCRSQK*M*MLYSMTFWARFCIDGAY 316
C +++ RHG+ SFTMTDKDSCL K M L+ M F A+FCI Y
Sbjct: 245 CS*PVVEVRRHGKETFRIRQMGSSFTMTDKDSCLWSFNKSMPKLFIMIFLAKFCIFRIY 421
>BP073853
Length = 386
Score = 28.9 bits (63), Expect = 1.2
Identities = 20/55 (36%), Positives = 27/55 (48%), Gaps = 1/55 (1%)
Frame = -1
Query: 266 ERHGEAISSHGTRK-*SFTMTDKDSCLCRSQK*M*MLYSMTFWARFCIDGAYRKN 319
+R GE T++ * +M DK SC C + M L SM F A FC + + N
Sbjct: 386 QRLGEETFKKRTKEV*ISSMMDKVSCQCN*LRLMQTLSSMMFLAMFCTESFLQSN 222
>BP051340
Length = 537
Score = 28.1 bits (61), Expect = 2.1
Identities = 13/28 (46%), Positives = 18/28 (63%)
Frame = -2
Query: 139 SILRQKDSRWVCLRSFILDGGIVEFFFV 166
SI+ K RWVCL+ FI + +FFF+
Sbjct: 89 SIVETK*LRWVCLQVFIYNVLTEKFFFI 6
>AI967308
Length = 443
Score = 26.9 bits (58), Expect = 4.6
Identities = 23/75 (30%), Positives = 37/75 (48%), Gaps = 7/75 (9%)
Frame = +1
Query: 149 VCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAEL-------LK 201
+CL+ F L+ +EF TT IEK T P E + P ++ + E+ LK
Sbjct: 52 MCLKLFQLEKVAIEFTKPATTDQIEKE-TQPSE-----DSTTPPDAPKPEIPCVDEMTLK 213
Query: 202 NVDLALVKSKAKWKI 216
N ++AL+K + K+
Sbjct: 214 NDEIALLKCNLEEKV 258
>TC8135 similar to UP|GUN4_ARATH (Q9LX31) Tetrapyrrole-binding protein,
chloroplast precursor (Genomes uncoulped 4), partial
(51%)
Length = 1213
Score = 26.6 bits (57), Expect = 6.1
Identities = 14/45 (31%), Positives = 25/45 (55%), Gaps = 4/45 (8%)
Frame = -1
Query: 45 QQSLNFL----VVGDWGRKGNYNQSLVAHQMGIVGDNLNIDFVIS 85
Q+ LN L V G W +K +S + + +VGD L I+++++
Sbjct: 1135 QEKLNKLQMKNVKGHW*KKRK**KSTIKQPLSLVGDKLTINYILN 1001
>TC14907 similar to GB|BAB10858.1|10177467|AB009053 permease 1 {Arabidopsis
thaliana;} , partial (28%)
Length = 734
Score = 26.6 bits (57), Expect = 6.1
Identities = 9/18 (50%), Positives = 12/18 (66%)
Frame = -2
Query: 173 EKYFTDPKEHTYDWNGVL 190
+KY T P + +DWNG L
Sbjct: 307 KKYATTPATNAFDWNGTL 254
>BU494131
Length = 381
Score = 26.6 bits (57), Expect = 6.1
Identities = 12/32 (37%), Positives = 19/32 (58%)
Frame = +2
Query: 68 AHQMGIVGDNLNIDFVISTGDNFYKDGLEGVD 99
+H + GD ++I FV+ D+ + G EGVD
Sbjct: 218 SHDVKYEGDTVHISFVVIKADSPWHYGDEGVD 313
>BU494484
Length = 386
Score = 26.2 bits (56), Expect = 7.9
Identities = 15/49 (30%), Positives = 23/49 (46%)
Frame = -2
Query: 43 PQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGDNLNIDFVISTGDNFY 91
PQ+ ++F WG N+ LVA + I+G F+ T NF+
Sbjct: 232 PQKALISF*RTIRWGNHSNFK*KLVAFCLPIIGATKLYPFINPT*LNFF 86
>AV770859
Length = 198
Score = 26.2 bits (56), Expect = 7.9
Identities = 12/27 (44%), Positives = 17/27 (62%)
Frame = -3
Query: 148 WVCLRSFILDGGIVEFFFVDTTPFIEK 174
++CL F IV FF+V +TPFI +
Sbjct: 184 FICLFPFYPTPYIVLFFYVTSTPFIRE 104
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.327 0.141 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,383,639
Number of Sequences: 28460
Number of extensions: 93119
Number of successful extensions: 507
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 503
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 507
length of query: 322
length of database: 4,897,600
effective HSP length: 90
effective length of query: 232
effective length of database: 2,336,200
effective search space: 541998400
effective search space used: 541998400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146550.4


